BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= QAY3d01.yg.3.1
(1174 letters)
Database: swall
1,381,838 sequences; 439,479,560 total letters
Searching..................................................done
Score E
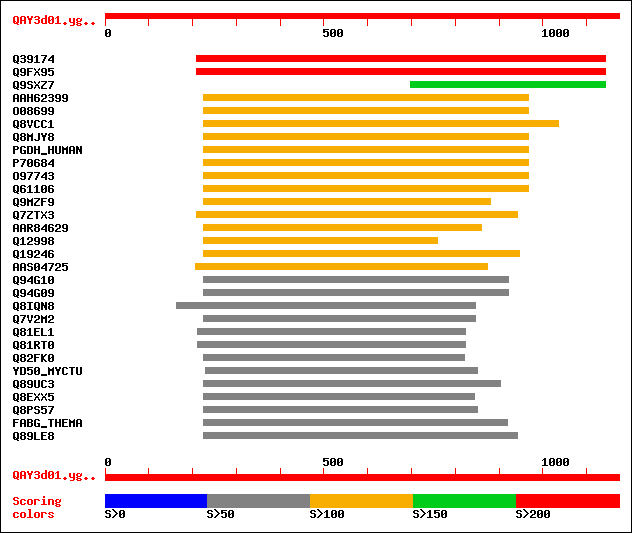
Sequences producing significant alignments: (bits) Value
sptr|Q39174|Q39174 Putative alcohol dehydrogenase (EC 1.1.1.1) /... 385 e-106
sptr|Q9FX95|Q9FX95 ARP protein (At1g49670). 384 e-105
sptr|Q9SXZ7|Q9SXZ7 NADPH oxidoreductase homolog (Fragment). 183 5e-45
sptrnew|AAH62399|AAH62399 NAD-dependent 15-hydroxyprostaglandin ... 139 6e-32
sptr|O08699|O08699 NAD-dependent 15-hydroxyprostaglandin dehydro... 139 6e-32
sptr|Q8VCC1|Q8VCC1 Similar to hydroxyprostaglandin dehydrogenase... 139 6e-32
sptr|Q8MJY8|Q8MJY8 Prostaglandin dehydrogenase I. 136 7e-31
sw|P15428|PGDH_HUMAN 15-hydroxyprostaglandin dehydrogenase [NAD(... 136 7e-31
sptr|P70684|P70684 15-hydroxyprostaglandin dehydrogenase (NAD+) ... 135 2e-30
sptr|O97743|O97743 NAD+-dependent 15-hydoxyprostaglandin dehydro... 132 8e-30
sptr|Q61106|Q61106 NAD(+)-dependent 15-hydroxyprostaglandin dehy... 131 2e-29
sptr|Q9MZF9|Q9MZF9 Prostaglandin dehydrogenase (Fragment). 125 1e-27
sptr|Q7ZTX3|Q7ZTX3 Similar to hydroxyprostaglandin dehydrogenase... 123 5e-27
sptrnew|AAR84629|AAR84629 Putative alcohol dehydrogenase. 108 1e-22
sptr|Q12998|Q12998 15-hydroxy prostaglandin dehydrogenase. 107 5e-22
sptr|Q19246|Q19246 Hypothetical protein F09E10.3. 103 7e-21
sptrnew|AAS04725|AAS04725 FabG2_2. 101 2e-20
sptr|Q94G10|Q94G10 CTA. 100 3e-20
sptr|Q94G09|Q94G09 Sex determination protein. 100 4e-20
sptr|Q8IQN8|Q8IQN8 CG4899-PB. 100 4e-20
sptr|Q7V2M2|Q7V2M2 3-oxoacyl-[acyl-carrier protein] reductase (E... 100 7e-20
sptr|Q81EL1|Q81EL1 3-oxoacyl-[acyl-carrier protein] reductase (E... 99 1e-19
sptr|Q81RT0|Q81RT0 Oxidoreductase, short-chain dehydrogenase/red... 99 1e-19
sptr|Q82FK0|Q82FK0 Putative oxidoreductase. 99 1e-19
sw|Q11020|YD50_MYCTU Putative oxidoreductase Rv1350/MT1393/Mb138... 99 1e-19
sptr|Q89UC3|Q89UC3 D-beta-hydroxybutyrate dehydrogenase. 99 1e-19
sptr|Q8EXX5|Q8EXX5 3-ketoacyl-acyl carrier protein reductase (EC... 99 2e-19
sptr|Q8PS57|Q8PS57 Putative ketoreductase. 98 2e-19
sw|Q9X248|FABG_THEMA 3-oxoacyl-[acyl-carrier protein] reductase ... 98 3e-19
sptr|Q89LE8|Q89LE8 Short chain dehydrogenase. 97 4e-19
>sptr|Q39174|Q39174 Putative alcohol dehydrogenase (EC 1.1.1.1) /
probable NADP-dependent oxidoreductase P2 (EC 1.-.-.-).
Length = 629
Score = 385 bits (989), Expect = e-106
Identities = 177/312 (56%), Positives = 244/312 (78%)
Frame = +2
Query: 209 MELKPGMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGD 388
ME+KPG+SALVTGGASGIG+ALC+A A +G+FVTV DFSEE G+E +LV+K N+KFH
Sbjct: 1 MEIKPGLSALVTGGASGIGRALCLALAEKGVFVTVADFSEEKGQETTSLVRKANAKFHHG 60
Query: 389 LRIPSSIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWR 568
L PS+IFVKCDV+N +L A F+KH+ T+ LDICIN AGI+ D +DG+++W+
Sbjct: 61 LNFPSAIFVKCDVTNRGDLLAAFDKHLATFGTLDICINNAGISTPLRFDKDDTDGSKSWK 120
Query: 569 HAVNVNLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTR 748
H +NV+L+AV++GT++A + M+ ++KPGVIIN+GSAAGLYPM +DP+Y+A+K GVV+FTR
Sbjct: 121 HTINVDLIAVVEGTQLAIKAMKAKQKPGVIINMGSAAGLYPMPIDPIYAASKAGVVLFTR 180
Query: 749 SLSPLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSSGGYLEMEDVVNGTFELIQDESKAG 928
SL+ +R G+R+NVLCPEF++T++AE + +++S GGY+ M+ ++ G FELI DE K G
Sbjct: 181 SLAYYRRQGIRINVLCPEFIKTDLAEAIDASILESIGGYMSMDMLIKGAFELITDEKKVG 240
Query: 929 ACLWITKRRGMEYWPTPEEQRKYMVNPNKSKRMLTNNIYPSIRMPEFFEKIVVHTLSHNF 1108
ACLWITKRRG+EYWPTP E+ KY+V + KR + + I P+ FEK++VHTLSH F
Sbjct: 241 ACLWITKRRGLEYWPTPMEETKYLVGSSSRKRP-SFKVSTKIEFPQSFEKMIVHTLSHKF 299
Query: 1109 RNATRLERVQLR 1144
R+ATR+ R L+
Sbjct: 300 RSATRIVRAPLQ 311
>sptr|Q9FX95|Q9FX95 ARP protein (At1g49670).
