BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
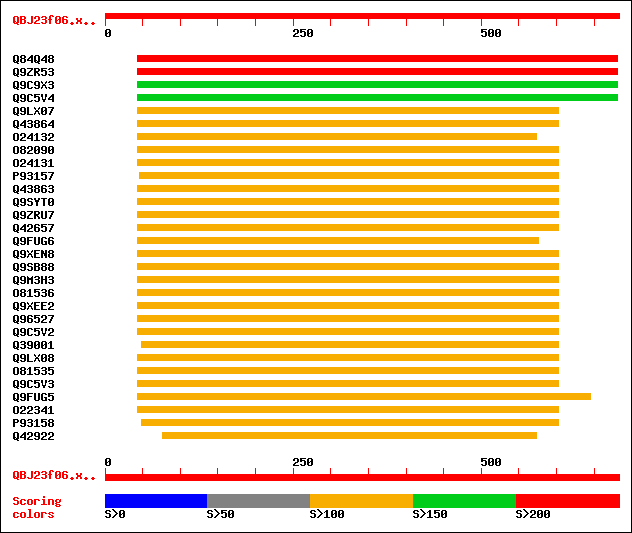
Query= QBJ23f06.xg.2.1
(683 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q84Q48|Q84Q48 P0456B03.13 protein. 261 6e-69
sptr|Q9ZR53|Q9ZR53 Annexin-like protein. 238 7e-62
sptr|Q9C9X3|Q9C9X3 Putative annexin. 177 1e-43
sptr|Q9C5V4|Q9C5V4 Calcium-binding protein annexin 5. 176 3e-43
sptr|Q9LX07|Q9LX07 Annexin-like protein. 135 4e-31
sptr|Q43864|Q43864 Annexin p35. 135 4e-31
sptr|O24132|O24132 Annexin. 135 5e-31
sptr|O82090|O82090 Fiber annexin. 134 1e-30
sptr|O24131|O24131 Annexin. 134 1e-30
sptr|P93157|P93157 Annexin (Fragment). 134 1e-30
sptr|Q43863|Q43863 Annexin p33. 132 5e-30
sptr|Q9SYT0|Q9SYT0 Ca2+-dependent membrane-binding protein annex... 132 5e-30
sptr|Q9ZRU7|Q9ZRU7 Annexin P38. 131 7e-30
sptr|Q42657|Q42657 Annexin. 131 7e-30
sptr|Q9FUG6|Q9FUG6 Annexin. 131 7e-30
sptr|Q9XEN8|Q9XEN8 Vacuole-associated annexin VCaB42. 131 9e-30
sptr|Q9SB88|Q9SB88 Annexin cap32. 131 9e-30
sptr|Q9M3H3|Q9M3H3 Annexin p34. 131 9e-30
sptr|O81536|O81536 Annexin p34. 130 1e-29
sptr|Q9XEE2|Q9XEE2 Putative annexin. 130 2e-29
sptr|Q96527|Q96527 Annexin-like protein. 130 2e-29
sptr|Q9C5V2|Q9C5V2 Calcium-binding protein annexin 7. 130 2e-29
sptr|Q39001|Q39001 Annexin (Fragment). 128 6e-29
sptr|Q9LX08|Q9LX08 Annexin-like protein. 128 8e-29
sptr|O81535|O81535 Annexin p35. 127 1e-28
sptr|Q9C5V3|Q9C5V3 Calcium-binding protein annexin 6. 127 1e-28
sptr|Q9FUG5|Q9FUG5 Annexin. 126 2e-28
sptr|O22341|O22341 Annexin. 126 3e-28
sptr|P93158|P93158 Annexin (Fragment). 124 1e-27
sptr|Q42922|Q42922 Annexin (Fragment). 123 2e-27
>sptr|Q84Q48|Q84Q48 P0456B03.13 protein.
Length = 321
Score = 261 bits (667), Expect = 6e-69
Identities = 125/218 (57%), Positives = 163/218 (74%), Gaps = 5/218 (2%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASL++PP +PR+DAIDL +AFKGFGCD+T V IL HRD+ QR LI++ Y A+YH++
Sbjct: 1 MASLSVPPVPTDPRRDAIDLHRAFKGFGCDATAVTAILAHRDASQRALIRRHYAAVYHQD 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L HR+++EL+G+HK+A+LLW+LDPA RD VL +AL+GD D+RAAT+++CSRTPSQL +
Sbjct: 61 LLHRLAAELSGHHKRAVLLWVLDPASRDAAVLHQALNGDVTDMRAATEVVCSRTPSQLLV 120
Query: 403 MKQTYYARF----GTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPE-VDPTIXTHDAKDL 567
++Q Y ARF G LEHD+ SGDHQ LLLAY+ PRYEGPE VD DA++L
Sbjct: 121 VRQAYLARFGGGGGGGLEHDVAVRASGDHQRLLLAYLRSPRYEGPEVVDMAAAARDAREL 180
Query: 568 YKAGEXRLGTDEXXXXXXFTXRXWAHXASXSSAXHHMY 681
Y+AGE RLGTDE F+ R AH A+ ++A HHMY
Sbjct: 181 YRAGERRLGTDERTFIRVFSERSAAHMAAVAAAYHHMY 218
Score = 45.4 bits (106), Expect = 7e-04
Identities = 25/69 (36%), Positives = 37/69 (53%)
Frame = +1
Query: 100 LQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNHKKAMLL 279
L +A KG G + TT+I ++T R V I+ EY Y L+ + SE +GN++ +L
Sbjct: 256 LHEAMKGLGTNDTTLIRVVTTRAEVDMQYIKAEYHRSYKRSLADAVHSETSGNYRTFLLS 315
Query: 280 WILDPAGRD 306
I GRD
Sbjct: 316 LI----GRD 320
>sptr|Q9ZR53|Q9ZR53 Annexin-like protein.
Length = 333
Score = 238 bits (606), Expect = 7e-62
Identities = 120/213 (56%), Positives = 151/213 (70%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MA+L LPP +PR DA+ L +AFKGFGCD+T VINIL HRD+ QR +QQEY+A Y EE
Sbjct: 1 MATLILPPIPPSPRDDAMQLYRAFKGFGCDTTAVINILAHRDATQRAYLQQEYKATYSEE 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
LS R+ SEL G + A+LLW+ DPA RD ++R++L D L AAT++ICSRTPSQL
Sbjct: 61 LSKRLVSELKGKLETAVLLWLPDPAARDAEIIRKSLVVDR-SLEAATEVICSRTPSQLQY 119
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+KQ Y+++FG YLEH+I +TSGDHQ +LL Y+ PR+EG EV+ I DAK LYKAGE
Sbjct: 120 LKQLYHSKFGVYLEHEIELNTSGDHQKILLRYLTTPRHEGLEVNREIAQKDAKVLYKAGE 179
Query: 583 XRLGTDEXXXXXXFTXRXWAHXASXSSAXHHMY 681
+LGTDE F+ R AH A+ SS H MY
Sbjct: 180 KKLGTDEKTFVQIFSERSSAHLAAVSSYYHDMY 212
>sptr|Q9C9X3|Q9C9X3 Putative annexin.
