BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= QAJ4a11.xg.3.2
(2249 letters)
Database: swall
1,381,838 sequences; 439,479,560 total letters
Searching..................................................done
Score E
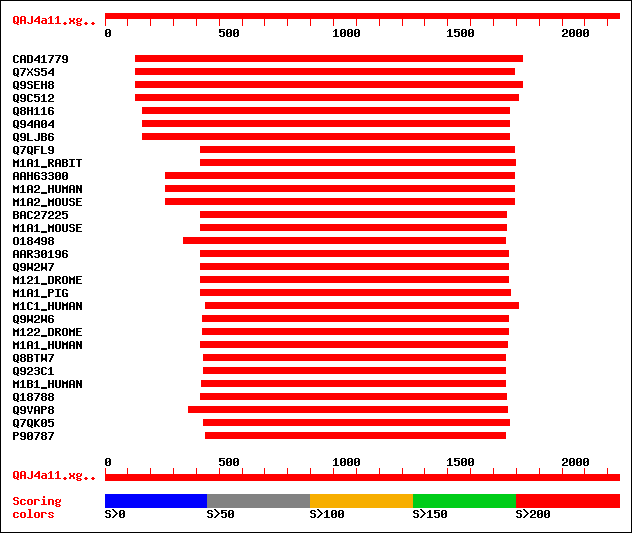
Sequences producing significant alignments: (bits) Value
sptrnew|CAD41779|CAD41779 OSJNBa0035M09.7 protein. 1022 0.0
sptr|Q7XS54|Q7XS54 OSJNBa0035M09.10 protein. 857 0.0
sptr|Q9SEH8|Q9SEH8 Mannosyl-oligosaccharide 1,2-alpha-mannosidas... 821 0.0
sptr|Q9C512|Q9C512 Mannosyl-oligosaccharide 1,2-alpha-mannosidas... 810 0.0
sptr|Q8H116|Q8H116 Putative mannosidase. 789 0.0
sptr|Q94A04|Q94A04 Putative alpha 1,2-mannosidase. 787 0.0
sptr|Q9LJB6|Q9LJB6 Alpha 1,2-mannosidase-like protein. 785 0.0
sptr|Q7QFL9|Q7QFL9 AgCP13734 (Fragment). 402 e-110
sw|P45701|M1A1_RABIT Mannosyl-oligosaccharide 1,2-alpha-mannosid... 399 e-109
sptrnew|AAH63300|AAH63300 MAN1A2 protein. 396 e-109
sw|O60476|M1A2_HUMAN Mannosyl-oligosaccharide 1,2-alpha-mannosid... 396 e-109
sw|P39098|M1A2_MOUSE Mannosyl-oligosaccharide 1,2-alpha-mannosid... 395 e-108
sptrnew|BAC27225|BAC27225 Adult male thymus cDNA, RIKEN full-len... 391 e-107
sw|P45700|M1A1_MOUSE Mannosyl-oligosaccharide 1,2-alpha-mannosid... 391 e-107
sptr|O18498|O18498 Alpha 1,2-mannosidase. 388 e-106
sptrnew|AAR30196|AAR30196 RE43942p. 386 e-106
sptr|Q9W2W7|Q9W2W7 Alpha-MAN-I protein. 386 e-106
sw|P53624|M121_DROME Mannosyl-oligosaccharide alpha-1,2-mannosid... 386 e-106
sw|O02773|M1A1_PIG Mannosyl-oligosaccharide 1,2-alpha-mannosidas... 382 e-104
sw|Q9NR34|M1C1_HUMAN Mannosyl-oligosaccharide 1,2-alpha-mannosid... 382 e-104
sptr|Q9W2W6|Q9W2W6 CG32684 protein. 380 e-104
sw|P53625|M122_DROME Mannosyl-oligosaccharide alpha-1,2-mannosid... 380 e-104
sw|P33908|M1A1_HUMAN Mannosyl-oligosaccharide 1,2-alpha-mannosid... 378 e-103
sptr|Q8BTW7|Q8BTW7 Similar to alpha 1 (Fragment). 376 e-103
sptr|Q923C1|Q923C1 Similar to alpha 1,2-mannosidase (Fragment). 376 e-103
sw|Q9UKM7|M1B1_HUMAN Endoplasmic reticulum mannosyl-oligosacchar... 374 e-102
sptr|Q18788|Q18788 C52E4.5 protein. 370 e-101
sptr|Q9VAP8|Q9VAP8 CG11874 protein (SD05769P). 369 e-101
sptr|Q7QK05|Q7QK05 AgCP2314 (Fragment). 362 2e-98
sptr|P90787|P90787 D2030.1 protein. 339 1e-91
>sptrnew|CAD41779|CAD41779 OSJNBa0035M09.7 protein.
Length = 572
Score = 1022 bits (2643), Expect = 0.0
Identities = 506/574 (88%), Positives = 524/574 (91%), Gaps = 9/574 (1%)
Frame = +3
Query: 132 MARRXXXXXX--GTWRYLNPAYYLKRPKRXXXXXXXXXXXXXXXWDRQSLVSEYESEISR 305
MARR G WRYLNPAYYLKRPKR WDRQSLV EYESEISR
Sbjct: 1 MARRSSSSSSSSGAWRYLNPAYYLKRPKRLALLFFVFVAATFAFWDRQSLVREYESEISR 60
Query: 306 LEDDINRLHDQLRKAGVNLDENPVVSNKNSRKDLVVEIDPVNNEKREKVKEAMLHAWNSY 485
L+D +N+L DQLRKAGV+LDENP +K SR+ LV EIDP+NNE+REKVKEAM HAWNSY
Sbjct: 61 LDDKVNQLRDQLRKAGVHLDENPT-GDKVSREKLV-EIDPINNERREKVKEAMAHAWNSY 118
Query: 486 VKYAWGMDELQPQSKNGINSFGGLGATLVDSLDTLYIMGLKDEFQKARDWVAESLDFDKD 665
VKYAWGMDELQPQSKNG+NSFGGLGATLVDSLDTLYIMGLKDEFQ+ARDWVA+SL FDKD
Sbjct: 119 VKYAWGMDELQPQSKNGVNSFGGLGATLVDSLDTLYIMGLKDEFQRARDWVADSLSFDKD 178
Query: 666 YDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDITDRLLPAWDTTSGIPYNRINLAHGR 845
YDASVFETTIRVVGGLLS+YDLSGDKVFL+KAKDITDRLLPAWDT SGIPYNRINLAHGR
Sbjct: 179 YDASVFETTIRVVGGLLSAYDLSGDKVFLEKAKDITDRLLPAWDTPSGIPYNRINLAHGR 238
Query: 846 AHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQKAENVIKQLQKIYPSDGLLPIYI 1025
AHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQKAENVI+QLQKIYPSDGLLPIYI
Sbjct: 239 AHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQKAENVIRQLQKIYPSDGLLPIYI 298
Query: 1026 NPHSGTASYSTITFGAMGDSFYEYLLKVWIQGNKTEHVKHYRQMWETSMEGLLSLTKKTT 1205
NPHSGTASYSTITFGAMGDSFYEYLLKVW+QGNKTEHVKHYRQMWETSMEGLLSLTKKTT
Sbjct: 299 NPHSGTASYSTITFGAMGDSFYEYLLKVWVQGNKTEHVKHYRQMWETSMEGLLSLTKKTT 358
Query: 1206 PSNYYYICEKNGGSLSDKMDELACFAPGMLALGASGY-GPEKSEQIMNLAKELARTCYNF 1382
PSNYYYICEKNGGSLSDKMDELACFAPGMLALGASGY EK+E+IMNLAKELARTCYNF
Sbjct: 359 PSNYYYICEKNGGSLSDKMDELACFAPGMLALGASGYEETEKAEEIMNLAKELARTCYNF 418
Query: 1383 YQTTPTKLAGENYFFHTGQDMGVGTSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQA 1562
YQTTPTKLAGENYFFHTGQDM VGTSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQA
Sbjct: 419 YQTTPTKLAGENYFFHTGQDMNVGTSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQA 478
Query: 1563 FEKNSRIESGYVGLRDVNTGEKDNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAH 1742
FEKNSRIESGYVGLRDVNTGEKDNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAH
Sbjct: 479 FEKNSRIESGYVGLRDVNTGEKDNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAH 538
Query: 1743 PLRIVPVNDNK-----GIGTP-VRQFGRKQGKPE 1826
PLRIVP+NDN GI TP VR FGRKQGK E
Sbjct: 539 PLRIVPLNDNSKAHSVGIATPTVRPFGRKQGKQE 572
>sptr|Q7XS54|Q7XS54 OSJNBa0035M09.10 protein.
