BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= QBL17f07.xg.2.1
(657 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
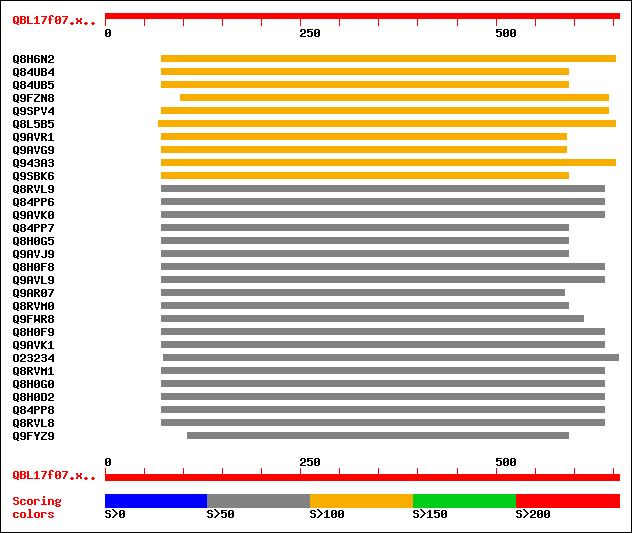
Sequences producing significant alignments: (bits) Value
sptr|Q8H6N2|Q8H6N2 S-adenosyl-L-methionine:salicylic acid methyl... 113 2e-24
sptr|Q84UB4|Q84UB4 S-adenosyl-L-methionine:benzoic acid/salicyli... 112 5e-24
sptr|Q84UB5|Q84UB5 S-adenosyl-L-methionine:benzoic acid/salicyli... 112 5e-24
sptr|Q9FZN8|Q9FZN8 Caffeine synthase. 109 3e-23
sptr|Q9SPV4|Q9SPV4 S-adenosyl-L-methionine:salicylic acid carbox... 107 1e-22
sptr|Q8L5B5|Q8L5B5 Putative S-adenosyl-L-methionine:salicylic ac... 107 1e-22
sptr|Q9AVR1|Q9AVR1 S-adenosyl-L-methionine:salicylic acid carbox... 106 3e-22
sptr|Q9AVG9|Q9AVG9 S-adenosyl-L-methionine:salicylic acid carbox... 105 5e-22
sptr|Q943A3|Q943A3 Putative S-adenosyl-L-methionine:salicylic ac... 105 5e-22
sptr|Q9SBK6|Q9SBK6 Floral nectary-specific protein. 101 7e-21
sptr|Q8RVL9|Q8RVL9 N-methyltransferase. 99 3e-20
sptr|Q84PP6|Q84PP6 Xanthosine N-methyltransferase. 99 5e-20
sptr|Q9AVK0|Q9AVK0 Theobromine synthase (N-methyltransferase) (T... 99 5e-20
sptr|Q84PP7|Q84PP7 7-methylxanthine N-methyltransferase. 98 8e-20
sptr|Q8H0G5|Q8H0G5 Theobromine synthase 1. 98 8e-20
sptr|Q9AVJ9|Q9AVJ9 7-methylxanthine N-methyltransferase. 98 8e-20
sptr|Q8H0F8|Q8H0F8 Tentative caffeine synthase 4. 98 1e-19
sptr|Q9AVL9|Q9AVL9 Caffeine synthase. 98 1e-19
sptr|Q9AR07|Q9AR07 S-adenosyl-L-methionine:jasmonic acid carboxy... 98 1e-19
sptr|Q8RVM0|Q8RVM0 N-methyltransferase. 98 1e-19
sptr|Q9FWR8|Q9FWR8 F14P1.3 protein. 98 1e-19
sptr|Q8H0F9|Q8H0F9 Tentative caffeine synthase 3. 97 2e-19
sptr|Q9AVK1|Q9AVK1 Theobromine synthase. 97 2e-19
sptr|O23234|O23234 Hypothetical 42.0 kDa protein. 96 4e-19
sptr|Q8RVM1|Q8RVM1 N-methyltransferase. 96 5e-19
sptr|Q8H0G0|Q8H0G0 Theobromine synthse 2. 95 7e-19
sptr|Q8H0D2|Q8H0D2 Tentative caffeine synthase 7. 93 3e-18
sptr|Q84PP8|Q84PP8 3,7-dimethylxanthine N-methyltransferase. 92 4e-18
sptr|Q8RVL8|Q8RVL8 N-methyltransferase. 92 6e-18
sptr|Q9FYZ9|Q9FYZ9 SAM:benzoic acid carboxyl methyltransferase. 90 3e-17
>sptr|Q8H6N2|Q8H6N2 S-adenosyl-L-methionine:salicylic acid
methyltransferase.