Length = 316
Score = 177 bits (449), Expect = 1e-43
Identities = 90/213 (42%), Positives = 130/213 (61%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MA++ +P + +PR DA L KAFKG GCD++ +INIL HR++ QR LI+QEY + ++
Sbjct: 1 MATMKIPMTVPSPRVDADQLFKAFKGTGCDTSVIINILAHRNATQRALIEQEYETKFSDD 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L R+ SEL+G+ KKA+LLW+ + RD ++L+ +L G D +A +IIC+R+ SQL
Sbjct: 61 LRKRLHSELHGHLKKAVLLWMPEAVERDASILKRSLRGAVTDHKAIAEIICTRSGSQLRQ 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+KQ Y FG LE DI SG+H+ +LLAY+ RYEGPE+D +DA+ L A
Sbjct: 121 IKQVYSNTFGVKLEEDIESEASGNHKRVLLAYLNTTRYEGPEIDNASVENDARTLKSAVA 180
Query: 583 XRLGTDEXXXXXXFTXRXWAHXASXSSAXHHMY 681
+ +D+ FT R H + S MY
Sbjct: 181 RKHKSDDQTLIQIFTDRSRTHLVAVRSTYRSMY 213
Score = 46.6 bits (109), Expect = 3e-04
Identities = 40/149 (26%), Positives = 65/149 (43%), Gaps = 5/149 (3%)
Frame = +1
Query: 73 ANPRQDAIDLQKAF-KGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELSHRISSEL 249
A+ DA L+ A + D T+I I T R ++ YR+MY +EL I E
Sbjct: 166 ASVENDARTLKSAVARKHKSDDQTLIQIFTDRSRTHLVAVRSTYRSMYGKELGKAIRDET 225
Query: 250 NGNHKKAMLLWILDPAGRD----XTVLREALSGDTMDLRAATDIICSRTPSQLXIMKQTY 417
GN + +LL IL A LR+++ G D A I+ +R + + Y
Sbjct: 226 RGNFEH-VLLTILQCAENSCFYFAKALRKSMKGLGTDDTALIRIVVTRAEVDMQFIITEY 284
Query: 418 YARFGTYLEHDIGHHTSGDHQXLLLAYVG 504
R+ L + + T+ ++ LL+ +G
Sbjct: 285 RKRYKKTLYNAVHSDTTSHYRTFLLSLLG 313
Score = 40.8 bits (94), Expect = 0.016
Identities = 23/65 (35%), Positives = 34/65 (52%)
Frame = +1
Query: 100 LQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNHKKAMLL 279
L+K+ KG G D T +I I+ R V I EYR Y + L + + S+ +H + LL
Sbjct: 251 LRKSMKGLGTDDTALIRIVVTRAEVDMQFIITEYRKRYKKTLYNAVHSDTT-SHYRTFLL 309
Query: 280 WILDP 294
+L P
Sbjct: 310 SLLGP 314
>sptr|Q9C5V4|Q9C5V4 Calcium-binding protein annexin 5.
Length = 316
Score = 176 bits (445), Expect = 3e-43
Identities = 89/213 (41%), Positives = 130/213 (61%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MA++ +P + +PR DA L KAFKG GCD++ +INIL HR++ QR LI+QEY + ++
Sbjct: 1 MATMKIPMTVPSPRVDADQLFKAFKGRGCDTSVIINILAHRNATQRALIEQEYETKFSDD 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L R+ SEL+G+ KKA+LLW+ + RD ++L+ +L G D +A +I+C+R+ SQL
Sbjct: 61 LRKRLHSELHGHLKKAVLLWMPEAVERDASILKRSLRGAVTDHKAIAEIMCTRSGSQLRQ 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+KQ Y FG LE DI SG+H+ +LLAY+ RYEGPE+D +DA+ L A
Sbjct: 121 IKQVYSNTFGVKLEEDIESEASGNHKRVLLAYLNTTRYEGPEIDNASVENDARTLKSAVA 180
Query: 583 XRLGTDEXXXXXXFTXRXWAHXASXSSAXHHMY 681
+ +D+ FT R H + S MY
Sbjct: 181 RKHKSDDQTLIQIFTDRSRTHLVAVRSTYRSMY 213
Score = 46.6 bits (109), Expect = 3e-04
Identities = 40/149 (26%), Positives = 65/149 (43%), Gaps = 5/149 (3%)
Frame = +1
Query: 73 ANPRQDAIDLQKAF-KGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELSHRISSEL 249
A+ DA L+ A + D T+I I T R ++ YR+MY +EL I E
Sbjct: 166 ASVENDARTLKSAVARKHKSDDQTLIQIFTDRSRTHLVAVRSTYRSMYGKELGKAIRDET 225
Query: 250 NGNHKKAMLLWILDPAGRD----XTVLREALSGDTMDLRAATDIICSRTPSQLXIMKQTY 417
GN + +LL IL A LR+++ G D A I+ +R + + Y
Sbjct: 226 RGNFEH-VLLTILQCAENSCFYFAKALRKSMKGLGTDDTALIRIVVTRAEVDMQFIITEY 284
Query: 418 YARFGTYLEHDIGHHTSGDHQXLLLAYVG 504
R+ L + + T+ ++ LL+ +G
Sbjct: 285 RKRYKKTLYNAVHSDTTSHYRTFLLSLLG 313
Score = 40.8 bits (94), Expect = 0.016
Identities = 23/65 (35%), Positives = 34/65 (52%)
Frame = +1
Query: 100 LQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNHKKAMLL 279
L+K+ KG G D T +I I+ R V I EYR Y + L + + S+ +H + LL
Sbjct: 251 LRKSMKGLGTDDTALIRIVVTRAEVDMQFIITEYRKRYKKTLYNAVHSDTT-SHYRTFLL 309
Query: 280 WILDP 294
+L P
Sbjct: 310 SLLGP 314
>sptr|Q9LX07|Q9LX07 Annexin-like protein.
Length = 316
Score = 135 bits (341), Expect = 4e-31
Identities = 70/187 (37%), Positives = 113/187 (60%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASL +P + P +DA L KAFKG+G + +I+IL HR++ QR I+ Y A Y+++
Sbjct: 1 MASLKVPATVPLPEEDAEQLYKAFKGWGTNERMIISILAHRNATQRSFIRAVYAANYNKD 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL+G+ ++A++LW +PA RD + +E+ T + +I C+R+ +L
Sbjct: 61 LLKELDRELSGDFERAVMLWTFEPAERDAYLAKESTKMFTKNNWVLVEIACTRSALELFN 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
KQ Y AR+ T LE D+ +HTSGD + LL+ V RY+G EV+ T+ +AK L++ +
Sbjct: 121 AKQAYQARYKTSLEEDVAYHTSGDIRKLLVPLVSTFRYDGDEVNMTLARSEAKILHEKIK 180
Query: 583 XRLGTDE 603
+ D+
Sbjct: 181 EKAYADD 187
Score = 42.7 bits (99), Expect = 0.004
Identities = 35/148 (23%), Positives = 61/148 (41%), Gaps = 3/148 (2%)
Frame = +1
Query: 70 MANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELSHRISSEL 249
M R +A L + K +I ILT R Q Y+ + +S + +
Sbjct: 165 MTLARSEAKILHEKIKEKAYADDDLIRILTTRSKAQISATLNHYKNNFGTSMSKYLKEDS 224
Query: 250 NGNH---KKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYY 420
+ KA++ + P VLR+A++ D T ++ +R + +K+ Y
Sbjct: 225 ENEYIQLLKAVIKCLTYPEKYFEKVLRQAINKLGTDEWGLTRVVTTRAEFDMERIKEEYI 284
Query: 421 ARFGTYLEHDIGHHTSGDHQXLLLAYVG 504
R L+ I T GD++ +LLA +G
Sbjct: 285 RRNSVPLDRAIAKDTHGDYEDILLALLG 312
>sptr|Q43864|Q43864 Annexin p35.