Length = 543
Score = 857 bits (2215), Expect = 0.0
Identities = 440/561 (78%), Positives = 459/561 (81%), Gaps = 8/561 (1%)
Frame = +3
Query: 132 MARRXXXXXX--GTWRYLNPAYYLKRPKRXXXXXXXXXXXXXXXWDRQSLVSEYESEISR 305
MARR G WRYLNPAYYLKRPKR WDRQSLV EYESEISR
Sbjct: 1 MARRSSSSSSSSGAWRYLNPAYYLKRPKRLALLFFVFVAATFAFWDRQSLVREYESEISR 60
Query: 306 LEDDINRLHDQLRKAGVNLDENPVVSNKNSRKDLVVEIDPVNNEKREKVKEAMLHAWNSY 485
L+D +N+L DQLRKAGV+LDENP +K SR+ LV EIDP+NNE+REKVKEAM HAWNSY
Sbjct: 61 LDDKVNQLRDQLRKAGVHLDENPT-GDKVSREKLV-EIDPINNERREKVKEAMAHAWNSY 118
Query: 486 VKYAWGMDELQPQSKNGINSFGGLGATLVDSLDTLYIMGLKDEFQKARDWVAESLDFDKD 665
VKYAWGMDELQPQSKNG+NSFGGLGATLVDSLDTLYIMGLKDEFQ+AR+
Sbjct: 119 VKYAWGMDELQPQSKNGVNSFGGLGATLVDSLDTLYIMGLKDEFQRARE----------- 167
Query: 666 YDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDITDRLLPAWDTTSGIPYNRINLAHGR 845
VVGGLLS+YDLSGDKVFL+KAKDITDRLLPAWDT SGIPYNRINLAHGR
Sbjct: 168 -----------VVGGLLSAYDLSGDKVFLEKAKDITDRLLPAWDTPSGIPYNRINLAHGR 216
Query: 846 AHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQKAENVIKQLQKIYPSDGLLPIYI 1025
AHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQK N L+P I
Sbjct: 217 AHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQKILN------------NLVPYPI 264
Query: 1026 NPHSGTASYSTITFGAMGDSFYEYLLKVWIQGNKTEHVKHYRQMWETSMEGLLSLTKKTT 1205
Y T + FYEYLLKVW+QGNKTEHVKHYRQMWETSMEGLLSLTKKTT
Sbjct: 265 --------YLTCYCMDQINCFYEYLLKVWVQGNKTEHVKHYRQMWETSMEGLLSLTKKTT 316
Query: 1206 PSNYYYICEKNGGSLSDKMDELACFAPGMLALGASGYGP-EKSEQIMNLAKELARTCYNF 1382
PSNYYYICEKNGGSLSDKMDELACFAPGMLALGASGY EK+E+IMNLAKELARTCYNF
Sbjct: 317 PSNYYYICEKNGGSLSDKMDELACFAPGMLALGASGYEETEKAEEIMNLAKELARTCYNF 376
Query: 1383 YQTTPTKLAGENYFFHTGQDMGVGTSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQA 1562
YQTTPTKLAGENYFFHTGQDM VGTSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQA
Sbjct: 377 YQTTPTKLAGENYFFHTGQDMNVGTSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQA 436
Query: 1563 FEKNSRIESGYVGLRDVNTGEKDNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAH 1742
FEKNSRIESGYVGLRDVNTGEKDNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAH
Sbjct: 437 FEKNSRIESGYVGLRDVNTGEKDNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAH 496
Query: 1743 PLRIVPVNDNK-----GIGTP 1790
PLRIVP+NDN GI TP
Sbjct: 497 PLRIVPLNDNSKAHSVGIATP 517
>sptr|Q9SEH8|Q9SEH8 Mannosyl-oligosaccharide 1,2-alpha-mannosidase (EC
3.2.1.113).