Length = 629
Score = 384 bits (987), Expect = e-105
Identities = 177/312 (56%), Positives = 245/312 (78%)
Frame = +2
Query: 209 MELKPGMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGD 388
ME+KPG+SALVTGGASGIG+ALC+A A +G+FVTV DFSEE G+E +LV++ N+KFH
Sbjct: 1 MEIKPGLSALVTGGASGIGRALCLALAEKGVFVTVADFSEEKGQETTSLVREANAKFHQG 60
Query: 389 LRIPSSIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWR 568
L PS+IFVKCDV+N +L A F+KH+ T+ LDICIN AGI+ D +DG+++W+
Sbjct: 61 LSFPSAIFVKCDVTNRGDLLAAFDKHLATFGTLDICINNAGISTPLRFDKDDTDGSKSWK 120
Query: 569 HAVNVNLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTR 748
H +NV+L+AV++GT++A + M+ ++KPGVIIN+GSAAGLYPM +DP+Y+A+K GVV+FTR
Sbjct: 121 HTINVDLIAVVEGTQLAIKAMKAKQKPGVIINMGSAAGLYPMPVDPIYAASKAGVVLFTR 180
Query: 749 SLSPLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSSGGYLEMEDVVNGTFELIQDESKAG 928
SL+ +R G+R+NVLCPEF++T++AE + +++S GGY+ M+ ++ G FELI DE KAG
Sbjct: 181 SLAYYRRQGIRINVLCPEFIKTDLAEAIDASILESIGGYMSMDMLIKGAFELITDEKKAG 240
Query: 929 ACLWITKRRGMEYWPTPEEQRKYMVNPNKSKRMLTNNIYPSIRMPEFFEKIVVHTLSHNF 1108
ACLWITKRRG+EYWPTP E+ KY+V + KR + + I P+ FEK++VHTLSH F
Sbjct: 241 ACLWITKRRGLEYWPTPMEETKYLVGSSSRKRP-SFKVSTKIEFPQSFEKMIVHTLSHKF 299
Query: 1109 RNATRLERVQLR 1144
R+ATR+ R L+
Sbjct: 300 RSATRIVRAPLQ 311
>sptr|Q9SXZ7|Q9SXZ7 NADPH oxidoreductase homolog (Fragment).
Length = 470
Score = 183 bits (464), Expect = 5e-45
Identities = 87/149 (58%), Positives = 110/149 (73%)
Frame = +2
Query: 698 LDPVYSAAKGGVVMFTRSLSPLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSSGGYLEME 877
LDP+YS +KGGV+MF+R+L KR G+RVNVLCPEFV+T+M + K + GG++ ME
Sbjct: 1 LDPIYSGSKGGVIMFSRALRLYKRQGIRVNVLCPEFVETDMGTMIGPKFLSMMGGFVPME 60
Query: 878 DVVNGTFELIQDESKAGACLWITKRRGMEYWPTPEEQRKYMVNPNKSKRMLTNNIYPSIR 1057
VV G FELI DE+KAG CLWIT RRG+EYWPTP E+ KY++ +S+R T P I+
Sbjct: 61 MVVKGAFELITDENKAGDCLWITNRRGLEYWPTPSEEAKYLLRSTRSRRR-TEYKAPPIK 119
Query: 1058 MPEFFEKIVVHTLSHNFRNATRLERVQLR 1144
+PE FEKIVV TL+HNFRNAT + R LR
Sbjct: 120 LPESFEKIVVQTLTHNFRNATSVVRAPLR 148
>sptrnew|AAH62399|AAH62399 NAD-dependent 15-hydroxyprostaglandin
dehydrogenase.
Length = 266
Score = 139 bits (351), Expect = 6e-32
Identities = 90/269 (33%), Positives = 138/269 (51%), Gaps = 21/269 (7%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G ALVTG A GIGKA A G V +VD++ E G + + ++
Sbjct: 5 GKVALVTGAAQGIGKAFTEALLLHGAKVALVDWNLETGVKCKAALDEQ-------FEPQK 57
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
++F++CDV++ L F K V + LDI +N AG+ N+ + W + +
Sbjct: 58 TLFIQCDVADQKQLRDTFRKVVDHFGRLDILVNNAGVNNE-----------KNWEQTLQI 106
Query: 584 NLVAVIDGTRIASQIMRNQK--KPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRS-- 751
NLV+VI GT + M Q + G+IINI S AGL P+ PVY A+K G++ FTRS
Sbjct: 107 NLVSVISGTYLGLDYMSKQNGGEGGIIINISSIAGLMPVAQQPVYCASKHGIIGFTRSAA 166
Query: 752 -LSPLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSSGGYLEMED---------------- 880
+ L + GVR+NV+CP FV+T + E + ++ ++ G Y+E D
Sbjct: 167 MAANLMKSGVRLNVICPGFVKTPILESIEKE--ENMGQYIEYTDQIKAMMKFYGILDPSA 224
Query: 881 VVNGTFELIQDESKAGACLWITKRRGMEY 967
+ NG LI+D++ GA + IT +G+ +
Sbjct: 225 IANGLINLIEDDALNGAIMKITASKGIHF 253
>sptr|O08699|O08699 NAD-dependent 15-hydroxyprostaglandin
dehydrogenase.