Length = 314
Score = 135 bits (341), Expect = 4e-31
Identities = 69/187 (36%), Positives = 110/187 (58%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MA+LT+P ++ +D L KAF+G+G + +I+IL HR++ Q I++ Y Y +E
Sbjct: 1 MATLTVPSSVPAVAEDCEQLHKAFEGWGTNEKLIISILAHRNAAQARAIRRGYAEAYGKE 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + E++G ++A++LW LDPA RD + E RA +I C+RTP+QL
Sbjct: 61 LLRALGDEIHGKFERAVILWTLDPAERDAVLANEEAKKSHPGGRALVEIACARTPAQLFA 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+KQ Y+ RF LE D+ H +GD + LL+ V RY+GPEV+ ++ +AK L++
Sbjct: 121 VKQAYHDRFKRSLEEDVAAHVTGDFRKLLVPLVSAYRYDGPEVNTSLAHSEAKILHEKIH 180
Query: 583 XRLGTDE 603
+ +DE
Sbjct: 181 KKAYSDE 187
>sptr|O24132|O24132 Annexin.
Length = 314
Score = 135 bits (340), Expect = 5e-31
Identities = 67/177 (37%), Positives = 108/177 (61%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASLT+P + + +D L+ AFKG+G + +I+IL HR++ QR LIQQ Y + E+
Sbjct: 1 MASLTVPAEVPSVAEDCEQLRSAFKGWGTNEKLIISILAHRNAAQRKLIQQTYAETFGED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL + +K +++W LDP+ RD + +EA T +I C+R+P +L +
Sbjct: 61 LLKELDRELTNDFEKLVVVWTLDPSERDAYLAKEATKRWTKSNFVLVEIACTRSPKELVL 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYK 573
++ Y+AR+ LE D+ +HT+G+H+ LL+A V RY G EVD + +AK L++
Sbjct: 121 AREAYHARYKKSLEEDVAYHTTGEHRKLLVALVSSYRYGGDEVDLRLAKAEAKILHE 177
Score = 35.0 bits (79), Expect = 0.88
Identities = 30/123 (24%), Positives = 52/123 (42%), Gaps = 2/123 (1%)
Frame = +1
Query: 142 VINILTHRDSVQRGLIQQEYRAMYHEELSHRISS--ELNGNHKKAMLLWILDPAGRDXTV 315
VI IL R Q Y+ Y E++ ++ E G +A + ++ V
Sbjct: 189 VIRILATRSKAQINATLNHYKDEYEEDILKQLEEGDEFVGL-LRATIKGLVYTEHYFVEV 247
Query: 316 LREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFGTYLEHDIGHHTSGDHQXLLLA 495
LR+A++ + T +I +R + + Y R +L I T GD++ +LLA
Sbjct: 248 LRDAINRRGTEEDHLTRVIATRAEVDMKTIADEYQKRDSIHLGRAIAKDTRGDYESMLLA 307
Query: 496 YVG 504
+G
Sbjct: 308 LLG 310
>sptr|O82090|O82090 Fiber annexin.
Length = 316
Score = 134 bits (337), Expect = 1e-30
Identities = 70/187 (37%), Positives = 111/187 (59%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MA+LT+P + + +D L+KAF G+G + +I+IL HR++ QR LI++ Y Y E+
Sbjct: 1 MATLTVPTTVPSVSEDCEQLRKAFSGWGTNEGLIIDILGHRNAEQRNLIRKTYAETYGED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL+ + ++ +LLW LDPA RD + EA T + +I C+R+ +QL
Sbjct: 61 LLKALDKELSNDFERLVLLWALDPAERDALLANEATKRWTSSNQVLMEIACTRSANQLLH 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+Q Y+AR+ LE D+ HHT+GD + LLL V RYEG EV+ + +AK L++
Sbjct: 121 ARQAYHARYKKSLEEDVAHHTTGDFRKLLLPLVSSYRYEGEEVNMNLAKTEAKLLHEKIS 180
Query: 583 XRLGTDE 603
+ +D+
Sbjct: 181 DKAYSDD 187
Score = 38.9 bits (89), Expect = 0.061
Identities = 30/124 (24%), Positives = 51/124 (41%), Gaps = 3/124 (2%)
Frame = +1
Query: 142 VINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNHK---KAMLLWILDPAGRDXT 312
VI +L R Q Y+ Y +++ + ++ ++ + ++ P
Sbjct: 189 VIRVLATRSKAQINATLNHYKNEYGNDINKDLKADPKDEFLALLRSTVKCLVYPEKYFEK 248
Query: 313 VLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFGTYLEHDIGHHTSGDHQXLLL 492
VLR A++ D A T ++C+R L I+ Y R L I T GD++ LLL
Sbjct: 249 VLRLAINRRGTDEGALTRVVCTRAEVDLKIIADEYQRRNSVPLTRAIVKDTHGDYEKLLL 308
Query: 493 AYVG 504
G
Sbjct: 309 VLAG 312
>sptr|O24131|O24131 Annexin.
Length = 314
Score = 134 bits (337), Expect = 1e-30
Identities = 68/187 (36%), Positives = 110/187 (58%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASLT+P + + +D L+ AFKG+G + +I+IL HR++ QR LIQQ Y + E+
Sbjct: 1 MASLTVPAEVPSVAEDCEQLRSAFKGWGTNEKLIISILAHRNAAQRKLIQQTYAETFGED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL + +K +++W LDP+ RD + +EA T +I C+R+P +L +
Sbjct: 61 LLKELDRELTNDFEKLVVVWTLDPSERDAYLAKEATKRWTKSNFVLVEIACTRSPKELVL 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
++ Y+ARF LE D+ +HT+G+H LL+ V RY G EVD + +AK L++
Sbjct: 121 AREAYHARFKKSLEEDVAYHTTGEHPQLLVPLVSSYRYGGDEVDLRLAKAEAKILHEKIS 180
Query: 583 XRLGTDE 603
+ +D+
Sbjct: 181 DKAYSDD 187
Score = 39.7 bits (91), Expect = 0.036
Identities = 33/123 (26%), Positives = 53/123 (43%), Gaps = 2/123 (1%)
Frame = +1
Query: 142 VINILTHRDSVQRGLIQQEYRAMYHEELSHRISS--ELNGNHKKAMLLWILDPAGRDXTV 315
VI IL R Q Y+ Y E++ ++ E G +A + ++ P V
Sbjct: 189 VIRILATRSKAQINATLNHYKDEYEEDILKQLEEGDEFVGL-LRATIKGLVYPEHYFVEV 247
Query: 316 LREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFGTYLEHDIGHHTSGDHQXLLLA 495
LR+A++ D T +I +R + I+ Y R L I T GD++ +LLA
Sbjct: 248 LRDAINRRGTDEDHLTRVIATRAEVDMKIIADEYQKRDSIPLGRAIAKDTRGDYESMLLA 307
Query: 496 YVG 504
+G
Sbjct: 308 LLG 310
>sptr|P93157|P93157 Annexin (Fragment).