Length = 383
Score = 113 bits (283), Expect = 2e-24
Identities = 71/200 (35%), Positives = 100/200 (50%), Gaps = 6/200 (3%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP--RTMVVADLG 245
M + + HM G G+ SY+ N Q+ + T+P++E A+ + Y +LP T+ +ADLG
Sbjct: 12 MKLAQVLHMNGGLGKSSYASNSLVQRKVISITKPIIEEAMTEFYTRMLPSPHTISIADLG 71
Query: 246 CSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRS 425
CS GPNTLL + ++ I + + P E Q LNDLPGNDFN +FR
Sbjct: 72 CSCGPNTLLVAAELVKIIVKLRQKLDREPPP---------EFQIHLNDLPGNDFNSIFRY 122
Query: 426 V----EEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEG 593
+ EE R G +V G+P S+Y RLFP +S+H SSY + PEG
Sbjct: 123 LLPMFREELREEIGGGEEAGRRCFVSGVPGSFYGRLFPTKSLHFVHSSYSLMWLSKVPEG 182
Query: 594 XXAXRKPCLNXXXIYIAXXT 653
+N IYIA +
Sbjct: 183 VK------MNKENIYIASTS 196
>sptr|Q84UB4|Q84UB4 S-adenosyl-L-methionine:benzoic acid/salicylic
acid carboxyl methyltransferase.
Length = 357
Score = 112 bits (279), Expect = 5e-24
Identities = 65/178 (36%), Positives = 93/178 (52%), Gaps = 4/178 (2%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRTMVVADLGCS 251
M++ HM G G+ SY+ N QQ + T+P+ E A+ +Y++L P T+ +ADLGCS
Sbjct: 1 MEVVEVLHMNGGNGDSSYANNSLVQQKVILMTKPITEQAMIDLYSSLFPETLCIADLGCS 60
Query: 252 AGPNTLLFISSVLSSIXXXXXEQ-CKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRSV 428
G NT L +S ++ + + K P E F NDLPGNDFN LF+S+
Sbjct: 61 LGANTFLVVSQLVKIVEKERKKHGFKSP-----------EFYFHFNDLPGNDFNTLFQSL 109
Query: 429 ---EEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEG 593
+E+ R+ G P + G+P S+Y RLFP +S+H SSY Q P G
Sbjct: 110 GAFQEDLRKHIG---ESFGPCFFSGVPGSFYTRLFPSKSLHFVYSSYSLMWLSQVPNG 164
>sptr|Q84UB5|Q84UB5 S-adenosyl-L-methionine:benzoic acid/salicylic
acid carboxyl methyltransferase.
Length = 357
Score = 112 bits (279), Expect = 5e-24
Identities = 65/178 (36%), Positives = 93/178 (52%), Gaps = 4/178 (2%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRTMVVADLGCS 251
M++ HM G G+ SY+ N QQ + T+P+ E A+ +Y++L P T+ +ADLGCS
Sbjct: 1 MEVVEVLHMNGGNGDSSYANNSLVQQKVILMTKPITEQAMIDLYSSLFPETLCIADLGCS 60
Query: 252 AGPNTLLFISSVLSSIXXXXXEQ-CKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRSV 428
G NT L +S ++ + + K P E F NDLPGNDFN LF+S+
Sbjct: 61 LGANTFLVVSQLVKIVEKERKKHGFKSP-----------EFYFHFNDLPGNDFNTLFQSL 109
Query: 429 ---EEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEG 593
+E+ R+ G P + G+P S+Y RLFP +S+H SSY Q P G
Sbjct: 110 GAFQEDLRKHIG---ESFGPCFFSGVPGSFYTRLFPSKSLHFVYSSYSLMWLSQVPNG 164
>sptr|Q9FZN8|Q9FZN8 Caffeine synthase.
Length = 369
Score = 109 bits (272), Expect = 3e-23
Identities = 70/184 (38%), Positives = 98/184 (53%), Gaps = 1/184 (0%)
Frame = +3
Query: 96 MAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP-RTMVVADLGCSAGPNTLL 272
M GEGE SY++N Q +P +E AV+ +++ + + ADLGC+AGPNT
Sbjct: 15 MNRGEGESSYAQNSSFTQQVASMAQPALENAVETLFSRDFHLQALNAADLGCAAGPNTF- 73
Query: 273 FISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRSVEEEFRRAA 452
+V+S+I ++C+ + +ELQ LNDL GNDFN LF+ + E
Sbjct: 74 ---AVISTIKRMMEKKCRELNCQT------LELQVYLNDLFGNDFNTLFKGLSSEVI-GN 123
Query: 453 GCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXAXRKPCLNXXX 632
CE P YVMG+P S++ RLFPR S+HL SSY Q P+G + LN
Sbjct: 124 KCEEVP---CYVMGVPGSFHGRLFPRNSLHLVHSSYSVHWLTQAPKGLTSREGLALNKGK 180
Query: 633 IYIA 644
IYI+
Sbjct: 181 IYIS 184
>sptr|Q9SPV4|Q9SPV4 S-adenosyl-L-methionine:salicylic acid carboxyl
methyltransferase.