Length = 266
Score = 139 bits (351), Expect = 6e-32
Identities = 90/269 (33%), Positives = 138/269 (51%), Gaps = 21/269 (7%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G ALVTG A GIGKA A G V +VD++ E G + + ++
Sbjct: 5 GKVALVTGAAQGIGKAFTEALLLHGAKVALVDWNLETGVKCKAALDEQ-------FEPQK 57
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
++F++CDV++ L F K V + LDI +N AG+ N+ + W + +
Sbjct: 58 TLFIQCDVADQKQLRDTFRKVVDHFGRLDILVNNAGVNNE-----------KNWEQTLQI 106
Query: 584 NLVAVIDGTRIASQIMRNQK--KPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRS-- 751
NLV+VI GT + M Q + G+IINI S AGL P+ PVY A+K G++ FTRS
Sbjct: 107 NLVSVISGTYLGLDYMSKQNGGEGGIIINISSIAGLMPVTQQPVYCASKHGIIGFTRSAA 166
Query: 752 -LSPLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSSGGYLEMED---------------- 880
+ L + GVR+NV+CP FV+T + E + ++ ++ G Y+E D
Sbjct: 167 MAANLMKSGVRLNVICPGFVKTPILESIEKE--ENMGQYIEYTDQIKAMMKFYGILDPSA 224
Query: 881 VVNGTFELIQDESKAGACLWITKRRGMEY 967
+ NG LI+D++ GA + IT +G+ +
Sbjct: 225 IANGLINLIEDDALNGAIMKITASKGIHF 253
>sptr|Q8VCC1|Q8VCC1 Similar to hydroxyprostaglandin dehydrogenase 15
(NAD).
Length = 269
Score = 139 bits (351), Expect = 6e-32
Identities = 95/292 (32%), Positives = 147/292 (50%), Gaps = 21/292 (7%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G ALVTG A GIGKA A G V +VD++ E G + + ++
Sbjct: 5 GKVALVTGAAQGIGKAFAEALLLHGAKVALVDWNLEAGVKCKAALDEQ-------FEPQK 57
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
++FV+CDV++ L F K V + LDI +N AG+ N+ + W + +
Sbjct: 58 TLFVQCDVADQKQLRDTFRKVVDHFGRLDILVNNAGVNNE-----------KNWEQTLQI 106
Query: 584 NLVAVIDGTRIASQIMRNQK--KPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRS-- 751
NLV+VI GT + M Q + G+IIN+ S AGL P+ PVY A+K G++ FTRS
Sbjct: 107 NLVSVISGTYLGLDYMSKQNGGEGGIIINMSSLAGLMPVAQQPVYCASKHGIIGFTRSAA 166
Query: 752 -LSPLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSSGGYLEMED---------------- 880
+ L + GVR+NV+CP FV T + E + ++ ++ G Y+E +D
Sbjct: 167 MAANLMKSGVRLNVICPGFVDTPILESIEKE--ENMGQYIEYKDQIKAMMKFYGVLHPST 224
Query: 881 VVNGTFELIQDESKAGACLWITKRRGMEYWPTPEEQRKYMVNPNKSKRMLTN 1036
+ NG LI+D++ GA + IT +G+ + + Y ++P K LT+
Sbjct: 225 IANGLINLIEDDALNGAIMKITASKGIHF-------QDYDISPLLVKAPLTS 269
>sptr|Q8MJY8|Q8MJY8 Prostaglandin dehydrogenase I.
Length = 266
Score = 136 bits (342), Expect = 7e-31
Identities = 90/269 (33%), Positives = 138/269 (51%), Gaps = 21/269 (7%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G ALVTG A GIG+A A +G V +VD++ E G + + + KF
Sbjct: 5 GKVALVTGAAQGIGRAFAEALLLKGAKVALVDWNLEAGVQCKAALDE---KFEPQ----K 57
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
++F++CDV++ L F K V + LDI +N AG+ N+ + W + +
Sbjct: 58 TLFIQCDVADQQQLRDTFRKVVDHFGRLDILVNNAGVNNE-----------KNWEKTLQI 106
Query: 584 NLVAVIDGTRIASQIMRNQK--KPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRS-- 751
NLV+VI GT + M Q + G+IIN+ S AGL P+ PVY A+K G+V FTRS
Sbjct: 107 NLVSVISGTYLGLDYMSKQNGGEGGIIINMSSLAGLMPVAQQPVYCASKHGIVGFTRSAA 166
Query: 752 -LSPLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSSGGYLEMED---------------- 880
+ L GVR+N +CP FV T + E + ++ ++ G Y+E +D
Sbjct: 167 LAANLMNSGVRLNAICPGFVNTAILESIEKE--ENMGQYIEYKDHIKDMIKYYGILDPPL 224
Query: 881 VVNGTFELIQDESKAGACLWITKRRGMEY 967
+ NG LI+D++ GA + IT +G+ +
Sbjct: 225 IANGLITLIEDDALNGAIMKITTSKGIHF 253
>sw|P15428|PGDH_HUMAN 15-hydroxyprostaglandin dehydrogenase [NAD(+)]
(EC 1.1.1.141) (PGDH).
Length = 266
Score = 136 bits (342), Expect = 7e-31
Identities = 88/269 (32%), Positives = 137/269 (50%), Gaps = 21/269 (7%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G ALVTG A GIG+A A +G V +VD++ E G + + ++
Sbjct: 5 GKVALVTGAAQGIGRAFAEALLLKGAKVALVDWNLEAGVQCKAALDEQ-------FEPQK 57
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
++F++CDV++ L F K V + LDI +N AG+ N+ + W + +
Sbjct: 58 TLFIQCDVADQQQLRDTFRKVVDHFGRLDILVNNAGVNNE-----------KNWEKTLQI 106
Query: 584 NLVAVIDGTRIASQIMRNQK--KPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRS-- 751
NLV+VI GT + M Q + G+IIN+ S AGL P+ PVY A+K G+V FTRS
Sbjct: 107 NLVSVISGTYLGLDYMSKQNGGEGGIIINMSSLAGLMPVAQQPVYCASKHGIVGFTRSAA 166
Query: 752 -LSPLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSSGGYLEMED---------------- 880
+ L GVR+N +CP FV T + E + ++ ++ G Y+E +D
Sbjct: 167 LAANLMNSGVRLNAICPGFVNTAILESIEKE--ENMGQYIEYKDHIKDMIKYYGILDPPL 224
Query: 881 VVNGTFELIQDESKAGACLWITKRRGMEY 967
+ NG LI+D++ GA + IT +G+ +
Sbjct: 225 IANGLITLIEDDALNGAIMKITTSKGIHF 253
>sptr|P70684|P70684 15-hydroxyprostaglandin dehydrogenase (NAD+) (EC
1.1.1.141).