Length = 578
Score = 821 bits (2120), Expect = 0.0
Identities = 405/574 (70%), Positives = 458/574 (79%), Gaps = 9/574 (1%)
Frame = +3
Query: 132 MAR--RXXXXXXGTWRYLNPAYYLKRPKRXXXXXXXXXXXXXXXWDRQSLVSEYESEISR 305
MAR R WRY NP+YYLKRPKR WDRQ+LV E++ EIS
Sbjct: 1 MARGSRSVGSSSSKWRYCNPSYYLKRPKRLALLFIVFVCVSFVFWDRQTLVREHQVEISE 60
Query: 306 LED---DINRLHDQLRKAGVNLDENPVVSNKNSRKDLVVEIDPVNNEKREKVKEAMLHAW 476
L+ D+ L D L + K ++ V DP++ E+REKVKEAMLHAW
Sbjct: 61 LQKEVTDLKNLVDDLNNKQGGTSGKTDLGRKATKSSKDVLDDPIDIERREKVKEAMLHAW 120
Query: 477 NSYVKYAWGMDELQPQSKNGINSFGGLGATLVDSLDTLYIMGLKDEFQKARDWVAESLDF 656
SY KYAWG DELQPQSKNG+NSFGGLGATL+DSLDTLYIMGL ++FQKAR+WVA SLDF
Sbjct: 121 GSYEKYAWGQDELQPQSKNGVNSFGGLGATLIDSLDTLYIMGLNEQFQKAREWVANSLDF 180
Query: 657 DKDYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDITDRLLPAWDTTSGIPYNRINLA 836
+KDY+ASVFETTIRVVGGLLS+YDLSGDKVFLDKA +I DRLLPAW+T +GIPYN INL+
Sbjct: 181 NKDYEASVFETTIRVVGGLLSAYDLSGDKVFLDKAIEIADRLLPAWNTPTGIPYNIINLS 240
Query: 837 HGRAHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQKAENVIKQLQKIYPSDGLLP 1016
HGRAHNP WT G+SILADSGTEQLEFI LSQRTGD KYQQK ENVI QL K +P DGLLP
Sbjct: 241 HGRAHNPSWTGGESILADSGTEQLEFIVLSQRTGDLKYQQKVENVIAQLNKTFPDDGLLP 300
Query: 1017 IYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQGNKTEHVKHYRQMWETSMEGLLSLTK 1196
IYINPHSG A YS ITFGAMGDSFYEYLLKVWIQGNKT +KHYR MWE SM+GL SL +
Sbjct: 301 IYINPHSGAAGYSPITFGAMGDSFYEYLLKVWIQGNKTSSIKHYRDMWEKSMKGLSSLIR 360
Query: 1197 KTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALGASGY-GPEKSEQIMNLAKELARTC 1373
++TPS++ YICEKNGGSL+DKMDELACFAPGM+ALG+ GY + S++ ++LA+ELA TC
Sbjct: 361 RSTPSSFTYICEKNGGSLTDKMDELACFAPGMIALGSFGYSAADDSQKFLSLAEELAWTC 420
Query: 1374 YNFYQTTPTKLAGENYFFHTGQDMGVGTSWNILRPETVESLMYLWRLTGNKTYQDWGWDI 1553
YNFYQ+TPTKLAGENYFFH+GQDM VGTSWNILRPETVESL YLWRLTGNKTYQ+WGW+I
Sbjct: 421 YNFYQSTPTKLAGENYFFHSGQDMSVGTSWNILRPETVESLFYLWRLTGNKTYQEWGWNI 480
Query: 1554 FQAFEKNSRIESGYVGLRDVNTGEKDNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNT 1733
FQAFEKNSRIESGYVGL+DVN+G KDNMMQSFFLAETLKY YLLFSP SVIS DEWVFNT
Sbjct: 481 FQAFEKNSRIESGYVGLKDVNSGVKDNMMQSFFLAETLKYFYLLFSPSSVISLDEWVFNT 540
Query: 1734 EAHPLRIVPVNDN---KGIGTPVRQFGRKQGKPE 1826
EAHPLRIV ++ K + + F R G+ E
Sbjct: 541 EAHPLRIVTRHEEGLVKNLNEKQKPFSRIGGRKE 574
>sptr|Q9C512|Q9C512 Mannosyl-oligosaccharide 1,2-alpha-mannosidase,
putative (Mannosyl- oligosaccharide
alpha-1,2-mannosidase, putative).
Length = 560
Score = 810 bits (2092), Expect = 0.0
Identities = 393/563 (69%), Positives = 457/563 (81%), Gaps = 3/563 (0%)
Frame = +3
Query: 132 MARRXXXXXXGTWRYLNPAYYLKRPKRXXXXXXXXXXXXXXXWDRQSLVSEYESEISRLE 311
MAR G W+YLNPAYYL+RP+R WDR +L E+E E+ +L
Sbjct: 1 MARSRSISGYGIWKYLNPAYYLRRPRRLALLFIVFVSVSMLVWDRINLAREHEVEVFKLN 60
Query: 312 DDINRLHDQLRKAGVNLDENPVVSNKNSRKDLVVEIDPVNNEKREKVKEAMLHAWNSYVK 491
++++RL L + + P+ + K++ +D PV+ ++R+KVKEAM+HAW+SY K
Sbjct: 61 EEVSRLEQMLEELNGGVGNKPLKTLKDAPED------PVDKQRRQKVKEAMIHAWSSYEK 114
Query: 492 YAWGMDELQPQSKNGINSFGGLGATLVDSLDTLYIMGLKDEFQKARDWVAESLDFDKDYD 671
YAWG DELQP++K+G +SFGGLGAT+VDSLDTLYIMGL ++FQKAR+WVA SLDFDKDYD
Sbjct: 115 YAWGKDELQPRTKDGTDSFGGLGATMVDSLDTLYIMGLDEQFQKAREWVASSLDFDKDYD 174
Query: 672 ASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDITDRLLPAWDTTSGIPYNRINLAHGRAH 851
AS+FETTIRVVGGLLS+YDLSGDK+FL+KAKDI DRLLPAW+T +GIPYN INL +G AH
Sbjct: 175 ASMFETTIRVVGGLLSAYDLSGDKMFLEKAKDIADRLLPAWNTPTGIPYNIINLRNGNAH 234
Query: 852 NPGWT-NGDSILADSGTEQLEFIALSQRTGDPKYQQKAENVIKQLQKIYPSDGLLPIYIN 1028
NP W GDSILADSGTEQLEFIALSQRTGDPKYQQK E VI +L K +P+DGLLPIYIN
Sbjct: 235 NPSWAAGGDSILADSGTEQLEFIALSQRTGDPKYQQKVEKVITELNKNFPADGLLPIYIN 294
Query: 1029 PHSGTASYSTITFGAMGDSFYEYLLKVWIQGNKTEHVKHYRQMWETSMEGLLSLTKKTTP 1208
P + SYST TFGAMGDSFYEYLLKVW+QGNKT VK YR MWE SM+GLLSL KK+TP
Sbjct: 295 PDNANPSYSTTTFGAMGDSFYEYLLKVWVQGNKTSAVKPYRDMWEKSMKGLLSLVKKSTP 354
Query: 1209 SNYYYICEKNGGSLSDKMDELACFAPGMLALGASGYGPEKSEQIMNLAKELARTCYNFYQ 1388
S++ YICEKNG +L DKMDELACFAPGMLALGASGYGP++ ++ ++LA ELA TCYNFYQ
Sbjct: 355 SSFTYICEKNGNNLIDKMDELACFAPGMLALGASGYGPDEEKKFLSLAGELAWTCYNFYQ 414
Query: 1389 TTPTKLAGENYFFHTGQDMGVGTSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQAFE 1568
+TPTKLAGENYFF GQDM VGTSWNILRPETVESL YLWRLTGNKTYQ+WGW+IFQAFE
Sbjct: 415 STPTKLAGENYFFTAGQDMSVGTSWNILRPETVESLFYLWRLTGNKTYQEWGWNIFQAFE 474
Query: 1569 KNSRIESGYVGLRDVNTGEKDNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPL 1748
KNSR+ESGYVGL+DVNTG KDN MQSFFLAETLKYLYLLFSP SVIS DEWVFNTEAHPL
Sbjct: 475 KNSRVESGYVGLKDVNTGAKDNKMQSFFLAETLKYLYLLFSPSSVISLDEWVFNTEAHPL 534
Query: 1749 RIVPVNDNK--GIGTPVRQFGRK 1811
+IV ND + I R+FG +
Sbjct: 535 KIVARNDPRKPTIALRQRKFGHQ 557
>sptr|Q8H116|Q8H116 Putative mannosidase.