Length = 359
Score = 107 bits (268), Expect = 1e-22
Identities = 66/192 (34%), Positives = 97/192 (50%), Gaps = 1/192 (0%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRT-MVVADLGC 248
MD+R+ HM G GE SY+ N Q+ + T+P+ E A+ +Y+ T + +ADLGC
Sbjct: 1 MDVRQVLHMKGGAGENSYAMNSFIQRQVISITKPITEAAITALYSGDTVTTRLAIADLGC 60
Query: 249 SAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRSV 428
S+GPN L ++ ++ ++ + + S E Q LNDLPGNDFN +FRS+
Sbjct: 61 SSGPNALFAVTELIKTVEELRKKMGRENSP---------EYQIFLNDLPGNDFNAIFRSL 111
Query: 429 EEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXAXR 608
E C ++ G+P S+Y RLFPR ++H SSY Q P G +
Sbjct: 112 PIENDVDGVC--------FINGVPGSFYGRLFPRNTLHFIHSSYSLMWLSQVPIGIES-- 161
Query: 609 KPCLNXXXIYIA 644
N IY+A
Sbjct: 162 ----NKGNIYMA 169
>sptr|Q8L5B5|Q8L5B5 Putative S-adenosyl-L-methionine:salicylic acid
carboxyl methyltransferase.
Length = 375
Score = 107 bits (267), Expect = 1e-22
Identities = 63/198 (31%), Positives = 93/198 (46%), Gaps = 3/198 (1%)
Frame = +3
Query: 69 YMDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRTMVVADLGC 248
+M++ HM EG GE SY++N R + + ++ + VY + +P VADLGC
Sbjct: 14 FMNVETVLHMKEGLGETSYAQNSR----GMDTLKSLITNSAADVYLSQMPERFAVADLGC 69
Query: 249 SAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRSV 428
S+GPN L ++ SI +PP E +LNDLP NDFN +F S+
Sbjct: 70 SSGPNALCLAEDIIGSIGRICCRSSRPPP----------EFSVLLNDLPTNDFNTIFFSL 119
Query: 429 EE---EFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXX 599
E + AA + P ++ G+P S+Y RLFP +SVH S Q P G
Sbjct: 120 PEFTDRLKAAAKSDEWGRPMVFLSGVPGSFYGRLFPAKSVHFVCSCSSLHWLSQVPSGLL 179
Query: 600 AXRKPCLNXXXIYIAXXT 653
+N +YI+ +
Sbjct: 180 DEMNRPINKGKMYISSTS 197
>sptr|Q9AVR1|Q9AVR1 S-adenosyl-L-methionine:salicylic acid carboxyl
methyltransferase.
Length = 366
Score = 106 bits (264), Expect = 3e-22
Identities = 57/176 (32%), Positives = 92/176 (52%), Gaps = 3/176 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRTMVVADLGCS 251
M++ HM G G+ SY+ N Q+ + T+P+ E A+ ++Y L P+++ +AD+GCS
Sbjct: 1 MEVVEVLHMNGGTGDASYASNSLLQKKVILLTKPITEEAITELYTRLFPKSICIADMGCS 60
Query: 252 AGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRSV- 428
+GPNT L +S ++ ++ E Q LNDLP NDFN +FRS+
Sbjct: 61 SGPNTFLAVSELIKNV----------EKKRTSLGHESPEYQIHLNDLPSNDFNTIFRSLP 110
Query: 429 --EEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPE 590
++ F + G + + G+P S+Y RLFP +S+H SSY + P+
Sbjct: 111 SFQKSFSKQMG---SGFGHCFFTGVPGSFYGRLFPNKSLHFVHSSYSLMWLSRVPD 163
>sptr|Q9AVG9|Q9AVG9 S-adenosyl-L-methionine:salicylic acid carboxyl
methyltransferase.
Length = 357
Score = 105 bits (262), Expect = 5e-22
Identities = 59/176 (33%), Positives = 92/176 (52%), Gaps = 3/176 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRTMVVADLGCS 251
M + HM G G+ SY+ N Q+ + T+P+ E A+ +Y + P T+ +ADLGCS
Sbjct: 1 MKVVEVLHMNGGNGDISYANNSLVQRKVILMTKPITEQAISDLYCSFFPETLCIADLGCS 60
Query: 252 AGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRSV- 428
+G NT L +S ++ + ++ K + H NDLPGNDFN +F+S+
Sbjct: 61 SGANTFLVVSELVKIV----EKERKIHNLQSAGNLFH------FNDLPGNDFNTIFQSLG 110
Query: 429 --EEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPE 590
+++ R+ G E P + G+P S+Y RLFP +S+H SSY Q P+
Sbjct: 111 KFQQDLRKQIGEE---FGPCFFSGVPGSFYTRLFPSESLHFVHSSYSLMWLSQVPD 163
>sptr|Q943A3|Q943A3 Putative S-adenosyl-L-methionine:salicylic acid
carboxyl methyltransferase.