Length = 265
Score = 135 bits (339), Expect = 2e-30
Identities = 88/267 (32%), Positives = 136/267 (50%), Gaps = 19/267 (7%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G ALVTG A GIG+A +G V +VD++ E G + + +E
Sbjct: 5 GKVALVTGAAQGIGRAFAEGLLHKGAKVALVDWNLEAGVKCKAALDEE-------FEPQK 57
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
++F++CDV++ + L F K V + LDI +N AG+ N+ + W + +
Sbjct: 58 TLFIQCDVADQEQLRDTFTKVVDYFGRLDILVNNAGVNNE-----------KNWEKTLQI 106
Query: 584 NLVAVIDGTRIASQIMRNQK--KPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSLS 757
NLV+VI GT + M Q + GVIIN+ S AGL P+ PVY A+K G++ FTRS +
Sbjct: 107 NLVSVISGTYLGLDYMSKQHGGEGGVIINMSSLAGLMPVAQQPVYCASKHGIIGFTRSAA 166
Query: 758 ---PLKRHGVRVNVLCPEFVQTNMAEQMSRK-----VIDSSG---------GYLEMEDVV 886
L GVR+N +CP FV T++ + + ++ I+ +G G L+ E +
Sbjct: 167 MARKLMNSGVRMNAICPGFVNTSILQSIEKEENMGPYIEYTGHIKDMMKCYGILDPEMIA 226
Query: 887 NGTFELIQDESKAGACLWITKRRGMEY 967
NG LI+D+ GA + IT G+ +
Sbjct: 227 NGLITLIEDDDLNGAIMKITTSNGIHF 253
>sptr|O97743|O97743 NAD+-dependent 15-hydoxyprostaglandin
dehydrogenase.
Length = 266
Score = 132 bits (333), Expect = 8e-30
Identities = 88/267 (32%), Positives = 135/267 (50%), Gaps = 19/267 (7%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G ALVTG A GIG+A A +G V +VD++ E G + + ++
Sbjct: 5 GKVALVTGAAQGIGRAFAEALLLKGAKVALVDWNLEAGVKCKAALDEQ-------FEPQK 57
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
++F++CDV++ L F K V + LDI +N AG+ N+ + W + +
Sbjct: 58 TLFIQCDVADQQQLRDTFRKVVDHFGKLDILVNNAGVNNE-----------KNWEKTLQI 106
Query: 584 NLVAVIDGTRIASQIMRNQK--KPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRS-- 751
NLV+VI GT + M Q + G+IIN+ S AGL P+ PVY A+K G+V FTRS
Sbjct: 107 NLVSVISGTYLGLDYMSKQNGGEGGIIINMSSLAGLMPVAQQPVYCASKHGIVGFTRSAA 166
Query: 752 -LSPLKRHGVRVNVLCPEFVQTNMAEQMSR-----KVIDSSG---------GYLEMEDVV 886
+ L GVR+N +CP FV T + + + + K I+ G G L+ +
Sbjct: 167 MAANLMNSGVRLNAICPGFVDTPILKSIEKEENMGKYIEYMGPIKDMMKYYGILDPSMIA 226
Query: 887 NGTFELIQDESKAGACLWITKRRGMEY 967
NG LI+D++ GA + IT +G+ +
Sbjct: 227 NGLITLIEDDALNGAIMKITTSKGIHF 253
>sptr|Q61106|Q61106 NAD(+)-dependent 15-hydroxyprostaglandin
dehydrogenase.
Length = 266
Score = 131 bits (329), Expect = 2e-29
Identities = 86/269 (31%), Positives = 136/269 (50%), Gaps = 21/269 (7%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G ALVTG A GIGKA A G V +VD++ E G + + ++
Sbjct: 5 GKVALVTGAAQGIGKAFAEALLLHGAKVALVDWNLEAGVKCKAALDEQ-------FEPQK 57
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
++FV+CDV++ L F K V + LDI +N AG+ N+ + W + +
Sbjct: 58 TLFVQCDVADQKQLRDTFRKVVDHFGRLDILVNNAGVNNE-----------KNWEQTLQI 106
Query: 584 NLVAVIDGTRIASQIMRNQK--KPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRS-- 751
NLV+VI GT + M Q + G+IIN+ S AGL P+ PVY ++K G++ FTRS
Sbjct: 107 NLVSVIGGTYLGLDYMSKQNGGEGGIIINMSSLAGLMPVAQQPVYCSSKHGIIGFTRSAA 166
Query: 752 -LSPLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSSGGYLEMED---------------- 880
+ L + GVR+NV+CP FV T + E + ++ ++ G Y+ +D
Sbjct: 167 MAANLMKSGVRLNVICPGFVDTPILESIEKE--ENMGQYIGYKDQIKAMMKFYGVLHPSA 224
Query: 881 VVNGTFELIQDESKAGACLWITKRRGMEY 967
+ NG +LI+D++ G + I+ G+ +
Sbjct: 225 IANGLIDLIEDDALNGPIMKISASEGIHF 253
>sptr|Q9MZF9|Q9MZF9 Prostaglandin dehydrogenase (Fragment).
Length = 228
Score = 125 bits (314), Expect = 1e-27
Identities = 78/224 (34%), Positives = 119/224 (53%), Gaps = 5/224 (2%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G ALVTG A GIG+A A +G V +VD++ E G + + ++
Sbjct: 5 GKVALVTGAAQGIGRAFAEALLLKGAKVALVDWNLEAGVQCKAALDEQ-------FEPQK 57
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
++F++CDV++ L F K V + LDI +N AG+ N+ + W + +
Sbjct: 58 TLFIQCDVADQQQLRDTFRKVVDHFGRLDILVNNAGVNNE-----------KNWEKTLQI 106
Query: 584 NLVAVIDGTRIASQIMRNQK--KPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRS-- 751
NLV+VI GT + M Q + G+IIN+ S AGL P+ PVY A+K G+V FTRS
Sbjct: 107 NLVSVISGTYLGLDYMSKQNGGEGGIIINMSSLAGLMPVAQQPVYCASKHGIVGFTRSAA 166
Query: 752 -LSPLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSSGGYLEMED 880
+ L GVR+N +CP FV T + E + ++ ++ G Y+E +D
Sbjct: 167 LAANLMNSGVRLNAICPGFVNTAILESIEKE--ENMGQYIEYKD 208
>sptr|Q7ZTX3|Q7ZTX3 Similar to hydroxyprostaglandin dehydrogenase
15-(NAD).