Length = 572
Score = 789 bits (2038), Expect = 0.0
Identities = 382/537 (71%), Positives = 440/537 (81%), Gaps = 1/537 (0%)
Frame = +3
Query: 162 GTWRYLNPAYYLKRPKRXXXXXXXXXXXXXXXWDRQSLVSEYESEISRLEDDINRLHDQL 341
G W+Y NPA+YL+RP+R WDRQSL +Y+ E+S+L +++ RL L
Sbjct: 12 GIWKYFNPAFYLRRPRRLALLIILFVSVSMVVWDRQSLSRDYQFEVSKLNEEVLRLQQML 71
Query: 342 RKAGVNLDENPVVSNKNSRKDLVVEIDPVNNEKREKVKEAMLHAWNSYVKYAWGMDELQP 521
+ ++ V NS KD V+ DPV+ ++ ++VKEAM+HAW+SY KYAWG DELQP
Sbjct: 72 EEIKSVTEDVSV----NSLKD--VQEDPVDAQRMQRVKEAMVHAWSSYEKYAWGQDELQP 125
Query: 522 QSKNGINSFGGLGATLVDSLDTLYIMGLKDEFQKARDWVAESLDFDKDYDASVFETTIRV 701
Q+K+G++SFGGLGAT++D+LDTLYIMGL ++FQKAR+WVA SLDFDKDY AS+FETTIRV
Sbjct: 126 QTKDGVDSFGGLGATMIDALDTLYIMGLDEQFQKAREWVASSLDFDKDYAASMFETTIRV 185
Query: 702 VGGLLSSYDLSGDKVFLDKAKDITDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTNGDSI 881
VGGLLS+YDLSGDK+FL+KA DI DRLLPAWDT SGIPYN INL HG AHNP W GDSI
Sbjct: 186 VGGLLSAYDLSGDKIFLEKAMDIADRLLPAWDTQSGIPYNIINLKHGNAHNPTWAGGDSI 245
Query: 882 LADSGTEQLEFIALSQRTGDPKYQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTI 1061
LADSGTEQLEFIALSQRTGDPKYQQK E VI L K +P+DGLLPIYINP + S STI
Sbjct: 246 LADSGTEQLEFIALSQRTGDPKYQQKVEKVISVLNKNFPADGLLPIYINPDTANPSQSTI 305
Query: 1062 TFGAMGDSFYEYLLKVWIQGNKTEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNG 1241
TFGAMGDSFYEYLLKVW+ GNKT VKHYR MWE SM GLLSL KK+TP ++ YICEK+G
Sbjct: 306 TFGAMGDSFYEYLLKVWVFGNKTSAVKHYRDMWEKSMNGLLSLVKKSTPLSFTYICEKSG 365
Query: 1242 GSLSDKMDELACFAPGMLALGASGYG-PEKSEQIMNLAKELARTCYNFYQTTPTKLAGEN 1418
SL DKMDELACFAPGMLALGASGY P + ++ + LA+ELA TCYNFYQ+TPTKLAGEN
Sbjct: 366 NSLIDKMDELACFAPGMLALGASGYSDPAEGKKFLTLAEELAWTCYNFYQSTPTKLAGEN 425
Query: 1419 YFFHTGQDMGVGTSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYV 1598
YFF++G DM VGTSWNILRPETVESL YLWRLTGNKTYQ+WGW+IF+AFEKNSRIESGYV
Sbjct: 426 YFFNSGSDMSVGTSWNILRPETVESLFYLWRLTGNKTYQEWGWNIFEAFEKNSRIESGYV 485
Query: 1599 GLRDVNTGEKDNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVND 1769
GL+DVNTG KDN MQSFFLAETLKYLYLLFSP +VI DEWVFNTEAHPL+I ND
Sbjct: 486 GLKDVNTGVKDNKMQSFFLAETLKYLYLLFSPTTVIPLDEWVFNTEAHPLKIKSRND 542
>sptr|Q94A04|Q94A04 Putative alpha 1,2-mannosidase.
Length = 572
Score = 787 bits (2033), Expect = 0.0
Identities = 381/537 (70%), Positives = 440/537 (81%), Gaps = 1/537 (0%)
Frame = +3
Query: 162 GTWRYLNPAYYLKRPKRXXXXXXXXXXXXXXXWDRQSLVSEYESEISRLEDDINRLHDQL 341
G W+Y NPA+YL+RP+R WDRQSL +Y+ E+S+L +++ RL L
Sbjct: 12 GIWKYFNPAFYLRRPRRLALLIILFVSVSMVVWDRQSLSRDYQFEVSKLNEEVLRLQQML 71
Query: 342 RKAGVNLDENPVVSNKNSRKDLVVEIDPVNNEKREKVKEAMLHAWNSYVKYAWGMDELQP 521
+ ++ V NS KD V+ DPV+ ++ ++VKEAM+HAW+S+ KYAWG DELQP
Sbjct: 72 EEIKSVTEDVSV----NSLKD--VQEDPVDAQRMQRVKEAMVHAWSSHEKYAWGQDELQP 125
Query: 522 QSKNGINSFGGLGATLVDSLDTLYIMGLKDEFQKARDWVAESLDFDKDYDASVFETTIRV 701
Q+K+G++SFGGLGAT++D+LDTLYIMGL ++FQKAR+WVA SLDFDKDY AS+FETTIRV
Sbjct: 126 QTKDGVDSFGGLGATMIDALDTLYIMGLDEQFQKAREWVASSLDFDKDYAASMFETTIRV 185
Query: 702 VGGLLSSYDLSGDKVFLDKAKDITDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTNGDSI 881
VGGLLS+YDLSGDK+FL+KA DI DRLLPAWDT SGIPYN INL HG AHNP W GDSI
Sbjct: 186 VGGLLSAYDLSGDKIFLEKAMDIADRLLPAWDTQSGIPYNIINLKHGNAHNPTWAGGDSI 245
Query: 882 LADSGTEQLEFIALSQRTGDPKYQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTI 1061
LADSGTEQLEFIALSQRTGDPKYQQK E VI L K +P+DGLLPIYINP + S STI
Sbjct: 246 LADSGTEQLEFIALSQRTGDPKYQQKVEKVISVLNKNFPADGLLPIYINPDTANPSQSTI 305
Query: 1062 TFGAMGDSFYEYLLKVWIQGNKTEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNG 1241
TFGAMGDSFYEYLLKVW+ GNKT VKHYR MWE SM GLLSL KK+TP ++ YICEK+G
Sbjct: 306 TFGAMGDSFYEYLLKVWVFGNKTSAVKHYRDMWEKSMNGLLSLVKKSTPLSFTYICEKSG 365
Query: 1242 GSLSDKMDELACFAPGMLALGASGYG-PEKSEQIMNLAKELARTCYNFYQTTPTKLAGEN 1418
SL DKMDELACFAPGMLALGASGY P + ++ + LA+ELA TCYNFYQ+TPTKLAGEN
Sbjct: 366 NSLIDKMDELACFAPGMLALGASGYSDPAEGKKFLTLAEELAWTCYNFYQSTPTKLAGEN 425
Query: 1419 YFFHTGQDMGVGTSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYV 1598
YFF++G DM VGTSWNILRPETVESL YLWRLTGNKTYQ+WGW+IF+AFEKNSRIESGYV
Sbjct: 426 YFFNSGSDMSVGTSWNILRPETVESLFYLWRLTGNKTYQEWGWNIFEAFEKNSRIESGYV 485
Query: 1599 GLRDVNTGEKDNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVND 1769
GL+DVNTG KDN MQSFFLAETLKYLYLLFSP +VI DEWVFNTEAHPL+I ND
Sbjct: 486 GLKDVNTGVKDNKMQSFFLAETLKYLYLLFSPTTVIPLDEWVFNTEAHPLKIKSRND 542
>sptr|Q9LJB6|Q9LJB6 Alpha 1,2-mannosidase-like protein.