Length = 378
Score = 105 bits (262), Expect = 5e-22
Identities = 60/197 (30%), Positives = 94/197 (47%), Gaps = 3/197 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRTMVVADLGCS 251
M++ HM EG GE SY+KN Q+ ++ + +V + + VYA+L P +ADLGCS
Sbjct: 15 MNVEAVLHMKEGVGETSYAKNSTLQKKSMDTVKSLVTESARDVYASLKPERFTLADLGCS 74
Query: 252 AGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRSVE 431
+G N L + ++ S+ PP E +LNDLP NDFN +F +
Sbjct: 75 SGTNALGMVEEIVRSVAEVCRGSSPPP-----------EFSVLLNDLPTNDFNTIFSRLP 123
Query: 432 E---EFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
E + + A + P ++ G+P S+Y RLFP ++VH S Q P G
Sbjct: 124 EFTGKLKADADADAGDDPMVFLSGVPGSFYGRLFPSKNVHFVCSFSSLHWLSQVPPGLLD 183
Query: 603 XRKPCLNXXXIYIAXXT 653
+N ++I+ +
Sbjct: 184 ETNGPVNKGKMFISSTS 200
>sptr|Q9SBK6|Q9SBK6 Floral nectary-specific protein.
Length = 392
Score = 101 bits (252), Expect = 7e-21
Identities = 65/186 (34%), Positives = 98/186 (52%), Gaps = 12/186 (6%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVY---AALLPRTMVVADL 242
M++ R HM +G GE SY+KN Q + R +++ A+K++ + +L + +ADL
Sbjct: 1 MEVMRILHMNKGNGETSYAKNSIVQSNIISLGRRVMDEALKKLMIRNSEIL--SFGIADL 58
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GCS+GPN+LL IS+++ +I + +P EL LNDLP NDFN++F
Sbjct: 59 GCSSGPNSLLSISNIVETIQNLCHDLDRPVP----------ELSLSLNDLPSNDFNYIFA 108
Query: 423 SVEEEFRR---------AAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXR 575
S+ E + R + G E P +V +P S+Y RLFPR+S+H SS
Sbjct: 109 SLPEFYDRVKKRDNNYESLGFEHGSGGPCFVSAVPGSFYGRLFPRRSLHFVHSSSSLHWL 168
Query: 576 XQEPEG 593
Q P G
Sbjct: 169 SQVPCG 174
>sptr|Q8RVL9|Q8RVL9 N-methyltransferase.
Length = 372
Score = 99.4 bits (246), Expect = 3e-20
Identities = 60/192 (31%), Positives = 93/192 (48%), Gaps = 3/192 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ M GEG+ SY+KN Q+ + +P++E V+++ A LP + + VADL
Sbjct: 1 MELQEVLRMNGGEGDTSYAKNSAYNQLVLAKVKPVLEQCVRELLRANLPNINKCIKVADL 60
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E+ +Q LNDL NDFN +F+
Sbjct: 61 GCASGPNTLLTVRDIVQSIDKVGQEK--------KNELERPTIQIFLNDLFPNDFNSVFK 112
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
+ +R+ + +P S+Y+RLFP +S+H S Y Q Q P G
Sbjct: 113 LLPSFYRKLEKENGRKIGSCLIGAMPGSFYSRLFPEESMHFLHSCYCLQWLSQVPSGLVT 172
Query: 603 XRKPCLNXXXIY 638
N IY
Sbjct: 173 ESGISTNKGSIY 184
>sptr|Q84PP6|Q84PP6 Xanthosine N-methyltransferase.
Length = 385
Score = 99.0 bits (245), Expect = 5e-20
Identities = 60/192 (31%), Positives = 93/192 (48%), Gaps = 3/192 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ M GEG+ SY+KN Q+ + +P++E V+++ A LP + + VADL
Sbjct: 1 MELQEVLRMNGGEGDTSYAKNSAYNQLVLAKVKPVLEQCVRELLRANLPNINKCIKVADL 60
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E+ +Q LNDL NDFN +F+
Sbjct: 61 GCASGPNTLLTVRDIVQSIDKVGQEK--------KNELERPTIQIFLNDLFPNDFNSVFK 112
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
+ +R+ + +P S+Y+RLFP +S+H S Y Q Q P G
Sbjct: 113 LLPSFYRKLEKENGRKIGSCLIGAMPGSFYSRLFPEESMHFLHSCYCLQWLSQVPSGLVT 172
Query: 603 XRKPCLNXXXIY 638
N IY
Sbjct: 173 ELGISTNKGSIY 184
>sptr|Q9AVK0|Q9AVK0 Theobromine synthase (N-methyltransferase)
(Tentative caffeine synthase 1).