Length = 315
Score = 134 bits (336), Expect = 1e-30
Identities = 70/186 (37%), Positives = 110/186 (59%)
Frame = +1
Query: 46 ASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEEL 225
A+LT+P + + +D L+KAF G+G + +I+IL HR++ QR LI++ Y Y E+L
Sbjct: 1 ATLTVPTTVPSVSEDCEQLRKAFSGWGTNEGLIIDILGHRNAEQRNLIRKTYAETYGEDL 60
Query: 226 SHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXIM 405
+ EL+ + ++ +LLW LDPA RD + EA T + +I C+R+ +QL
Sbjct: 61 LKALDKELSNDFERLVLLWALDPAERDALLANEATKRWTSSNQVLMEIACTRSANQLLHA 120
Query: 406 KQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGEX 585
+Q Y+AR+ LE D+ HHT+GD LLL V RYEG EV+ T+ +AK L++
Sbjct: 121 RQAYHARYKKSLEEDVAHHTTGDFHKLLLPLVSSYRYEGEEVNMTLAKTEAKLLHEKISN 180
Query: 586 RLGTDE 603
+ +D+
Sbjct: 181 KAYSDD 186
Score = 38.5 bits (88), Expect = 0.080
Identities = 29/124 (23%), Positives = 51/124 (41%), Gaps = 3/124 (2%)
Frame = +1
Query: 142 VINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNHK---KAMLLWILDPAGRDXT 312
VI +L R Q Y+ Y +++ + ++ ++ + ++ P
Sbjct: 188 VIRVLATRSKAQINATLNHYKNEYGNDINKDLKADPKDEFLALLRSTVKCLVYPEKYFEK 247
Query: 313 VLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFGTYLEHDIGHHTSGDHQXLLL 492
VLR A++ D A T ++C+R L ++ Y R L I T GD++ LLL
Sbjct: 248 VLRLAINRRGTDEGALTRVVCTRAEVDLKVIADEYQRRNSVPLTRAIVKDTHGDYEKLLL 307
Query: 493 AYVG 504
G
Sbjct: 308 VLAG 311
>sptr|Q43863|Q43863 Annexin p33.
Length = 314
Score = 132 bits (331), Expect = 5e-30
Identities = 69/187 (36%), Positives = 108/187 (57%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MA+L +P + D L+KAF+G+G + +I+IL HRD+ QR I++ Y Y EE
Sbjct: 1 MATLKVPATVPPVADDCDQLRKAFQGWGTNEALIISILGHRDAAQRRAIRRAYAEAYGEE 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L I+ E++G+ ++A++LW LDPA RD + EA R +I C+RT +Q+
Sbjct: 61 LLRSITDEISGDFERAVILWTLDPAERDAVLANEAARKWKPGNRVLVEIACTRTSAQIFA 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+Q Y+ RF LE DI H +GD + LL+ V RY+GPEV+ + +AK L++
Sbjct: 121 TRQAYHERFKRSLEEDIAAHVTGDFRKLLVPLVSTYRYDGPEVNTRLAHSEAKLLHEKIH 180
Query: 583 XRLGTDE 603
+ +D+
Sbjct: 181 HKAYSDD 187
Score = 47.8 bits (112), Expect = 1e-04
Identities = 37/128 (28%), Positives = 61/128 (47%), Gaps = 7/128 (5%)
Frame = +1
Query: 142 VINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNHK-------KAMLLWILDPAG 300
+I ILT R Q LI Y++ HRI+ +L + + +A++ P
Sbjct: 189 IIRILTTRSKPQ--LIATFNH--YNDAFGHRINKDLKADPQDEYLRTLRAIIRCFSCPDR 244
Query: 301 RDXTVLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFGTYLEHDIGHHTSGDHQ 480
V R+A++G D + T +I +R L ++K+ Y R LE + TSGD++
Sbjct: 245 YFEKVARQAIAGLGTDENSLTRVITTRAEVDLKLIKEAYQKRNSVPLERAVAGDTSGDYE 304
Query: 481 XLLLAYVG 504
+LLA +G
Sbjct: 305 SMLLALLG 312
>sptr|Q9SYT0|Q9SYT0 Ca2+-dependent membrane-binding protein annexin
(F14D7.2 protein) (Putative annexin protein).
Length = 317
Score = 132 bits (331), Expect = 5e-30
Identities = 68/187 (36%), Positives = 111/187 (59%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MA+L + ++ P DA L+ AF+G+G + +I+IL HR + QR +I+Q Y Y E+
Sbjct: 1 MATLKVSDSVPAPSDDAEQLRTAFEGWGTNEDLIISILAHRSAEQRKVIRQAYHETYGED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL+ + ++A+LLW L+P RD + EA T + ++ C+RT +QL
Sbjct: 61 LLKTLDKELSNDFERAILLWTLEPGERDALLANEATKRWTSSNQVLMEVACTRTSTQLLH 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+Q Y+AR+ LE D+ HHT+GD + LL++ V RYEG EV+ T+ +AK +++ +
Sbjct: 121 ARQAYHARYKKSLEEDVAHHTTGDFRKLLVSLVTSYRYEGDEVNMTLAKQEAKLVHEKIK 180
Query: 583 XRLGTDE 603
+ DE
Sbjct: 181 DKHYNDE 187
>sptr|Q9ZRU7|Q9ZRU7 Annexin P38.
Length = 316
Score = 131 bits (330), Expect = 7e-30
Identities = 69/187 (36%), Positives = 107/187 (57%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASL +P ++ +P +DA L+KAFKG+G + +I IL HR++ QR LI+ Y A Y E+
Sbjct: 1 MASLKVPASVPDPCEDAEQLKKAFKGWGTNEELIIQILAHRNAAQRKLIRDSYAAAYGED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + SEL + ++ +LLW L PA RD + EA T +I C+R+ +L
Sbjct: 61 LLKDLDSELTSDFQRIVLLWTLSPAERDAYLANEATKRLTASNWVIMEIACTRSSDELFK 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+Q Y+ R+ E D+ +HT+GD + LL+ + RYEG EV+ T+ +A L++
Sbjct: 121 ARQAYHTRYKKSFEEDVAYHTTGDFRKLLVPLITAFRYEGEEVNMTLARKEANILHEKVS 180
Query: 583 XRLGTDE 603
+ DE
Sbjct: 181 GKAYNDE 187
Score = 43.5 bits (101), Expect = 0.002
Identities = 35/148 (23%), Positives = 63/148 (42%), Gaps = 3/148 (2%)
Frame = +1
Query: 70 MANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELSHRISSEL 249
M R++A L + G + +I I++ R Q Y + E+ + ++
Sbjct: 165 MTLARKEANILHEKVSGKAYNDEELIRIISTRSKTQLNATFNHYNDQHGHEIIKDLEADD 224
Query: 250 NGNHKK---AMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYY 420
+ + K A + + P VLR A+ G D T ++ +R + +K+ Y
Sbjct: 225 DDEYLKLLRAAIECLKTPEKYFEKVLRVAIKGLGTDEWDLTRVVATRAEVDMERIKEEYN 284
Query: 421 ARFGTYLEHDIGHHTSGDHQXLLLAYVG 504
R L+ I TSGD++ +LLA +G
Sbjct: 285 KRNSVTLDRAITGDTSGDYERMLLALIG 312
>sptr|Q42657|Q42657 Annexin.