Length = 581
Score = 785 bits (2028), Expect = 0.0
Identities = 383/542 (70%), Positives = 442/542 (81%), Gaps = 6/542 (1%)
Frame = +3
Query: 162 GTWRYLNPAYYLKRPKRXXXXXXXXXXXXXXXWDRQSLVSEYESEISRLEDDINRLHDQL 341
G W+Y NPA+YL+RP+R WDRQSL +Y+ E+S+L +++ RL +
Sbjct: 12 GIWKYFNPAFYLRRPRRLALLIILFVSVSMVVWDRQSLSRDYQFEVSKLNEEVLRLQQMV 71
Query: 342 RKAGV--NLDENPVVSNK---NSRKDLVVEIDPVNNEKREKVKEAMLHAWNSYVKYAWGM 506
+ L+E V+ NS KD V+ DPV+ ++ ++VKEAM+HAW+SY KYAWG
Sbjct: 72 SYLFLFPMLEEIKSVTEDVSVNSLKD--VQEDPVDAQRMQRVKEAMVHAWSSYEKYAWGQ 129
Query: 507 DELQPQSKNGINSFGGLGATLVDSLDTLYIMGLKDEFQKARDWVAESLDFDKDYDASVFE 686
DELQPQ+K+G++SFGGLGAT++D+LDTLYIMGL ++FQKAR+WVA SLDFDKDY AS+FE
Sbjct: 130 DELQPQTKDGVDSFGGLGATMIDALDTLYIMGLDEQFQKAREWVASSLDFDKDYAASMFE 189
Query: 687 TTIRVVGGLLSSYDLSGDKVFLDKAKDITDRLLPAWDTTSGIPYNRINLAHGRAHNPGWT 866
TTIRVVGGLLS+YDLSGDK+FL+KA DI DRLLPAWDT SGIPYN INL HG AHNP W
Sbjct: 190 TTIRVVGGLLSAYDLSGDKIFLEKAMDIADRLLPAWDTQSGIPYNIINLKHGNAHNPTWA 249
Query: 867 NGDSILADSGTEQLEFIALSQRTGDPKYQQKAENVIKQLQKIYPSDGLLPIYINPHSGTA 1046
GDSILADSGTEQLEFIALSQRTGDPKYQQK E VI L K +P+DGLLPIYINP +
Sbjct: 250 GGDSILADSGTEQLEFIALSQRTGDPKYQQKVEKVISVLNKNFPADGLLPIYINPDTANP 309
Query: 1047 SYSTITFGAMGDSFYEYLLKVWIQGNKTEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYI 1226
S STITFGAMGDSFYEYLLKVW+ GNKT VKHYR MWE SM GLLSL KK+TP ++ YI
Sbjct: 310 SQSTITFGAMGDSFYEYLLKVWVFGNKTSAVKHYRDMWEKSMNGLLSLVKKSTPLSFTYI 369
Query: 1227 CEKNGGSLSDKMDELACFAPGMLALGASGYG-PEKSEQIMNLAKELARTCYNFYQTTPTK 1403
CEK+G SL DKMDELACFAPGMLALGASGY P + ++ + LA+ELA TCYNFYQ+TPTK
Sbjct: 370 CEKSGNSLIDKMDELACFAPGMLALGASGYSDPAEGKKFLTLAEELAWTCYNFYQSTPTK 429
Query: 1404 LAGENYFFHTGQDMGVGTSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRI 1583
LAGENYFF++G DM VGTSWNILRPETVESL YLWRLTGNKTYQ+WGW+IF+AFEKNSRI
Sbjct: 430 LAGENYFFNSGSDMSVGTSWNILRPETVESLFYLWRLTGNKTYQEWGWNIFEAFEKNSRI 489
Query: 1584 ESGYVGLRDVNTGEKDNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPV 1763
ESGYVGL+DVNTG KDN MQSFFLAETLKYLYLLFSP +VI DEWVFNTEAHPL+I
Sbjct: 490 ESGYVGLKDVNTGVKDNKMQSFFLAETLKYLYLLFSPTTVIPLDEWVFNTEAHPLKIKSR 549
Query: 1764 ND 1769
ND
Sbjct: 550 ND 551
>sptr|Q7QFL9|Q7QFL9 AgCP13734 (Fragment).
Length = 727
Score = 402 bits (1033), Expect = e-110
Identities = 212/468 (45%), Positives = 294/468 (62%), Gaps = 11/468 (2%)
Frame = +3
Query: 420 DPVNNEKREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGINS--FGG--LGATLVDSLDT 587
D EKR KVKE M+HAW++Y YAWG +EL+P SK G ++ FG LGAT+VD LDT
Sbjct: 258 DRAVREKRNKVKEMMVHAWSNYKLYAWGKNELRPLSKRGHSNSIFGSFDLGATIVDGLDT 317
Query: 588 LYIMGLKDEFQKARDWVAESLDFDK-DYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAK 764
LY+MGL EF + R+WV D D + SVFET IR +GG L+ Y +GD++FL+KA+
Sbjct: 318 LYLMGLHKEFDEGREWVERKFTLDNVDAELSVFETNIRFIGGFLTCYAFTGDRMFLEKAR 377
Query: 765 DITDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTNG-DSILADSGTEQLEFIALSQRTGD 941
+ D+LLPA+ T +GIPY +N+ +G + N GW +G SIL++ GT +EF LS TGD
Sbjct: 378 YVADKLLPAFQTPTGIPYALVNVHNGVSKNYGWASGGSSILSEFGTLHMEFAYLSDITGD 437
Query: 942 PKYQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQG 1121
Y+++ + + L+ I GL P Y+NP +G ++ GA+GDSFYEYLLK WIQ
Sbjct: 438 SVYRERVQTIRAVLKDIEKPKGLYPNYLNPKTGKWGQQHMSLGALGDSFYEYLLKAWIQS 497
Query: 1122 NKTEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLAL 1301
+ + R+M++ +M+ ++ + TPS Y+ + L KMD LACFA G+ L
Sbjct: 498 GQEDW--EAREMYDDAMQAIIKHMVRNTPSGLTYVSDMKFDRLEHKMDHLACFAGGLFGL 555
Query: 1302 GASGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQD---MGVGTSWNIL 1472
G + + S+Q M + + L TC+ Y T T+L E++ F+ G + + + IL
Sbjct: 556 GGTSLNNQYSQQYMEIGEGLTNTCHESYIRTYTRLGPESFRFNDGVEAKALKAQEKYYIL 615
Query: 1473 RPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDVNTGE--KDNMMQS 1646
RPET ES +WRLT ++ Y+DWGWD QA EK+ R +GY GL++V E KD++ QS
Sbjct: 616 RPETFESYFIMWRLTHDQKYRDWGWDAVQALEKHCRTPTGYTGLKNVYLTEPVKDDVQQS 675
Query: 1647 FFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVNDNKGIGTP 1790
FFLAETLKYLYLLFS S++ D+WVFNTEAHPL PV D + P
Sbjct: 676 FFLAETLKYLYLLFSDDSLLPLDDWVFNTEAHPL---PVKDRNPLYRP 720
>sw|P45701|M1A1_RABIT Mannosyl-oligosaccharide 1,2-alpha-mannosidase
IA (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IA)
(Alpha-1,2-mannosidase IA) (Mannosidase alpha class 1A
member 1) (Man(9)-alpha-mannosidase) (Fragment).