Length = 372
Score = 99.0 bits (245), Expect = 5e-20
Identities = 60/192 (31%), Positives = 93/192 (48%), Gaps = 3/192 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ M GEG+ SY+KN Q+ + +P++E V+++ A LP + + VADL
Sbjct: 1 MELQEVLRMNGGEGDTSYAKNSAYNQLVLAKVKPVLEQCVRELLRANLPNINKCIKVADL 60
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E+ +Q LNDL NDFN +F+
Sbjct: 61 GCASGPNTLLTVRDIVQSIDKVGQEK--------KNELERPTIQIFLNDLFPNDFNSVFK 112
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
+ +R+ + +P S+Y+RLFP +S+H S Y Q Q P G
Sbjct: 113 LLPSFYRKLEKENGRKIGSCLIGAMPGSFYSRLFPEESMHFLHSCYCLQWLSQVPSGLVT 172
Query: 603 XRKPCLNXXXIY 638
N IY
Sbjct: 173 ELGISTNKGSIY 184
>sptr|Q84PP7|Q84PP7 7-methylxanthine N-methyltransferase.
Length = 384
Score = 98.2 bits (243), Expect = 8e-20
Identities = 57/177 (32%), Positives = 89/177 (50%), Gaps = 3/177 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ HM EGEG+ SY+KN +A+ +P +E ++++ A LP + + VADL
Sbjct: 1 MELQEVLHMNEGEGDTSYAKNAS-YNLALAKVKPFLEQCIRELLRANLPNINKCIKVADL 59
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E+ +Q LNDL NDFN +F+
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEE--------KNELERPTIQIFLNDLFQNDFNSVFK 111
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEG 593
+ +R+ + +P S+Y RLFP +S+H S Y Q P G
Sbjct: 112 LLPSFYRKLEKENGRKIGSCLISAMPGSFYGRLFPEESMHFLHSCYSVHWLSQVPSG 168
>sptr|Q8H0G5|Q8H0G5 Theobromine synthase 1.
Length = 378
Score = 98.2 bits (243), Expect = 8e-20
Identities = 57/177 (32%), Positives = 89/177 (50%), Gaps = 3/177 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ HM EGEG+ SY+KN +A+ +P +E ++++ A LP + + VADL
Sbjct: 1 MELQEVLHMNEGEGDTSYAKNAS-YNLALAKVKPFLEQCIRELLRANLPNINKCIKVADL 59
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E+ +Q LNDL NDFN +F+
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEE--------KNELERPTIQIFLNDLFQNDFNSVFK 111
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEG 593
+ +R+ + +P S+Y RLFP +S+H S Y Q P G
Sbjct: 112 LLPSFYRKLEKENGRKIGSCLISAMPGSFYGRLFPEESMHFLHSCYSVHWLSQVPSG 168
>sptr|Q9AVJ9|Q9AVJ9 7-methylxanthine N-methyltransferase.
Length = 378
Score = 98.2 bits (243), Expect = 8e-20
Identities = 57/177 (32%), Positives = 89/177 (50%), Gaps = 3/177 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ HM EGEG+ SY+KN +A+ +P +E ++++ A LP + + VADL
Sbjct: 1 MELQEVLHMNEGEGDTSYAKNAS-YNLALAKVKPFLEQCIRELLRANLPNINKCIKVADL 59
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E+ +Q LNDL NDFN +F+
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEE--------KNELERPTIQIFLNDLFQNDFNSVFK 111
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEG 593
+ +R+ + +P S+Y RLFP +S+H S Y Q P G
Sbjct: 112 LLPSFYRKLEKENGRKIGSCLISAMPGSFYGRLFPEESMHFLHSCYSVHWLSQVPSG 168
>sptr|Q8H0F8|Q8H0F8 Tentative caffeine synthase 4.
Length = 385
Score = 97.8 bits (242), Expect = 1e-19
Identities = 61/192 (31%), Positives = 90/192 (46%), Gaps = 3/192 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ HM GEGE SY+KN Q+ + +P++E V+++ A LP + + VADL
Sbjct: 1 MELQEVLHMNGGEGEASYAKNSSFNQLVLAKVKPVLEQCVRELLRANLPNINKCIKVADL 60
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + + SI E +Q L DL NDFN +F
Sbjct: 61 GCASGPNTLLTVWDTVQSIDKVRQEM--------KNELERPTIQVFLTDLFQNDFNSVFM 112
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
+ +R+ + +P S++ RLFP +S+H SSY Q Q P G
Sbjct: 113 LLPSFYRKLEKENGRKIGSCLIAAMPGSFHGRLFPEESMHFLHSSYSLQFLSQVPSGLVT 172
Query: 603 XRKPCLNXXXIY 638
N IY
Sbjct: 173 ELGITANKRSIY 184
>sptr|Q9AVL9|Q9AVL9 Caffeine synthase.