Length = 270
Score = 123 bits (309), Expect = 5e-27
Identities = 83/264 (31%), Positives = 135/264 (51%), Gaps = 19/264 (7%)
Frame = +2
Query: 209 MELKPGMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGD 388
M+LK + A+VTG A G+G++ + G V ++D ++ G E+ T + KE +
Sbjct: 1 MDLKDKV-AVVTGAAQGLGRSFVEILMKNGSKVALIDVNKSLGEELKTTLNKEYGPNRAE 59
Query: 389 LRIPSSIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWR 568
F DVS+ ++ +K V+ + +DI N AGI N+ + W
Sbjct: 60 -------FYTADVSSEEDFKGVLKKIVEQFGQIDIMCNNAGIINE-----------KHWE 101
Query: 569 HAVNVNLVAVIDGTRIASQIMRNQK--KPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMF 742
+ +NL V+ GT +A + M+ + GVI+N+ S AGL P + P+Y+A K GVV F
Sbjct: 102 KTIAINLGGVVRGTYLALEYMKKENGGSGGVIVNVASMAGLGPFPVAPIYTATKHGVVGF 161
Query: 743 TRSL---SPLKRHGVRVNVLCPEFVQTNMAEQMS--------------RKVIDSSGGYLE 871
+R++ S L +GVR+NVLCP FV+T++ ++ +++ S G LE
Sbjct: 162 SRAMAVVSKLSNYGVRINVLCPWFVKTSLLSLLNSEEHTGSFSQMKEITEMLMESEGCLE 221
Query: 872 MEDVVNGTFELIQDESKAGACLWI 943
++ V L++DESK G L I
Sbjct: 222 VDVVAKAFLVLVKDESKDGEALMI 245
>sptrnew|AAR84629|AAR84629 Putative alcohol dehydrogenase.
Length = 266
Score = 108 bits (271), Expect = 1e-22
Identities = 84/219 (38%), Positives = 108/219 (49%), Gaps = 7/219 (3%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGL-FVTVVDFSEENGREVATLVQKENSKFHGDLRIP 400
G ALVTGGA+GIG A R G+ V V D E G V++ N +F
Sbjct: 5 GKYALVTGGATGIGLAYVKELLRHGVQAVAVADLDERKGDNS---VKQLNDEFGAG---- 57
Query: 401 SSIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVN 580
+IF+KCDV+N D+L A F K T+ GLDI IN AGI DD+ W +
Sbjct: 58 KAIFIKCDVTNKDDLEATFRKAAATFKGLDIVINNAGI------LDDS-----RWETEIA 106
Query: 581 VNLVAVIDGTRIASQIMRNQK--KPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSL 754
+N+ AV GT +A +M K + GV++NI S GL M PVY A K VV +RS
Sbjct: 107 INVTAVARGTWLAFDLMGKNKGGRGGVVVNIASILGLQAMAGCPVYVATKHAVVGLSRSF 166
Query: 755 S---PLKRHGVRVNVLCPEFVQTNM-AEQMSRKVIDSSG 859
R GVRV +CP T + +E R+ D G
Sbjct: 167 GMPYHFDRTGVRVLTMCPGVTDTPLISEAHHRQFTDDLG 205
>sptr|Q12998|Q12998 15-hydroxy prostaglandin dehydrogenase.
Length = 178
Score = 107 bits (266), Expect = 5e-22
Identities = 65/181 (35%), Positives = 95/181 (52%), Gaps = 2/181 (1%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G ALVTG A GIG+A A +G V +VD++ E G + + ++
Sbjct: 5 GKVALVTGAAQGIGRAFAEALLLKGAKVALVDWNLEAGVQCKAALDEQ-------FEPQK 57
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
++F++CDV++ L F K V + LDI +N AG+ NK + W + +
Sbjct: 58 TLFIQCDVADQQQLRDTFRKVVDHFGRLDILVNNAGVNNK-----------KNWEKTLQI 106
Query: 584 NLVAVIDGTRIASQIMRNQK--KPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSLS 757
NLV+VI GT + M Q + G+IIN+ S AGL P+ PVY A+K G+V FTRS +
Sbjct: 107 NLVSVISGTYLGLDYMSKQNGGEGGIIINMSSLAGLMPVAQQPVYCASKHGIVGFTRSAA 166
Query: 758 P 760
P
Sbjct: 167 P 167
>sptr|Q19246|Q19246 Hypothetical protein F09E10.3.
Length = 248
Score = 103 bits (256), Expect = 7e-21
Identities = 83/251 (33%), Positives = 114/251 (45%), Gaps = 10/251 (3%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G A+VTGGASGIGKA+ A+ G V V D ++G AT S+ H
Sbjct: 7 GRIAVVTGGASGIGKAISQTLAKHGARVVVADL--DSGNAAATAKALPASQSHSSF---- 60
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
CDVSNAD++ E HV++ I +NCAGI + + + W + V
Sbjct: 61 ----ACDVSNADSVKGLSE-HVKSLGTPSILVNCAGITKDSTLLKMKQE---QWDSVIKV 112
Query: 584 NLVAVIDGTR-IASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSLS- 757
NL V ++ + N P IIN+ S G F Y+A K GV+ FT+S +
Sbjct: 113 NLTGVFHVSQAFVKASVDNNNHPLSIINVSSIVGKMGNFGQTNYAATKAGVIGFTKSAAK 172
Query: 758 PLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDS------SGGYLEMEDVVNGTFELIQDES 919
L + VRVN + P F++T M E M V+ G E E++ N L D S
Sbjct: 173 ELAKKNVRVNAVLPGFIKTPMTEAMPPTVLAEICKGIPMGRMGEAEEIANSVLYLASDLS 232
Query: 920 K--AGACLWIT 946
GA L +T
Sbjct: 233 SYVTGATLEVT 243
>sptrnew|AAS04725|AAS04725 FabG2_2.
Length = 249
Score = 101 bits (252), Expect = 2e-20
Identities = 70/224 (31%), Positives = 114/224 (50%), Gaps = 1/224 (0%)
Frame = +2
Query: 206 EMELKPGMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHG 385
++ L G +A++TGGA G+G A+ F G V + D + E + A + G
Sbjct: 3 QVSLLSGQTAVITGGAQGLGFAIAERFVAEGARVVLGDVNLEATQTAA-------KQLGG 55
Query: 386 DLRIPSSIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTW 565
D ++ V+CDV+ + + + V+ + GLDI +N AGI + T + +
Sbjct: 56 D---QVALAVRCDVTKSSEVETLIQTAVERFGGLDIMVNNAGITRDATMRKMTEE---QF 109
Query: 566 RHAVNVNLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFT 745
+ V+L +GTR+A+ IMR K+ G IIN+ S +G M YSAAK G+V T
Sbjct: 110 DQVIAVHLKGTWNGTRLAAAIMRENKR-GAIINMSSVSGKVGMVGQTNYSAAKAGIVGMT 168
Query: 746 RSLS-PLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSSGGYLEM 874
++ + L GVRVN + P +++ M E M +++ DS + M
Sbjct: 169 KAAAKELAYLGVRVNAIAPGLIRSAMTEAMPQRIWDSKVAEVSM 212
>sptr|Q94G10|Q94G10 CTA.