Length = 314
Score = 131 bits (330), Expect = 7e-30
Identities = 66/187 (35%), Positives = 110/187 (58%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASLT+P + + +D L+ AFKG+G + +I+IL HR + QR LI+Q Y + E+
Sbjct: 1 MASLTVPAHVPSAAEDCEQLRSAFKGWGTNEKLIISILAHRTAAQRKLIRQTYAETFGED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL + +K +L+W LDP+ RD + +EA T ++ C+R+P +L +
Sbjct: 61 LLKELDRELTHDFEKLVLVWTLDPSERDAHLAKEATKRWTKSNFVLVELACTRSPKELVL 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
++ Y+AR+ LE D+ +HT+GDH+ LL+ V RY G EVD + ++K L++
Sbjct: 121 AREAYHARYKKSLEEDVAYHTTGDHRKLLVPLVSSYRYGGEEVDLRLAKAESKILHEKIS 180
Query: 583 XRLGTDE 603
+ +D+
Sbjct: 181 DKAYSDD 187
Score = 36.2 bits (82), Expect = 0.40
Identities = 32/125 (25%), Positives = 54/125 (43%), Gaps = 4/125 (3%)
Frame = +1
Query: 142 VINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNHKKAMLL----WILDPAGRDX 309
VI IL R Q Y+ + E++ ++ +G+ A+L ++ P
Sbjct: 189 VIRILATRSKAQLNATLNHYKDEHGEDILKQLE---DGDEFVALLRATIKGLVYPEHYFV 245
Query: 310 TVLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFGTYLEHDIGHHTSGDHQXLL 489
VLR+A++ + T +I +R L I+ Y R L I T GD++ +L
Sbjct: 246 EVLRDAINRRGTEEDHLTRVIATRAEVDLKIIADEYQKRDSIPLGRAIAKDTRGDYESML 305
Query: 490 LAYVG 504
LA +G
Sbjct: 306 LALLG 310
>sptr|Q9FUG6|Q9FUG6 Annexin.
Length = 330
Score = 131 bits (330), Expect = 7e-30
Identities = 66/178 (37%), Positives = 102/178 (57%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
M+++T+P + + +D I L +AF+G GCD ++N++ HRD QR I+ Y Y E+
Sbjct: 1 MSTITVPNPVPDTNEDCIALHRAFEGIGCDKEALLNVICHRDQQQRQRIRHSYNIKYEED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L ++ SEL+GN +K +LW+ +PA RD T+L EAL G D RA T+++ RT ++L
Sbjct: 61 LLKKLKSELHGNLEKGAVLWMCNPAERDATILHEALGGLIKDYRALTEVLYLRTSAELLD 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKA 576
+++ Y + F LE +I G Q LLL + R E E+D D +DL A
Sbjct: 121 IRRAYSSSFDRSLEEEIATKIGGSEQKLLLGLLREERIEDDEIDTLEVEADTEDLLSA 178
>sptr|Q9XEN8|Q9XEN8 Vacuole-associated annexin VCaB42.
Length = 316
Score = 131 bits (329), Expect = 9e-30
Identities = 69/187 (36%), Positives = 108/187 (57%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASL +P ++ P +DA L+KAF G+G + +I IL HR++ QR LI++ Y A Y E+
Sbjct: 1 MASLKVPTSVPEPYEDAEQLKKAFAGWGTNEALIIQILAHRNAAQRKLIRETYAAAYGED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + +EL + ++A+LLW L PA RD ++ EA T +I C+R+ L
Sbjct: 61 LLKDLDAELTSDFQRAVLLWTLSPAERDAYLVNEATKRLTSSNWVILEIACTRSSDDLFK 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+Q Y+AR+ LE D+ +HT+GD + LL+ + RYEG E + T+ +A L++
Sbjct: 121 ARQAYHARYKKSLEEDVAYHTTGDFRKLLVPLLTAFRYEGEEANMTLARKEANILHEKIS 180
Query: 583 XRLGTDE 603
+ DE
Sbjct: 181 DKAYNDE 187
Score = 45.4 bits (106), Expect = 7e-04
Identities = 34/148 (22%), Positives = 64/148 (43%), Gaps = 3/148 (2%)
Frame = +1
Query: 70 MANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELSHRISSEL 249
M R++A L + + +I I++ R Q Y + E++ + ++
Sbjct: 165 MTLARKEANILHEKISDKAYNDEELIRIISTRSKAQLNATFNHYLDQHGSEINKDLETDS 224
Query: 250 NGNHKK---AMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYY 420
+ + K A + + P VLR A+ G D T ++ +R + +K+ Y+
Sbjct: 225 DDEYLKLLSAAIECLKTPEKHFEKVLRLAIKGTGTDEWDLTRVVTTRAEVDMERIKEEYH 284
Query: 421 ARFGTYLEHDIGHHTSGDHQXLLLAYVG 504
R L+ I TSGD++ +LLA +G
Sbjct: 285 KRNSVPLDRAIAGDTSGDYERMLLALIG 312
>sptr|Q9SB88|Q9SB88 Annexin cap32.
Length = 314
Score = 131 bits (329), Expect = 9e-30
Identities = 66/187 (35%), Positives = 110/187 (58%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASLT+P + + +D L+ AFKG+G + +I+IL HR + QR LI+Q Y + E+
Sbjct: 1 MASLTVPAHVPSAAEDCEQLRSAFKGWGTNHKLIISILAHRTAAQRKLIRQTYAETFGED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL + +K +L+W LDP+ RD + +EA T ++ C+R+P +L +
Sbjct: 61 LLKELDRELTHDFEKLVLVWTLDPSERDAHLAKEATKRWTKSNFVLVELACTRSPKELVL 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
++ Y+AR+ LE D+ +HT+GDH+ LL+ V RY G EVD + ++K L++
Sbjct: 121 AREAYHARYKKSLEEDVAYHTTGDHRKLLVPLVSSYRYGGEEVDLRLAKAESKILHEKIS 180
Query: 583 XRLGTDE 603
+ +D+
Sbjct: 181 DKAYSDD 187
Score = 36.2 bits (82), Expect = 0.40
Identities = 32/125 (25%), Positives = 54/125 (43%), Gaps = 4/125 (3%)
Frame = +1
Query: 142 VINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNHKKAMLL----WILDPAGRDX 309
VI IL R Q Y+ + E++ ++ +G+ A+L ++ P
Sbjct: 189 VIRILATRSKAQLNATLNHYKDEHGEDILKQLE---DGDEFVALLRATIKGLVYPEHYFV 245
Query: 310 TVLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFGTYLEHDIGHHTSGDHQXLL 489
VLR+A++ + T +I +R L I+ Y R L I T GD++ +L
Sbjct: 246 EVLRDAINRRGTEEDHLTRVIATRAEVDLKIIADEYQKRDSIPLGRAIAKDTRGDYESML 305
Query: 490 LAYVG 504
LA +G
Sbjct: 306 LALLG 310
>sptr|Q9M3H3|Q9M3H3 Annexin p34.