Length = 385
Score = 97.8 bits (242), Expect = 1e-19
Identities = 61/192 (31%), Positives = 90/192 (46%), Gaps = 3/192 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ HM GEGE SY+KN Q+ + +P++E V+++ A LP + + VADL
Sbjct: 1 MELQEVLHMNGGEGEASYAKNSSFNQLVLAKVKPVLEQCVRELLRANLPNINKCIKVADL 60
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + + SI E +Q L DL NDFN +F
Sbjct: 61 GCASGPNTLLTVWDTVQSIDKVKQEM--------KNELERPTIQVFLTDLFQNDFNSVFM 112
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
+ +R+ + +P S++ RLFP +S+H SSY Q Q P G
Sbjct: 113 LLPSFYRKLEKENGRKIGSCLIAAMPGSFHGRLFPEESMHFLHSSYSLQFLSQVPSGLVT 172
Query: 603 XRKPCLNXXXIY 638
N IY
Sbjct: 173 ELGITANKRSIY 184
>sptr|Q9AR07|Q9AR07 S-adenosyl-L-methionine:jasmonic acid carboxyl
methyltransferase.
Length = 389
Score = 97.8 bits (242), Expect = 1e-19
Identities = 61/179 (34%), Positives = 94/179 (52%), Gaps = 7/179 (3%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRTMV-VADLGC 248
M++ R HM +G GE SY+KN Q + R +++ A+K++ + + + +ADLGC
Sbjct: 1 MEVMRVLHMNKGNGETSYAKNSTAQSNIISLGRRVMDEALKKLMMSNSEISSIGIADLGC 60
Query: 249 SAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRSV 428
S+GPN+LL IS+++ +I + +P EL+ LNDLP NDFN++ S+
Sbjct: 61 SSGPNSLLSISNIVDTIHNLCPDLDRPVP----------ELRVSLNDLPSNDFNYICASL 110
Query: 429 EEEFRR------AAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEP 587
E + R G R +V +P S+Y RLFPR+S+H SS Q P
Sbjct: 111 PEFYDRVNNNKEGLGFGRGGGESCFVSAVPGSFYGRLFPRRSLHFVHSSSSLHWLSQVP 169
>sptr|Q8RVM0|Q8RVM0 N-methyltransferase.
Length = 371
Score = 97.8 bits (242), Expect = 1e-19
Identities = 57/177 (32%), Positives = 89/177 (50%), Gaps = 3/177 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ HM EGEG+ SY+KN +A+ +P +E ++++ A LP + + VADL
Sbjct: 1 MELQEVLHMNEGEGDTSYAKNAS-YNLALAKVKPFLEQCIRELLRANLPNINKYIKVADL 59
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E+ +Q LNDL NDFN +F+
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEE--------KNELERPTIQIFLNDLFQNDFNSVFK 111
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEG 593
+ +R+ + +P S+Y RLFP +S+H S Y Q P G
Sbjct: 112 LLPSFYRKLEKENGRKIGSCLISAMPGSFYGRLFPEESMHFLHSCYSVHWLSQVPSG 168
>sptr|Q9FWR8|Q9FWR8 F14P1.3 protein.
Length = 323
Score = 97.8 bits (242), Expect = 1e-19
Identities = 62/187 (33%), Positives = 97/187 (51%), Gaps = 7/187 (3%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRTMV-VADLGC 248
M++ R HM +G GE SY+KN Q + R +++ A+K++ + + + +ADLGC
Sbjct: 1 MEVMRVLHMNKGNGETSYAKNSTAQSNIISLGRRVMDEALKKLMMSNSEISSIGIADLGC 60
Query: 249 SAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRSV 428
S+GPN+LL IS+++ +I + +P EL+ LNDLP NDFN++ S+
Sbjct: 61 SSGPNSLLSISNIVDTIHNLCPDLDRPVP----------ELRVSLNDLPSNDFNYICASL 110
Query: 429 EEEFRR------AAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPE 590
E + R G R +V +P S+Y RLFPR+S+H SS Q+
Sbjct: 111 PEFYDRVNNNKEGLGFGRGGGESCFVSAVPGSFYGRLFPRRSLHFVHSSSSLHWLSQKIT 170
Query: 591 GXXAXRK 611
G R+
Sbjct: 171 GSHNRRE 177
>sptr|Q8H0F9|Q8H0F9 Tentative caffeine synthase 3.
Length = 385
Score = 97.1 bits (240), Expect = 2e-19
Identities = 60/192 (31%), Positives = 90/192 (46%), Gaps = 3/192 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ HM GEG+ SY+KN Q+ + +P++E V ++ A LP + + VADL
Sbjct: 1 MELQEVLHMNGGEGDASYAKNSSFNQLVLAKVKPVLEQCVGELLRANLPNINKCIKVADL 60
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E +Q L DL NDFN +F
Sbjct: 61 GCASGPNTLLTVRDIVQSIDKVRQEM--------KNELERPTIQVFLTDLFQNDFNSVFM 112
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
+ +R+ + +P S++ RLFP +S+H SSY Q Q P G
Sbjct: 113 LLPSFYRKLEKENGRKIGSCLIAAMPGSFHGRLFPEESMHFLHSSYSLQFLSQVPSGLVT 172
Query: 603 XRKPCLNXXXIY 638
N IY
Sbjct: 173 ELGITANKRSIY 184
>sptr|Q9AVK1|Q9AVK1 Theobromine synthase.