Length = 271
Score = 100 bits (250), Expect = 3e-20
Identities = 76/245 (31%), Positives = 115/245 (46%), Gaps = 12/245 (4%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G A++TGGA GIG+ F + G V + D + G+ + DL S
Sbjct: 15 GKVAVITGGARGIGEQTAKLFFKHGAKVVIADIQDHLGQTLCK-----------DLGQSS 63
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
S+FV CDV+ ++ + V Y LDI +N AG+ ++ +D D T++ VNV
Sbjct: 64 SVFVHCDVTKEKDVETAVDTAVSKYGKLDIMLNNAGVFEESPNFDILKDDPLTFQRVVNV 123
Query: 584 NLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSLS-P 760
NLV GTR A+++M+ + G I+ S + Y+++K GV+ R+ +
Sbjct: 124 NLVGASLGTRHAARVMKPAGR-GSIVTTASICSVIGGIGTHAYTSSKHGVLGLMRNAAVD 182
Query: 761 LKRHGVRVNVLCPEFVQTNMAEQMSRKVID-----------SSGGYLEMEDVVNGTFELI 907
L R+G+RVN + P V T M ++ KV D +G L EDV L
Sbjct: 183 LGRYGIRVNCVSPNVVPTEMGRKLF-KVKDGGEFPSFYWSLKNGDILREEDVGEAVVYLG 241
Query: 908 QDESK 922
DESK
Sbjct: 242 SDESK 246
>sptr|Q94G09|Q94G09 Sex determination protein.
Length = 271
Score = 100 bits (249), Expect = 4e-20
Identities = 75/245 (30%), Positives = 115/245 (46%), Gaps = 12/245 (4%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G A++TGGA GIG+ F + G V + D + G+ + DL S
Sbjct: 15 GKVAVITGGARGIGEQTAKLFFKHGAKVVIADIQDHLGQTLCK-----------DLGQSS 63
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
S+FV CDV+ ++ + V Y LDI +N AG+ ++ +D D T++ VNV
Sbjct: 64 SVFVHCDVTKEKDVETAVDTAVSKYGKLDIMLNNAGVFEESPNFDFLKDDPLTFQRVVNV 123
Query: 584 NLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSLS-P 760
NLV GT+ A+++M+ + G I+ S + Y+++K GV+ R+ +
Sbjct: 124 NLVGAFLGTKHAARVMKPAGR-GSIVTTASICSVIGGIGTHAYTSSKHGVLGLMRNAAVD 182
Query: 761 LKRHGVRVNVLCPEFVQTNMAEQMSRKVID-----------SSGGYLEMEDVVNGTFELI 907
L R+G+RVN + P V T M ++ KV D +G L EDV L
Sbjct: 183 LGRYGIRVNCVSPNVVPTEMGRKLF-KVKDGGEFPSFYWSLKNGDILREEDVGEAVVYLG 241
Query: 908 QDESK 922
DESK
Sbjct: 242 SDESK 246
>sptr|Q8IQN8|Q8IQN8 CG4899-PB.
Length = 277
Score = 100 bits (249), Expect = 4e-20
Identities = 73/233 (31%), Positives = 111/233 (47%), Gaps = 5/233 (2%)
Frame = +2
Query: 164 PQQLLDRSTHPCAGEMELKPGMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGRE 343
PQ + +T +M + G +A+VTGGA GIG + G V ++D +
Sbjct: 3 PQGIPPTTTRISQTKMSFR-GKNAVVTGGAGGIGLQVSKQLLAAGAAVAIIDLQDN---- 57
Query: 344 VATLVQKENSKFHGDLRIPSSIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANK 523
+E K S + +K DV+N + A +E+ +T+ +DI +N AGI N
Sbjct: 58 -----LEEFVKLRAAHPTQSVMIIKMDVANKKGVEATYEEIAKTFGNIDIVVNVAGIFND 112
Query: 524 TLVYDDTSDGTRTWRHAVNVNLVAVIDGTRIASQIMR--NQKKPGVIINIGSAAGLYPMF 697
D RT + VNL +I+ T A M N K G+++N+ S GL PMF
Sbjct: 113 -------KDVQRT----LLVNLGGIINSTLSALPYMGKDNGGKGGIVVNMSSVVGLDPMF 161
Query: 698 LDPVYSAAKGGVVMFTRSLSPLK---RHGVRVNVLCPEFVQTNMAEQMSRKVI 847
+ PVY A K G++ FTR L+ K R G++ +CP T+M + K+I
Sbjct: 162 IIPVYGATKAGIINFTRCLANEKYYQRSGIKFVTVCPGATMTDMFTNFTEKII 214
>sptr|Q7V2M2|Q7V2M2 3-oxoacyl-[acyl-carrier protein] reductase (EC
1.1.1.100).
Length = 249
Score = 99.8 bits (247), Expect = 7e-20
Identities = 67/210 (31%), Positives = 107/210 (50%), Gaps = 2/210 (0%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTV-VDFSEENGREVATLVQKENSKFHGDLRIP 400
G AL+TG + GIGK + + + G V + S+E EV L+++ K H
Sbjct: 9 GKVALITGASRGIGKEIALELSNLGAKVIINYSSSDEKAEEVVNLIKESGGKVHK----- 63
Query: 401 SSIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVN 580
+K DVS ++++ FE+ ++ +DI +N AGI L+ S+ W +N
Sbjct: 64 ----LKFDVSKEESVSKAFEEIIKINGAIDILVNNAGITRDGLLMRMKSE---QWDDVLN 116
Query: 581 VNLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSLS- 757
NL V T+ AS+ M +K+ G IINI S G+ YSAAK GV+ FT++ +
Sbjct: 117 TNLKGVFLCTKYASKFM-IKKRSGKIINISSIVGIIGNPGQANYSAAKAGVIGFTKTCAK 175
Query: 758 PLKRHGVRVNVLCPEFVQTNMAEQMSRKVI 847
G+ VN + P F++T M E+++ + I
Sbjct: 176 EFASRGINVNAIAPGFIETEMTEKLNNEEI 205
>sptr|Q81EL1|Q81EL1 3-oxoacyl-[acyl-carrier protein] reductase (EC
1.1.1.100).