Length = 314
Score = 131 bits (329), Expect = 9e-30
Identities = 67/187 (35%), Positives = 110/187 (58%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASLT+P + + +D L+ AFKG+G + +I+IL HR++ QR LI+Q Y + E+
Sbjct: 1 MASLTVPAEVPSVAEDCEQLRSAFKGWGTNEKLIISILAHRNAAQRKLIRQTYAETFGED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL + +K +L+W LDP+ RD + +EA T +I C+R+P +L +
Sbjct: 61 LLKELDRELTHDFEKLVLIWTLDPSERDAYLAKEATKRWTKSNFVLVEIACTRSPKELVL 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
++ Y+AR LE D+ +HT+GDH+ LL+ V RY G EVD + ++K L++
Sbjct: 121 AREAYHARNKKSLEEDVAYHTTGDHRKLLVPLVSSYRYGGDEVDLRLAKAESKVLHEKIS 180
Query: 583 XRLGTDE 603
+ +D+
Sbjct: 181 DKAYSDD 187
>sptr|O81536|O81536 Annexin p34.
Length = 314
Score = 130 bits (328), Expect = 1e-29
Identities = 67/187 (35%), Positives = 110/187 (58%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASLT+P + + +D L+ AFKG+G + +I+IL HR++ QR LI+Q Y + E+
Sbjct: 1 MASLTVPAEVPSVAEDCEQLRSAFKGWGTNEKLIISILAHRNAAQRKLIRQTYAETFGED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL + +K +++W LDPA RD + +EA T +I C+R+P +L +
Sbjct: 61 LLKELDRELTHDFEKLVVVWTLDPAERDAYLAKEATKRWTKSNFVLVEIACTRSPKELVL 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
++ Y+AR LE D+ +HT+GDH+ LL+ V RY G EVD + ++K L++
Sbjct: 121 AREAYHARNKKSLEEDVAYHTTGDHRKLLVPLVSSYRYGGDEVDLRLAKAESKVLHEKIS 180
Query: 583 XRLGTDE 603
+ +D+
Sbjct: 181 DKAYSDD 187
>sptr|Q9XEE2|Q9XEE2 Putative annexin.
Length = 317
Score = 130 bits (327), Expect = 2e-29
Identities = 69/187 (36%), Positives = 108/187 (57%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASL +P + P DA L KAF G+G + +I+IL HR++ QR LI+ Y A Y+E+
Sbjct: 1 MASLKVPSNVPLPEDDAEQLHKAFSGWGTNEKLIISILAHRNAAQRSLIRSVYAATYNED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL+ + ++A++LW LDP RD + +E+ T + +I C+R +L
Sbjct: 61 LLKALDKELSSDFERAVMLWTLDPPERDAYLAKESTKMFTKNNWVLVEIACTRPALELIK 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+KQ Y AR+ +E D+ HTSGD + LLL V RYEG +V+ + +AK L++
Sbjct: 121 VKQAYQARYKKSIEEDVAQHTSGDLRKLLLPLVSTFRYEGDDVNMMLARSEAKILHEKVS 180
Query: 583 XRLGTDE 603
+ +D+
Sbjct: 181 EKSYSDD 187
>sptr|Q96527|Q96527 Annexin-like protein.
Length = 317
Score = 130 bits (327), Expect = 2e-29
Identities = 67/187 (35%), Positives = 111/187 (59%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MA+L + ++ P DA L+ AF+G+G + +I+IL HR + QR +I+Q Y Y ++
Sbjct: 1 MATLKVSDSVPAPSDDAEQLRTAFEGWGTNEDLIISILAHRSAEQRKVIRQAYHETYGKD 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL+ + ++A+LLW L+P RD + EA T + ++ C+RT +QL
Sbjct: 61 LLKTLDKELSNDFERAILLWTLEPGERDALLANEATKRWTSSNQVLMEVACTRTSTQLLH 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+Q Y+AR+ LE D+ HHT+GD + LL++ V RYEG EV+ T+ +AK +++ +
Sbjct: 121 ARQAYHARYKKSLEEDVAHHTTGDFRKLLVSLVTSYRYEGDEVNMTLAKQEAKLVHEKIK 180
Query: 583 XRLGTDE 603
+ DE
Sbjct: 181 DKHYNDE 187
>sptr|Q9C5V2|Q9C5V2 Calcium-binding protein annexin 7.
Length = 316
Score = 130 bits (326), Expect = 2e-29
Identities = 67/187 (35%), Positives = 112/187 (59%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASL +P + P +DA L K+FKG+G + +I+IL HR++ QR I+ Y A Y+++
Sbjct: 1 MASLKVPATVPLPEEDAEQLYKSFKGWGTNERMIISILAHRNATQRSFIRAVYAANYNKD 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL+G+ ++A++LW +PA R + +E+ T + +I C+R+ +L
Sbjct: 61 LLKELDRELSGDFERAVMLWTFEPAERYAYLAKESTKMFTKNNWVLVEIACTRSALELFN 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+Q Y AR+ T LE D+ +HTSGD + LL+ V RY+G EV+ T+ +AK L++ +
Sbjct: 121 ARQAYQARYKTSLEEDVAYHTSGDIRKLLVPLVSTFRYDGDEVNMTLARSEAKILHEKIK 180
Query: 583 XRLGTDE 603
+ D+
Sbjct: 181 EKAYADD 187
Score = 42.0 bits (97), Expect = 0.007
Identities = 35/148 (23%), Positives = 61/148 (41%), Gaps = 3/148 (2%)
Frame = +1
Query: 70 MANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELSHRISSEL 249
M R +A L + K +I ILT R Q Y+ + +S + +
Sbjct: 165 MTLARSEAKILHEKIKEKAYADDDLIRILTTRSKAQISATLNHYKNNFGTSMSKYLKEDS 224
Query: 250 NGNH---KKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYY 420
+ KA++ + P VLR+A++ D T ++ +R + +K+ Y
Sbjct: 225 ENEYIQLLKAVIKCLTYPEKYFEKVLRQAINKLGTDEWGLTRVVTTRAEFVMERIKEEYI 284
Query: 421 ARFGTYLEHDIGHHTSGDHQXLLLAYVG 504
R L+ I T GD++ +LLA +G
Sbjct: 285 RRNSVPLDRAIAKDTHGDYEDILLALLG 312
>sptr|Q39001|Q39001 Annexin (Fragment).
Length = 314
Score = 128 bits (322), Expect = 6e-29
Identities = 66/185 (35%), Positives = 109/185 (58%)
Frame = +1
Query: 49 SLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELS 228
+L + ++ P DA L+ AF+G+G + +I+IL HR + QR +I+Q Y Y E+L
Sbjct: 1 TLKVSDSVPAPSDDAEQLRTAFEGWGTNEDLIISILAHRSAEQRKVIRQAYHETYGEDLL 60
Query: 229 HRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXIMK 408
+ EL+ + ++A+LLW L+P RD + EA T + ++ C+RT +QL +
Sbjct: 61 KTLDKELSNDFERAILLWTLEPGERDALLANEATKRWTSSNQVLMEVACTRTSTQLLHAR 120
Query: 409 QTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGEXR 588
Q Y+AR+ LE D+ HHT+GD + LL++ V RYEG EV+ T+ +AK +++ + +
Sbjct: 121 QAYHARYKKSLEEDVAHHTTGDFRKLLVSLVTSYRYEGDEVNMTLAKQEAKLVHEKIKDK 180
Query: 589 LGTDE 603
DE
Sbjct: 181 HYNDE 185
>sptr|Q9LX08|Q9LX08 Annexin-like protein.