Length = 385
Score = 97.1 bits (240), Expect = 2e-19
Identities = 60/192 (31%), Positives = 90/192 (46%), Gaps = 3/192 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ HM GEG+ SY+KN Q+ + +P++E V ++ A LP + + VADL
Sbjct: 1 MELQEVLHMNGGEGDASYAKNSSFNQLVLAKVKPVLEQCVGELLRANLPNINKCIKVADL 60
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E +Q L DL NDFN +F
Sbjct: 61 GCASGPNTLLTVRDIVQSIDKVRQEM--------KNELERPTIQVFLTDLFQNDFNSVFM 112
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
+ +R+ + +P S++ RLFP +S+H SSY Q Q P G
Sbjct: 113 LLPSFYRKLEKENGRKIGSCLIAAMPGSFHGRLFPEESMHFLHSSYSLQFLSQVPSGLVT 172
Query: 603 XRKPCLNXXXIY 638
N IY
Sbjct: 173 ELGITANKRSIY 184
>sptr|O23234|O23234 Hypothetical 42.0 kDa protein.
Length = 371
Score = 95.9 bits (237), Expect = 4e-19
Identities = 54/197 (27%), Positives = 95/197 (48%), Gaps = 3/197 (1%)
Frame = +3
Query: 75 DIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRTMVVADLGCSA 254
D+ R+F+M G+G+ SY++N Q+ A + + ++Q+Y P+++ +ADLGCS+
Sbjct: 5 DMEREFYMTGGDGKTSYARNSSLQKKASDTAKHITLETLQQLYKETRPKSLGIADLGCSS 64
Query: 255 GPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRSVEE 434
GPNTL I+ + ++ + E LNDLPGNDFN +F+S+ +
Sbjct: 65 GPNTLSTITDFIKTVQVAHHREIPIQPL--------PEFSIFLNDLPGNDFNFIFKSLPD 116
Query: 435 ---EFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXAX 605
E +R P ++ P S+Y RLFP ++H +S+ + P
Sbjct: 117 FHIELKR--DNNNGDCPSVFIAAYPGSFYGRLFPENTIHFVYASHSLHWLSKVPTALYDE 174
Query: 606 RKPCLNXXXIYIAXXTT 656
+ +N + I ++
Sbjct: 175 QGKSINKGCVSICSLSS 191
>sptr|Q8RVM1|Q8RVM1 N-methyltransferase.
Length = 378
Score = 95.5 bits (236), Expect = 5e-19
Identities = 59/192 (30%), Positives = 91/192 (47%), Gaps = 3/192 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRT---MVVADL 242
M+++ HM GEG+ SY+KN +A+ +P++E ++++ A LP + VADL
Sbjct: 1 MELQEVLHMNGGEGDTSYAKNSS-YNLALAKVKPVLEQCIRELLRANLPNINNCIKVADL 59
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E+ +Q LNDL NDFN +F+
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEE--------KNELERPTIQIFLNDLFQNDFNSVFK 111
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
+ +R+ + +P S+Y RLFP +S+H S Y Q P G
Sbjct: 112 LLPSFYRKLEKENGRKIGSCLISAMPGSFYGRLFPEESMHFIHSCYSFHWLSQVPSGLVI 171
Query: 603 XRKPCLNXXXIY 638
N IY
Sbjct: 172 ELGISANKGSIY 183
>sptr|Q8H0G0|Q8H0G0 Theobromine synthse 2.
Length = 384
Score = 95.1 bits (235), Expect = 7e-19
Identities = 59/192 (30%), Positives = 91/192 (47%), Gaps = 3/192 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRT---MVVADL 242
M+++ HM GEG+ SY+KN +A+ +P++E ++++ A LP + VADL
Sbjct: 1 MELQAVLHMNGGEGDTSYAKNSS-YNLALAKVKPVLEQCIRELLRANLPNINNCIKVADL 59
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E+ +Q LNDL NDFN +F+
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEE--------KNELERPTIQIFLNDLFQNDFNSVFK 111
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
+ +R+ + +P S+Y RLFP +S+H S Y Q P G
Sbjct: 112 LLPSFYRKLEKENGRKIGSCLISAMPGSFYGRLFPEESMHFIHSCYSFHWLSQVPSGLVI 171
Query: 603 XRKPCLNXXXIY 638
N IY
Sbjct: 172 ELGISANKGSIY 183
>sptr|Q8H0D2|Q8H0D2 Tentative caffeine synthase 7.