Length = 469
Score = 399 bits (1025), Expect = e-109
Identities = 216/469 (46%), Positives = 299/469 (63%), Gaps = 10/469 (2%)
Frame = +3
Query: 420 DPVNNEKREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGINS--FGGL-GATLVDSLDTL 590
D EKR K+KE M HAWNSY +YAWG++EL+P +K G +S FG + GAT+VD+LDTL
Sbjct: 5 DAAVREKRAKIKEMMEHAWNSYKRYAWGLNELKPITKEGHSSSLFGTIKGATIVDALDTL 64
Query: 591 YIMGLKDEFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDI 770
+IMG++ EFQ+A+ W+AE+LDF+ + + SVFE IR VGGLLS+Y LSG+++F KA ++
Sbjct: 65 FIMGMESEFQEAKSWIAENLDFNVNAEISVFEVNIRFVGGLLSAYYLSGEEIFRKKAVEL 124
Query: 771 TDRLLPAWDTTSGIPYNRINLAHGRAHNPGW-TNGDSILADSGTEQLEFIALSQRTGDPK 947
+LLPA+ T SGIP+ +N+ G N W + G SILA+ GT LEF+ LS +G+P
Sbjct: 125 GIKLLPAFHTPSGIPWALLNIKSGIGRNWPWASGGSSILAEFGTLHLEFMHLSHLSGNPI 184
Query: 948 YQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQGNK 1127
+ +K N+ K L K+ +GL P Y+NP SG ++ G +GDSFYEYLLK W+ K
Sbjct: 185 FAEKVMNIRKVLNKLEKPEGLYPNYLNPSSGQWGQHHVSIGGLGDSFYEYLLKAWLMSEK 244
Query: 1128 TEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALGA 1307
T+ ++M+ +++ + + + + YI E GG L KM L CFA GM ALGA
Sbjct: 245 TD--LEAKKMYFDAVQAIETHLIRKSSGGLTYIAEWKGGLLEHKMGHLTCFAGGMFALGA 302
Query: 1308 SGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMGVGTSWN----ILR 1475
G +++ + L E+ARTC+ Y T KL E + F G + + T N ILR
Sbjct: 303 DGAPEGRAQHYLELGAEIARTCHESYNRTFMKLGPEAFRFDGGVE-AIATRQNEKYYILR 361
Query: 1476 PETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDVN-TGEK-DNMMQSF 1649
PE VE+ MY+WRLT + Y+ W W+ +A E + R+ GY GLRDV T EK DN+ QSF
Sbjct: 362 PEVVETYMYMWRLTHDPKYRKWAWEAVEALESHCRVNGGYSGLRDVYFTHEKYDNVQQSF 421
Query: 1650 FLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVNDNKGIGTPVR 1796
FLAETLKYLYL+FS ++ + W+FNTEAH L I+P D K + V+
Sbjct: 422 FLAETLKYLYLIFSDDDLLPLEHWIFNTEAHLLPILP-TDQKEVEVKVK 469
>sptrnew|AAH63300|AAH63300 MAN1A2 protein.
Length = 641
Score = 396 bits (1018), Expect = e-109
Identities = 216/534 (40%), Positives = 317/534 (59%), Gaps = 25/534 (4%)
Frame = +3
Query: 264 RQSLVSEYESEISRLEDDINRLHDQLRKAGVNLDENPVVSNKNSRKDLVVEIDPVNN--- 434
R + +++E + ++ + + +++R A + ++N VV +++ + P+ N
Sbjct: 108 RNKIRADHEKALEEAKEKLRKSREEIR-AEIQTEKNKVVQEMKIKENKPLPPVPIPNLVG 166
Query: 435 ------------EKREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGI--NSFGG--LGAT 566
EKREK+KE M HAW++Y Y WG +EL+P ++ G N FG +GAT
Sbjct: 167 IRGGDPEDNDIREKREKIKEMMKHAWDNYRTYGWGHNELRPIARKGHSPNIFGSSQMGAT 226
Query: 567 LVDSLDTLYIMGLKDEFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKV 746
+VD+LDTLYIMGL DEF + W+ ++LDF + + SVFE IR +GGLL++Y LSG+++
Sbjct: 227 IVDALDTLYIMGLHDEFLDGQRWIEDNLDFSVNSEVSVFEVNIRFIGGLLAAYYLSGEEI 286
Query: 747 FLDKAKDITDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTN-GDSILADSGTEQLEFIAL 923
F KA + ++LLPA++T +GIP+ +NL G N GW + G SILA+ GT +EFI L
Sbjct: 287 FKIKAVQLAEKLLPAFNTPTGIPWAMVNLKSGVGRNWGWASAGSSILAEFGTLHMEFIHL 346
Query: 924 SQRTGDPKYQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLL 1103
S TGD Y +K ++ K LQK+ +GL P Y+NP +G + G +GDSFYEYLL
Sbjct: 347 SYLTGDLTYYKKVMHIRKLLQKMDRPNGLYPNYLNPRTGRWGQYHTSVGGLGDSFYEYLL 406
Query: 1104 KVWIQGNKTEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFA 1283
K W+ +KT+H R+M++ ++E + K + +I E G L KM LACFA
Sbjct: 407 KAWLMSDKTDH--EARKMYDDAIEAIEKHLIKKSRGGLTFIGEWKNGHLEKKMGHLACFA 464
Query: 1284 PGMLALGASGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQD---MGVG 1454
GM ALGA G +K+ + L E+ARTC+ Y T KL E++ F + +
Sbjct: 465 GGMFALGADGSRADKAGHYLELGAEIARTCHESYDRTALKLGPESFKFDGAVEAVAVRQA 524
Query: 1455 TSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDV--NTGEK 1628
+ ILRPE +E+ YLWR T + Y+ WGW+ A EK R+ G+ G++DV +T
Sbjct: 525 EKYYILRPEVIETYWYLWRFTHDPRYRQWGWEAALAIEKYCRVNGGFSGVKDVYSSTPTH 584
Query: 1629 DNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVNDNKGIGTP 1790
D++ QSFFLAETLKYLYLLFS ++ D WVFNTEAHPL ++ + + G P
Sbjct: 585 DDVQQSFFLAETLKYLYLLFSGDDLLPLDHWVFNTEAHPLPVLHLANTTLSGNP 638
>sw|O60476|M1A2_HUMAN Mannosyl-oligosaccharide 1,2-alpha-mannosidase
IB (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IB)
(Alpha-1,2-mannosidase IB) (Mannosidase alpha class 1A
member 2).