Length = 318
Score = 128 bits (321), Expect = 8e-29
Identities = 68/189 (35%), Positives = 109/189 (57%), Gaps = 2/189 (1%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASL +P + P +D+ L KAFKG+G + +I+IL HR++ QR I+ Y A Y+++
Sbjct: 1 MASLKIPANIPLPEEDSEQLHKAFKGWGTNEGMIISILAHRNATQRSFIRAVYAANYNKD 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL+G+ ++ ++LW LDP RD + E+ T ++ +I C+R +
Sbjct: 61 LLKELDGELSGDFERVVMLWTLDPTERDAYLANESTKLFTKNIWVLVEIACTRPSLEFFK 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEG--PEVDPTIXTHDAKDLYKA 576
KQ Y+ R+ T LE D+ +HTSG+ + LL+ V RY+G EV+ + +AK L+K
Sbjct: 121 TKQAYHVRYKTSLEEDVAYHTSGNIRKLLVPLVSTFRYDGNADEVNVKLARSEAKTLHKK 180
Query: 577 GEXRLGTDE 603
+ TDE
Sbjct: 181 ITEKAYTDE 189
Score = 44.7 bits (104), Expect = 0.001
Identities = 35/144 (24%), Positives = 61/144 (42%), Gaps = 3/144 (2%)
Frame = +1
Query: 82 RQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNH 261
R +A L K +I ILT R Q ++ + ++ + + N ++
Sbjct: 171 RSEAKTLHKKITEKAYTDEDLIRILTTRSKAQINATLNHFKDKFGSSINKFLKEDSNDDY 230
Query: 262 K---KAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFG 432
K + + P VLR A++ D A T ++ +R L +K+ Y R
Sbjct: 231 VQLLKTAIKCLTYPEKYFEKVLRRAINRMGTDEWALTRVVTTRAEVDLERIKEEYLRRNS 290
Query: 433 TYLEHDIGHHTSGDHQXLLLAYVG 504
L+ I + TSGD++ +LLA +G
Sbjct: 291 VPLDRAIANDTSGDYKDMLLALLG 314
>sptr|O81535|O81535 Annexin p35.
Length = 315
Score = 127 bits (320), Expect = 1e-28
Identities = 69/187 (36%), Positives = 107/187 (57%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
M+SL +P ++ +P +DA L+KAFKG+G + +I IL HR++ QR LI+ Y A Y E+
Sbjct: 1 MSSLKVPASVPDPYEDAEQLKKAFKGWGTNEELIIQILAHRNARQRKLIRDSYAAAYGED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + SEL + ++ +LLW L PA RD ++ EA T +I C+R+ L
Sbjct: 61 LLKDLDSELTSDFQRVVLLWTLSPAERDAYLVNEATKRLTASNWGIMEIACTRSSDDLFK 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+Q Y+A + LE D+ +HT GD + LL+ + RYEG EV+ T+ +K L++
Sbjct: 121 ARQAYHAPYKKSLEEDVAYHTVGDFRKLLVPLITAFRYEGDEVNMTLARKGSKYLHEKIS 180
Query: 583 XRLGTDE 603
+ DE
Sbjct: 181 DKAYHDE 187
Score = 40.8 bits (94), Expect = 0.016
Identities = 30/123 (24%), Positives = 54/123 (43%), Gaps = 2/123 (1%)
Frame = +1
Query: 142 VINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNHKKAMLLWI--LDPAGRDXTV 315
+I I++ R Q Y + E+ + ++ + + K + I L P V
Sbjct: 189 IIRIISTRSKAQLSATFNHYHDHHGHEIIKDLEADDDDEYLKLLRAAIECLKPREHFEKV 248
Query: 316 LREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFGTYLEHDIGHHTSGDHQXLLLA 495
LR A+ D T ++ +R + +K+ Y+ R L+ I TSGD++ +LLA
Sbjct: 249 LRLAIKKLGTDEWDLTRVVATRAEVDMERIKEEYHRRNSVTLDRAIAGDTSGDYEKMLLA 308
Query: 496 YVG 504
+G
Sbjct: 309 LIG 311
>sptr|Q9C5V3|Q9C5V3 Calcium-binding protein annexin 6.
Length = 318
Score = 127 bits (319), Expect = 1e-28
Identities = 68/189 (35%), Positives = 108/189 (57%), Gaps = 2/189 (1%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MASL +P + P +D+ L KAFKG+G + +I+IL HR++ QR I+ Y A Y+++
Sbjct: 1 MASLKIPANIPLPEEDSEQLHKAFKGWGTNEGMIISILAHRNATQRSFIRAVYAANYNKD 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL+G+ ++ ++LW LDP RD E+ T ++ +I C+R +
Sbjct: 61 LLKELDGELSGDFERVVMLWTLDPTERDAYSANESTKMFTKNIWVLVEIACTRPSLEFFK 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEG--PEVDPTIXTHDAKDLYKA 576
KQ Y+ R+ T LE D+ +HTSG+ + LL+ V RY+G EV+ + +AK L+K
Sbjct: 121 TKQAYHVRYKTSLEEDVAYHTSGNIRKLLVPLVSTFRYDGNADEVNVKLARSEAKTLHKK 180
Query: 577 GEXRLGTDE 603
+ TDE
Sbjct: 181 ITEKAYTDE 189
Score = 43.1 bits (100), Expect = 0.003
Identities = 35/144 (24%), Positives = 60/144 (41%), Gaps = 3/144 (2%)
Frame = +1
Query: 82 RQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNH 261
R +A L K +I ILT R Q + + ++ + + N ++
Sbjct: 171 RSEAKTLHKKITEKAYTDEDLIRILTTRSKAQINATLNHLKDKFGSSINKFLKEDSNDDY 230
Query: 262 K---KAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFG 432
K + + P VLR A++ D A T ++ +R L +K+ Y R
Sbjct: 231 VQLLKTAIKCLTYPEKYFEKVLRRAINRMGTDEWALTRVVTTRAEVDLERIKEEYLRRNS 290
Query: 433 TYLEHDIGHHTSGDHQXLLLAYVG 504
L+ I + TSGD++ +LLA +G
Sbjct: 291 VPLDRAIANDTSGDYKDMLLALLG 314
>sptr|Q9FUG5|Q9FUG5 Annexin.
Length = 334
Score = 126 bits (317), Expect = 2e-28
Identities = 66/201 (32%), Positives = 106/201 (52%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
M+++T+P + + +D I L KA + FGCD ++N++ HRD QR I+ Y Y E+
Sbjct: 1 MSTITVPNPVPDTNEDCITLHKALEDFGCDKEALLNVICHRDQQQRQRIRHSYNRKYEED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
+ + S+L+ +K +LW+ DPA RD T+L EAL + D A T+++ RT ++L
Sbjct: 61 ILKTLKSKLHAKLEKGAVLWMCDPAERDATILHEALRCMSKDYSALTEVLYLRTSAELLD 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+++ Y +RFG LE ++ G + LLL + R E E+D D KDL A
Sbjct: 121 IRRAYSSRFGRSLEEELATKIDGSEKKLLLGLLREARSEDDEIDTLQVEADTKDLLSAIS 180
Query: 583 XRLGTDEXXXXXXFTXRXWAH 645
++ FT R +H
Sbjct: 181 NTKEVNKSVIIRVFTTRSSSH 201
>sptr|O22341|O22341 Annexin.