Length = 384
Score = 92.8 bits (229), Expect = 3e-18
Identities = 59/192 (30%), Positives = 92/192 (47%), Gaps = 3/192 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ HM GEG+ SY+KN + + +P++E ++++ A LP + + VADL
Sbjct: 1 MELQEVLHMNGGEGDTSYAKNSF-YNLFLIRVKPILEQCIQELLRANLPNINKCIKVADL 59
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E+ +Q LNDL NDFN +F+
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEK--------KNELERPTIQIFLNDLFQNDFNSVFK 111
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
S+ +R+ + +P S+Y RLFP +S+H S Y Q P G
Sbjct: 112 SLPSFYRKLEKENGCKIGSCLIGAMPGSFYGRLFPEESMHFLHSCYCLHWLSQVPSGLVT 171
Query: 603 XRKPCLNXXXIY 638
N IY
Sbjct: 172 ELGISANKGCIY 183
>sptr|Q84PP8|Q84PP8 3,7-dimethylxanthine N-methyltransferase.
Length = 384
Score = 92.4 bits (228), Expect = 4e-18
Identities = 59/192 (30%), Positives = 92/192 (47%), Gaps = 3/192 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLP---RTMVVADL 242
M+++ HM GEG+ SY+KN + + +P++E ++++ A LP + + VADL
Sbjct: 1 MELQEVLHMNGGEGDTSYAKNSF-YNLFLIRVKPILEQCIQELLRANLPNINKCIKVADL 59
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E+ +Q LNDL NDFN +F+
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEK--------KNELERPTIQIFLNDLFQNDFNSVFK 111
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
S+ +R+ + +P S+Y RLFP +S+H S Y Q P G
Sbjct: 112 SLPSFYRKLEKENGRKIGSCLIGAMPGSFYGRLFPEESMHFLHSCYCLHWLSQVPSGLVT 171
Query: 603 XRKPCLNXXXIY 638
N IY
Sbjct: 172 ELGISANKGCIY 183
>sptr|Q8RVL8|Q8RVL8 N-methyltransferase.
Length = 386
Score = 92.0 bits (227), Expect = 6e-18
Identities = 58/192 (30%), Positives = 90/192 (46%), Gaps = 3/192 (1%)
Frame = +3
Query: 72 MDIRRDFHMAEGEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALLPRT---MVVADL 242
M+++ M GEG+ SY+KN +A+ +P++E ++++ A LP + VADL
Sbjct: 1 MELQEVLRMNGGEGDTSYAKNSS-YNLALAKVKPVLEQCIRELLRANLPNINNCIKVADL 59
Query: 243 GCSAGPNTLLFISSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFR 422
GC++GPNTLL + ++ SI E+ +Q LNDL NDFN +F+
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKLGLEE--------KNELERPTIQIFLNDLFQNDFNSVFK 111
Query: 423 SVEEEFRRAAGCERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEGXXA 602
+ +R+ + +P S+Y RLFP +S+H S Y Q P G
Sbjct: 112 LLPSFYRKLEKENGRKIGSCLISAMPGSFYGRLFPEESMHFLHSCYCLHWLSQVPSGLVT 171
Query: 603 XRKPCLNXXXIY 638
N IY
Sbjct: 172 ELGISANKGSIY 183
>sptr|Q9FYZ9|Q9FYZ9 SAM:benzoic acid carboxyl methyltransferase.
Length = 364
Score = 89.7 bits (221), Expect = 3e-17
Identities = 54/165 (32%), Positives = 84/165 (50%), Gaps = 2/165 (1%)
Frame = +3
Query: 105 GEGEWSYSKNCRRQQVAVXXTRPMVETAVKQVYAALL--PRTMVVADLGCSAGPNTLLFI 278
G+GE SY+ N Q+V + + +++ +K + + P+ + D+GCS+GPN LL +
Sbjct: 14 GDGETSYANNSGLQKVMMSKSLHVLDETLKDIIGDHVGFPKCFKMMDMGCSSGPNALLVM 73
Query: 279 SSVLSSIXXXXXEQCKPPSXXXXXXXHHVELQFVLNDLPGNDFNHLFRSVEEEFRRAAGC 458
S ++++I E+ E + LNDLP NDFN+LF+ + E C
Sbjct: 74 SGIINTIEDLYTEK---------NINELPEFEVFLNDLPDNDFNNLFKLLSHE---NGNC 121
Query: 459 ERAPHPPYYVMGLPESYYNRLFPRQSVHLFXSSYXXQXRXQEPEG 593
+V GLP S+Y RL P++S+H SSY Q PEG
Sbjct: 122 --------FVYGLPGSFYGRLLPKKSLHFAYSSYSIHWLSQVPEG 158
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 428,505,786
Number of Sequences: 1272877
Number of extensions: 8236710
Number of successful extensions: 20711
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 19918
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 20536
length of database: 406,886,216
effective HSP length: 118
effective length of database: 256,686,730
effective search space used: 25668673000
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