Length = 641
Score = 396 bits (1018), Expect = e-109
Identities = 216/534 (40%), Positives = 317/534 (59%), Gaps = 25/534 (4%)
Frame = +3
Query: 264 RQSLVSEYESEISRLEDDINRLHDQLRKAGVNLDENPVVSNKNSRKDLVVEIDPVNN--- 434
R + +++E + ++ + + +++R A + ++N VV +++ + P+ N
Sbjct: 108 RNKIRADHEKALEEAKEKLRKSREEIR-AEIQTEKNKVVQEMKIKENKPLPPVPIPNLVG 166
Query: 435 ------------EKREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGI--NSFGG--LGAT 566
EKREK+KE M HAW++Y Y WG +EL+P ++ G N FG +GAT
Sbjct: 167 IRGGDPEDNDIREKREKIKEMMKHAWDNYRTYGWGHNELRPIARKGHSPNIFGSSQMGAT 226
Query: 567 LVDSLDTLYIMGLKDEFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKV 746
+VD+LDTLYIMGL DEF + W+ ++LDF + + SVFE IR +GGLL++Y LSG+++
Sbjct: 227 IVDALDTLYIMGLHDEFLDGQRWIEDNLDFSVNSEVSVFEVNIRFIGGLLAAYYLSGEEI 286
Query: 747 FLDKAKDITDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTN-GDSILADSGTEQLEFIAL 923
F KA + ++LLPA++T +GIP+ +NL G N GW + G SILA+ GT +EFI L
Sbjct: 287 FKIKAVQLAEKLLPAFNTPTGIPWAMVNLKSGVGRNWGWASAGSSILAEFGTLHMEFIHL 346
Query: 924 SQRTGDPKYQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLL 1103
S TGD Y +K ++ K LQK+ +GL P Y+NP +G + G +GDSFYEYLL
Sbjct: 347 SYLTGDLTYYKKVMHIRKLLQKMDRPNGLYPNYLNPRTGRWGQYHTSVGGLGDSFYEYLL 406
Query: 1104 KVWIQGNKTEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFA 1283
K W+ +KT+H R+M++ ++E + K + +I E G L KM LACFA
Sbjct: 407 KAWLMSDKTDH--EARKMYDDAIEAIEKHLIKKSRGGLTFIGEWKNGHLEKKMGHLACFA 464
Query: 1284 PGMLALGASGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQD---MGVG 1454
GM ALGA G +K+ + L E+ARTC+ Y T KL E++ F + +
Sbjct: 465 GGMFALGADGSRADKAGHYLELGAEIARTCHESYDRTALKLGPESFKFDGAVEAVAVRQA 524
Query: 1455 TSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDV--NTGEK 1628
+ ILRPE +E+ YLWR T + Y+ WGW+ A EK R+ G+ G++DV +T
Sbjct: 525 EKYYILRPEVIETYWYLWRFTHDPRYRQWGWEAALAIEKYCRVNGGFSGVKDVYSSTPTH 584
Query: 1629 DNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVNDNKGIGTP 1790
D++ QSFFLAETLKYLYLLFS ++ D WVFNTEAHPL ++ + + G P
Sbjct: 585 DDVQQSFFLAETLKYLYLLFSGDDLLPLDHWVFNTEAHPLPVLHLANTTLSGNP 638
>sw|P39098|M1A2_MOUSE Mannosyl-oligosaccharide 1,2-alpha-mannosidase
IB (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IB)
(Alpha-1,2-mannosidase IB) (Mannosidase alpha class 1A
member 2).
Length = 641
Score = 395 bits (1015), Expect = e-108
Identities = 214/534 (40%), Positives = 316/534 (59%), Gaps = 25/534 (4%)
Frame = +3
Query: 264 RQSLVSEYESEISRLEDDINRLHDQLRKAGVNLDENPVVSNKNSRKDLVVEIDPVNN--- 434
R + +++E + ++ + + +++R A + ++N V +++ V+ PV
Sbjct: 108 RNKIRADHEKALEEAKEKLRKSREEIR-AEIQTEKNKVAQAMKTKETRVLPPVPVPQRVG 166
Query: 435 ------------EKREKVKEAMLHAWNSYVKYAWGMDELQPQSKNG--INSFGG--LGAT 566
+KR+K+KE M HAW++Y Y WG +EL+P ++ G N FG +GAT
Sbjct: 167 VSGGDPEDMEIKKKRDKIKEMMKHAWDNYRTYGWGHNELRPIARKGHSTNIFGSSQMGAT 226
Query: 567 LVDSLDTLYIMGLKDEFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKV 746
+VD+LDTLYIMGL DEF + W+ E+LDF + + SVFE IR +GGLL++Y LSG+++
Sbjct: 227 IVDALDTLYIMGLHDEFMDGQRWIEENLDFSVNSEVSVFEVNIRFIGGLLAAYYLSGEEI 286
Query: 747 FLDKAKDITDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTN-GDSILADSGTEQLEFIAL 923
F KA + ++LLPA++T +GIP+ +NL G N GW + G SILA+ GT +EF+ L
Sbjct: 287 FKTKAVQLAEKLLPAFNTPTGIPWAMVNLKSGVGRNWGWASAGSSILAEFGTLHMEFVHL 346
Query: 924 SQRTGDPKYQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLL 1103
S TGD Y K ++ K LQK+ +GL P Y+NP +G + G +GDSFYEYLL
Sbjct: 347 SYLTGDLTYYNKVMHIRKLLQKMERPNGLYPNYLNPRTGRWGQYHTSVGGLGDSFYEYLL 406
Query: 1104 KVWIQGNKTEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFA 1283
K W+ +KT+H R+M++ ++E + K + +I E G L KM LACFA
Sbjct: 407 KAWLMSDKTDH--EARRMYDDAVEAIEKHLIKKSRGGLVFIGEWKNGHLERKMGHLACFA 464
Query: 1284 PGMLALGASGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQD---MGVG 1454
GM ALGA G +K+ + L E+ARTC+ Y T KL E++ F + +
Sbjct: 465 GGMFALGADGSRKDKAGHYLELGAEIARTCHESYDRTALKLGPESFKFDGAVEAVAVRQA 524
Query: 1455 TSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDV--NTGEK 1628
+ ILRPE +E+ YLWR T + Y+ WGW+ A EK+ R+ G+ G++DV T
Sbjct: 525 EKYYILRPEVIETYWYLWRFTHDPRYRQWGWEAALAIEKSCRVSGGFSGVKDVYAPTPVH 584
Query: 1629 DNMMQSFFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVNDNKGIGTP 1790
D++ QSFFLAETLKYLYLLFS ++ D WVFNTEAHPL ++ + ++ G P
Sbjct: 585 DDVQQSFFLAETLKYLYLLFSGDDLLPLDHWVFNTEAHPLPVLRLANSTLSGNP 638
>sptrnew|BAC27225|BAC27225 Adult male thymus cDNA, RIKEN full-length
enriched library, clone:5832424A03 product:mannosidase 1,
alpha, full insert sequence.