Length = 316
Score = 126 bits (316), Expect = 3e-28
Identities = 66/187 (35%), Positives = 106/187 (56%)
Frame = +1
Query: 43 MASLTLPPAMANPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEE 222
MA+LT+P + + +D L+KAF G+G + +INIL HR++ +R I++ Y + E+
Sbjct: 1 MATLTVPSTLPSVSEDCEQLRKAFSGWGTNEDLIINILGHRNADERNSIRKAYTETHGED 60
Query: 223 LSHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXI 402
L + EL+ + ++ +LLW LDP RD + EA T + +I C + QL
Sbjct: 61 LLKALDKELSNDFERLVLLWTLDPPERDALLANEATKRWTSSNQVIMEIACRSSSDQLLR 120
Query: 403 MKQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGE 582
+Q Y+ R+ LE D+ HHT+GD + LLL V RYEG EV+ T+ +AK L++
Sbjct: 121 ARQAYHVRYKKSLEEDVAHHTTGDFRKLLLPLVSSYRYEGDEVNMTLAKTEAKLLHEKIS 180
Query: 583 XRLGTDE 603
+ +D+
Sbjct: 181 NKAYSDD 187
Score = 33.1 bits (74), Expect = 3.3
Identities = 29/124 (23%), Positives = 51/124 (41%), Gaps = 3/124 (2%)
Frame = +1
Query: 142 VINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNHK---KAMLLWILDPAGRDXT 312
VI +L R Q Y+ Y +++ + ++ ++ + ++ P
Sbjct: 189 VIRVLATRSKSQINERLNHYKNEYATDINKDLKADPKDEFLALLRSTVKCLVYPEKYFEK 248
Query: 313 VLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFGTYLEHDIGHHTSGDHQXLLL 492
VLR A++ D A T ++ +R L I+ Y R L I T+GD++ LLL
Sbjct: 249 VLRLAINKRGTDEGALTRVVSTRAEVDLKIIADEYQRRNSVPLTRAIVKDTNGDYEKLLL 308
Query: 493 AYVG 504
G
Sbjct: 309 VLAG 312
>sptr|P93158|P93158 Annexin (Fragment).
Length = 315
Score = 124 bits (310), Expect = 1e-27
Identities = 69/186 (37%), Positives = 110/186 (59%), Gaps = 1/186 (0%)
Frame = +1
Query: 49 SLTLPPAMANPRQDAI-DLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEEL 225
+L +P + +P +DA L+KAF+G+G + +I+IL HR++ QR I++ Y Y E+L
Sbjct: 1 TLKVPVHVPSPSEDAEWQLRKAFEGWGTNEQLIIDILAHRNAAQRNSIRKVYGEAYGEDL 60
Query: 226 SHRISSELNGNHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXIM 405
+ EL + ++A+LL+ LDPA RD + EA T +I CSR+ +L +
Sbjct: 61 LKCLEKELTSDFERAVLLFTLDPAERDAHLANEATKKFTSSNWILMEIACSRSSHELLNV 120
Query: 406 KQTYYARFGTYLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYKAGEX 585
K+ Y+AR+ LE D+ HHT+G+++ LL+ V RYEG EV+ T+ +AK L+
Sbjct: 121 KKAYHARYKKSLEEDVAHHTTGEYRKLLVPLVSAFRYEGEEVNMTLAKSEAKILHDKISD 180
Query: 586 RLGTDE 603
+ TDE
Sbjct: 181 KHYTDE 186
Score = 43.5 bits (101), Expect = 0.002
Identities = 31/124 (25%), Positives = 55/124 (44%), Gaps = 3/124 (2%)
Frame = +1
Query: 142 VINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNHKK---AMLLWILDPAGRDXT 312
VI I++ R Q Y + ++ + ++ + K A++ + P
Sbjct: 188 VIRIVSTRSKAQLNATLNHYNTSFGNAINKDLKADPSDEFLKLLRAVIKCLTTPEQYFEK 247
Query: 313 VLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFGTYLEHDIGHHTSGDHQXLLL 492
VLR+A++ D A T ++ +R + +K+ Y R LE I TSGD++ LL
Sbjct: 248 VLRQAINKLGSDEWALTRVVTTRAEVDMVRIKEAYQRRNSIPLEQAIAKDTSGDYEKFLL 307
Query: 493 AYVG 504
A +G
Sbjct: 308 ALIG 311
>sptr|Q42922|Q42922 Annexin (Fragment).
Length = 308
Score = 123 bits (309), Expect = 2e-27
Identities = 65/166 (39%), Positives = 101/166 (60%)
Frame = +1
Query: 76 NPRQDAIDLQKAFKGFGCDSTTVINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNG 255
+P +D+ L+ AF+G+G + +I+IL HR++ QR I++ Y + E+L + EL+
Sbjct: 5 SPSEDSEQLRGAFQGWGTNEGLIISILAHRNAAQRKSIRETYTQTHGEDLLKDLDKELSS 64
Query: 256 NHKKAMLLWILDPAGRDXTVLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFGT 435
+ +KA+LLW LDPA RD + +A T + +I +R+P +L KQ Y RF
Sbjct: 65 DFEKAVLLWTLDPAERDAFLANQATKMLTSNNSIIVEIASTRSPLELLKAKQAYQVRFKK 124
Query: 436 YLEHDIGHHTSGDHQXLLLAYVGIPRYEGPEVDPTIXTHDAKDLYK 573
LE D+ +HTSGD + LL+ VGI RYEG EV+ T+ +AK L++
Sbjct: 125 SLEEDVAYHTSGDIRKLLVPLVGIHRYEGDEVNMTLAKSEAKLLHE 170
Score = 38.5 bits (88), Expect = 0.080
Identities = 30/124 (24%), Positives = 53/124 (42%), Gaps = 3/124 (2%)
Frame = +1
Query: 142 VINILTHRDSVQRGLIQQEYRAMYHEELSHRISSELNGNHKK---AMLLWILDPAGRDXT 312
+I I+T R Q Y + + + ++ + + K A + + P
Sbjct: 182 LIRIVTTRSKAQLNATLNHYNNEFGNVIDKDLETDSDDEYLKLLRAAIKGLTYPEKYFEE 241
Query: 313 VLREALSGDTMDLRAATDIICSRTPSQLXIMKQTYYARFGTYLEHDIGHHTSGDHQXLLL 492
+LR A++ D A T ++ +R L + + Y R L+ I TSGD+Q +LL
Sbjct: 242 LLRLAINKMGTDENALTRVVTTRAEVDLQRIAEEYQRRNSVPLDRAIDKDTSGDYQKILL 301
Query: 493 AYVG 504
A +G
Sbjct: 302 ALMG 305
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 449,707,047
Number of Sequences: 1272877
Number of extensions: 8744581
Number of successful extensions: 20577
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 19595
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 20446
length of database: 406,886,216
effective HSP length: 118
effective length of database: 256,686,730
effective search space used: 27978853570
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