Length = 242
Score = 99.4 bits (246), Expect = 1e-19
Identities = 75/207 (36%), Positives = 104/207 (50%), Gaps = 3/207 (1%)
Frame = +2
Query: 212 ELKPGMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDL 391
EL G +AL+TG GIG+A+ IA A+ G+ V ++ SEEN + VA V+ E K
Sbjct: 6 ELLQGKNALITGAGRGIGRAVAIALAKEGVNVGLLARSEENLKAVAKEVEAEGVK----- 60
Query: 392 RIPSSIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIA--NKTLVYDDTSDGTRTW 565
++ DVS+ + + E +DI IN AGI+ K L D W
Sbjct: 61 ----AVIATADVSSYEEVTTAIETLKNGLGSIDILINNAGISKFGKFLELD-----VADW 111
Query: 566 RHAVNVNLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFT 745
+ VNL+ V TR A M Q+ G IINI S AG + YSA+K GV+ T
Sbjct: 112 EKIIQVNLMGVYYATRAALPSMIEQQS-GDIINISSTAGQKGAPVTSAYSASKFGVLGLT 170
Query: 746 RSLS-PLKRHGVRVNVLCPEFVQTNMA 823
SL+ +++H +RV L P V T+MA
Sbjct: 171 ESLAMEVRKHNIRVTALTPSTVATDMA 197
>sptr|Q81RT0|Q81RT0 Oxidoreductase, short-chain
dehydrogenase/reductase family.
Length = 242
Score = 99.4 bits (246), Expect = 1e-19
Identities = 75/207 (36%), Positives = 104/207 (50%), Gaps = 3/207 (1%)
Frame = +2
Query: 212 ELKPGMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDL 391
EL G +AL+TG GIG+A+ IA A+ G+ V ++ SEEN + VA V+ E K
Sbjct: 6 ELLQGKNALITGAGRGIGRAVAIALAKEGVNVGLLARSEENLKAVAKEVEAEGVK----- 60
Query: 392 RIPSSIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIA--NKTLVYDDTSDGTRTW 565
++ DVS+ + + E +DI IN AGI+ K L D W
Sbjct: 61 ----AVIATADVSSYEEVTTAIETLKNGLGSIDILINNAGISKFGKFLELD-----VADW 111
Query: 566 RHAVNVNLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFT 745
+ VNL+ V TR A M Q+ G IINI S AG + YSA+K GV+ T
Sbjct: 112 EKIIQVNLMGVYYATRAALPSMIEQQS-GDIINISSTAGQKGAPVTSAYSASKFGVLGLT 170
Query: 746 RSLS-PLKRHGVRVNVLCPEFVQTNMA 823
SL+ +++H +RV L P V T+MA
Sbjct: 171 ESLAMEVRKHNIRVTALTPSTVATDMA 197
>sptr|Q82FK0|Q82FK0 Putative oxidoreductase.
Length = 584
Score = 99.4 bits (246), Expect = 1e-19
Identities = 69/200 (34%), Positives = 96/200 (48%), Gaps = 1/200 (0%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G LVTG SGIG+A AFA G V VD + E A E S+ G P
Sbjct: 316 GQLVLVTGAGSGIGRATAYAFAEAGARVVAVDRNAEGAARTA-----EMSRLIG---APD 367
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
+ DVS+ + EK Y +D+ +N AGI +D T++ WR ++V
Sbjct: 368 AWAETVDVSDEQAMEKLAEKVATEYGVVDVLVNNAGIGLSGSFFDTTAED---WRKVLDV 424
Query: 584 NLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSL-SP 760
NL VI G R+ + M + + G I+N SAA P P YS +K V+M + L +
Sbjct: 425 NLWGVIHGCRLFGKQMAERGQGGHIVNTASAAAYQPSKALPAYSTSKAAVLMLSECLRAE 484
Query: 761 LKRHGVRVNVLCPEFVQTNM 820
L G+ V+ +CP FV TN+
Sbjct: 485 LAGQGIGVSAICPGFVHTNI 504
>sw|Q11020|YD50_MYCTU Putative oxidoreductase Rv1350/MT1393/Mb1385
(EC 1.-.-.-).
Length = 247
Score = 99.0 bits (245), Expect = 1e-19
Identities = 67/208 (32%), Positives = 110/208 (52%), Gaps = 1/208 (0%)
Frame = +2
Query: 230 SALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPSSI 409
+A++TGGA G+G A+ F G V + D + E EVA + GD ++
Sbjct: 9 TAVITGGAQGLGLAIGQRFVAEGARVVLGDVNLE-ATEVAA------KRLGGD---DVAL 58
Query: 410 FVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNVNL 589
V+CDV+ AD++ V+ + GLD+ +N AGI + T + + + V+L
Sbjct: 59 AVRCDVTQADDVDILIRTAVERFGGLDVMVNNAGITRDATMRTMTEE---QFDQVIAVHL 115
Query: 590 VAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSLSPLKR 769
+GTR+A+ IMR +K+ G I+N+ S +G M YSAAK G+V T++ +
Sbjct: 116 KGTWNGTRLAAAIMRERKR-GAIVNMSSVSGKVGMVGQTNYSAAKAGIVGMTKAAAKELA 174
Query: 770 H-GVRVNVLCPEFVQTNMAEQMSRKVID 850
H G+RVN + P +++ M E M +++ D
Sbjct: 175 HLGIRVNAIAPGLIRSAMTEAMPQRIWD 202
>sptr|Q89UC3|Q89UC3 D-beta-hydroxybutyrate dehydrogenase.
Length = 249
Score = 99.0 bits (245), Expect = 1e-19
Identities = 79/235 (33%), Positives = 113/235 (48%), Gaps = 8/235 (3%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G++ LVTG ASG+G+A FA G V V D+ E+ VA KE + G S
Sbjct: 3 GLTVLVTGAASGMGRATARVFAAEGANVAVTDYDEQGAISVA----KEIAASGG-----S 53
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
+ K DV++ + + GLDI +N AGI+ + + D+T + W + V
Sbjct: 54 AKAWKLDVADGGEIKRVVHDVAAHFGGLDIVVNNAGISVRVAIDDETYED--AWARGIAV 111
Query: 584 NLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSLS-P 760
L A R A +R K P I+NI S L L YSAAKGGV TRSL+
Sbjct: 112 MLTAHPRIIRAALPHLRKSKSPR-IVNIASTEALGATALHSPYSAAKGGVASLTRSLAVE 170
Query: 761 LKRHGVRVNVLCPEFVQTNMAEQMSR-------KVIDSSGGYLEMEDVVNGTFEL 904
L R G+ VN +CP ++T + +++S K + G Y + E+V + T L
Sbjct: 171 LGREGITVNCICPGPIRTAITDRISEEHKTIYAKRRTALGRYGDPEEVAHMTLSL 225
>sptr|Q8EXX5|Q8EXX5 3-ketoacyl-acyl carrier protein reductase (EC
1.1.1.100).
Length = 254
Score = 98.6 bits (244), Expect = 2e-19
Identities = 61/208 (29%), Positives = 102/208 (49%), Gaps = 1/208 (0%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G +A+VTG A GIGK+ + A+ G + + D +EE+ + A + K+
Sbjct: 6 GKNAVVTGAARGIGKSTALTLAKAGANLVIADLNEESSKATADEISKQTGV--------K 57
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
+I + +VS+ D+ A + V + +DI +N AGI TL+ + W + V
Sbjct: 58 AIGIGTNVSDVDSAAKAIQACVDEFGSVDILVNNAGITKDTLLMRMKKE---QWDAVIAV 114
Query: 584 NLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSLS-P 760
NL + T+ A + M G IIN+ S AG+ YSA+K GV+ FT++++
Sbjct: 115 NLTGTFNCTQAAIKFMMKNPNGGSIINLSSIAGVNGNIGQTNYSASKAGVIGFTKAVALE 174
Query: 761 LKRHGVRVNVLCPEFVQTNMAEQMSRKV 844
+ VR N + P F+ T M + + K+
Sbjct: 175 MASRKVRCNAIAPGFIATEMTDAIPEKI 202
>sptr|Q8PS57|Q8PS57 Putative ketoreductase.
Length = 236
Score = 98.2 bits (243), Expect = 2e-19
Identities = 71/221 (32%), Positives = 116/221 (52%), Gaps = 12/221 (5%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G +A+VTGG GIG+A+C+A AR G + + +E++ RE A +V+KE K L + +
Sbjct: 5 GQTAVVTGGGKGIGRAICLALAREGADIVIAARTEKDIRETARMVEKEGRK---ALPVST 61
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVNV 583
I V+ DV N + V + +DI +N AG+A + + + + T + + ++
Sbjct: 62 DIRVEEDVEN------MISEAVDAFGRIDILVNNAGVAYRKYMVETS---TEEYDNIMDT 112
Query: 584 NLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSLSPL 763
NL + T+ A + ++ G IINI S AG + + +YSA+K V+ FT S++
Sbjct: 113 NLKGMFFCTKYALPYLL-KRGEGRIINISSGAGKHGIPKLSIYSASKFAVIGFTESIAYE 171
Query: 764 KRHGVRVNVLCPEFVQTNM------------AEQMSRKVID 850
GVRV +CP V T+M E +SRKV++
Sbjct: 172 IGGGVRVYAVCPSSVDTDMYRSLHSDKPALKPEDVSRKVLE 212
>sw|Q9X248|FABG_THEMA 3-oxoacyl-[acyl-carrier protein] reductase (EC
1.1.1.100) (3-ketoacyl- acyl carrier protein reductase).
Length = 246
Score = 97.8 bits (242), Expect = 3e-19
Identities = 73/240 (30%), Positives = 117/240 (48%), Gaps = 8/240 (3%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G L+TG ASGIGKA + FA+ G V D S+EN + +LV++ +P
Sbjct: 5 GKVCLITGAASGIGKATTLLFAQEGATVIAGDISKEN---LDSLVKEAEG-------LPG 54
Query: 404 SIF-VKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTRTWRHAVN 580
+ +V++ D + EK VQ Y +D+ +N AGI L+ + W +N
Sbjct: 55 KVDPYVLNVTDRDQIKEVVEKVVQKYGRIDVLVNNAGITRDALLVRMKEE---DWDAVIN 111
Query: 581 VNLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRS-LS 757
VNL V + T++ M Q+ G I+N+ S G+Y Y+A+K GV+ T++
Sbjct: 112 VNLKGVFNVTQMVVPYMIKQRN-GSIVNVSSVVGIYGNPGQTNYAASKAGVIGMTKTWAK 170
Query: 758 PLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSS------GGYLEMEDVVNGTFELIQDES 919
L +RVN + P F++T M E++ K +++ G + + E+V L DES
Sbjct: 171 ELAGRNIRVNAVAPGFIETPMTEKLPEKARETALSRIPLGRFGKPEEVAQVILFLASDES 230
>sptr|Q89LE8|Q89LE8 Short chain dehydrogenase.
Length = 254
Score = 97.4 bits (241), Expect = 4e-19
Identities = 79/242 (32%), Positives = 112/242 (46%), Gaps = 2/242 (0%)
Frame = +2
Query: 224 GMSALVTGGASGIGKALCIAFARRGLFVTVVDFSEENGREVATLVQKENSKFHGDLRIPS 403
G ALVTG ASG+G A AFA G V + D++E+ R A + K
Sbjct: 7 GQVALVTGAASGLGLATAKAFAESGASVALADWNEDAVRAAAGELVTRGHK--------- 57
Query: 404 SIFVKCDVSNADNLAACFEKHVQTYNGLDICINCAGIANKTLVYDDTSDGTR-TWRHAVN 580
++ ++CDVSN + A + V + LD N AG+ N V +T+D TR + +
Sbjct: 58 AVAIRCDVSNDAQVEAMVAQTVSVFGRLDAAYNNAGVQN---VLAETADTTREDYDRVMG 114
Query: 581 VNLVAVIDGTRIASQIMRNQKKPGVIINIGSAAGLYPMFLDPVYSAAKGGVVMFTRSLS- 757
+NL + Q MR Q G I+N S GL +Y AAK GV+ FT+S +
Sbjct: 115 INLRGEWSCMKFELQQMRKQGS-GAIVNCSSLGGLVGGAERGIYHAAKHGVLGFTKSAAL 173
Query: 758 PLKRHGVRVNVLCPEFVQTNMAEQMSRKVIDSSGGYLEMEDVVNGTFELIQDESKAGACL 937
GVR+N +CP + T MA+QM V G L + + + E A A L
Sbjct: 174 EYAARGVRINAICPGLIWTPMADQM---VAAGQGDALRALEKSVPMGRVGRPEEIASAVL 230
Query: 938 WI 943
W+
Sbjct: 231 WL 232
Database: swall
Posted date: Feb 26, 2004 3:00 PM
Number of letters in database: 439,479,560
Number of sequences in database: 1,381,838
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 857,295,454
Number of Sequences: 1381838
Number of extensions: 18160798
Number of successful extensions: 65744
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 59738
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 62908
length of database: 439,479,560
effective HSP length: 125
effective length of database: 266,749,810
effective search space used: 70688699650
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