Length = 655
Score = 391 bits (1005), Expect = e-107
Identities = 205/456 (44%), Positives = 288/456 (63%), Gaps = 10/456 (2%)
Frame = +3
Query: 420 DPVNNEKREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGINS--FGGL-GATLVDSLDTL 590
D EKR K+KE M HAWN+Y +YAWG++EL+P SK G +S FG + GAT+VD+LDTL
Sbjct: 191 DATIREKRAKIKEMMTHAWNNYKRYAWGLNELKPISKEGHSSSLFGNIKGATIVDALDTL 250
Query: 591 YIMGLKDEFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDI 770
+IMG+K EFQ+A+ W+ + LDF+ + + SVFE IR VGGLLS+Y LSG+++F KA ++
Sbjct: 251 FIMGMKTEFQEAKSWIKKYLDFNVNAEVSVFEVNIRFVGGLLSAYYLSGEEIFRKKAVEL 310
Query: 771 TDRLLPAWDTTSGIPYNRINLAHGRAHNPGW-TNGDSILADSGTEQLEFIALSQRTGDPK 947
+LLPA+ T SGIP+ +N+ G N W + G SILA+ GT LEF+ LS +GDP
Sbjct: 311 GVKLLPAFHTPSGIPWALLNMKSGIGRNWPWASGGSSILAEFGTLHLEFMHLSHLSGDPV 370
Query: 948 YQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQGNK 1127
+ +K + L K+ +GL P Y+NP SG ++ G +GDSFYEYLLK W+ +K
Sbjct: 371 FAEKVMKIRTVLNKLDKPEGLYPNYLNPSSGQWGQHHVSVGGLGDSFYEYLLKAWLMSDK 430
Query: 1128 TEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALGA 1307
T+ ++M+ +++ + + + + YI E GG L KM L CFA GM ALGA
Sbjct: 431 TD--LEAKKMYFDAVQAIETHLIRKSSGGLTYIAEWKGGLLEHKMGHLTCFAGGMFALGA 488
Query: 1308 SGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMGVGTSWN----ILR 1475
G +++ + L E+ARTC+ Y T KL E + F G + + T N ILR
Sbjct: 489 DGAPEARAQHYLELGAEIARTCHESYNRTYVKLGPEAFRFDGGVE-AIATRQNEKYYILR 547
Query: 1476 PETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDVNTGEK--DNMMQSF 1649
PE +E+ MY+WRLT + Y+ W W+ +A E + R+ GY GLRDV + D++ QSF
Sbjct: 548 PEVIETYMYMWRLTHDPKYRTWAWEAVEALESHCRVNGGYSGLRDVYIARESYDDVQQSF 607
Query: 1650 FLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIV 1757
FLAETLKYLYL+FS ++ + W+FNTEAHP I+
Sbjct: 608 FLAETLKYLYLIFSDDDLLPLEHWIFNTEAHPFPIL 643
>sw|P45700|M1A1_MOUSE Mannosyl-oligosaccharide 1,2-alpha-mannosidase
IA (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IA)
(Alpha-1,2-mannosidase IA) (Mannosidase alpha class 1A
member 1) (Man(9)-alpha-mannosidase) (Man9-mannosidase).
Length = 655
Score = 391 bits (1005), Expect = e-107
Identities = 205/456 (44%), Positives = 288/456 (63%), Gaps = 10/456 (2%)
Frame = +3
Query: 420 DPVNNEKREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGINS--FGGL-GATLVDSLDTL 590
D EKR K+KE M HAWN+Y +YAWG++EL+P SK G +S FG + GAT+VD+LDTL
Sbjct: 191 DATIREKRAKIKEMMTHAWNNYKRYAWGLNELKPISKEGHSSSLFGNIKGATIVDALDTL 250
Query: 591 YIMGLKDEFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDI 770
+IMG+K EFQ+A+ W+ + LDF+ + + SVFE IR VGGLLS+Y LSG+++F KA ++
Sbjct: 251 FIMGMKTEFQEAKSWIKKYLDFNVNAEVSVFEVNIRFVGGLLSAYYLSGEEIFRKKAVEL 310
Query: 771 TDRLLPAWDTTSGIPYNRINLAHGRAHNPGW-TNGDSILADSGTEQLEFIALSQRTGDPK 947
+LLPA+ T SGIP+ +N+ G N W + G SILA+ GT LEF+ LS +GDP
Sbjct: 311 GVKLLPAFHTPSGIPWALLNMKSGIGRNWPWASGGSSILAEFGTLHLEFMHLSHLSGDPV 370
Query: 948 YQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQGNK 1127
+ +K + L K+ +GL P Y+NP SG ++ G +GDSFYEYLLK W+ +K
Sbjct: 371 FAEKVMKIRTVLNKLDKPEGLYPNYLNPSSGQWGQHHVSVGGLGDSFYEYLLKAWLMSDK 430
Query: 1128 TEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALGA 1307
T+ ++M+ +++ + + + + YI E GG L KM L CFA GM ALGA
Sbjct: 431 TD--LEAKKMYFDAVQAIETHLIRKSSGGLTYIAEWKGGLLEHKMGHLTCFAGGMFALGA 488
Query: 1308 SGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMGVGTSWN----ILR 1475
G +++ + L E+ARTC+ Y T KL E + F G + + T N ILR
Sbjct: 489 DGAPEARAQHYLELGAEIARTCHESYNRTYVKLGPEAFRFDGGVE-AIATRQNEKYYILR 547
Query: 1476 PETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDVNTGEK--DNMMQSF 1649
PE +E+ MY+WRLT + Y+ W W+ +A E + R+ GY GLRDV + D++ QSF
Sbjct: 548 PEVIETYMYMWRLTHDPKYRTWAWEAVEALESHCRVNGGYSGLRDVYIARESYDDVQQSF 607
Query: 1650 FLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIV 1757
FLAETLKYLYL+FS ++ + W+FNTEAHP I+
Sbjct: 608 FLAETLKYLYLIFSDDDLLPLEHWIFNTEAHPFPIL 643
>sptr|O18498|O18498 Alpha 1,2-mannosidase.
Length = 670
Score = 388 bits (996), Expect = e-106
Identities = 220/499 (44%), Positives = 291/499 (58%), Gaps = 29/499 (5%)
Frame = +3
Query: 345 KAGVNLDENPVVSNKNSRKD------------------LVVEIDPVNNEKREKVKEAMLH 470
K V+ E P V N + KD L DP K E VK+ MLH
Sbjct: 148 KPPVDAIEEPAVGNNAANKDVSPSGPKAESSDKFVAVALAPGADPEIKHKLETVKKMMLH 207
Query: 471 AWNSYVKYAWGMDELQPQSK----NGINSFGGLGATLVDSLDTLYIMGLKDEFQKARDWV 638
AW +Y YAWG +EL+P SK + + G LGAT+VD LDTLY+MGL DEF++ RDWV
Sbjct: 208 AWYNYKLYAWGKNELKPMSKRAHLSSVFGAGELGATIVDGLDTLYLMGLNDEFREGRDWV 267
Query: 639 AESLDFDK-DYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDITDRLLPAWDTTSGIP 815
AE L ++ D D SVFETTIR VGGLLS Y L+GD +F DKA ++ D LLPA+DT +G+P
Sbjct: 268 AEHLHINEIDSDLSVFETTIRFVGGLLSCYALTGDTMFRDKAAEVGDALLPAFDTPTGLP 327
Query: 816 YNRINLAHGRAHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQKAENVIKQLQKIY 995
Y IN + + W +SIL++ GT LEF LS TG Y+QK + + L +I
Sbjct: 328 YALINPSTKASRQYHWAGPNSILSELGTLHLEFTYLSDVTGRDIYRQKVSRIREVLDQID 387
Query: 996 PSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQGNKTEHVKHYRQMWETSME 1175
L P +INP +G ++ GA+GDSFYEYLLK W+ + + R M++T+M+
Sbjct: 388 KPGDLYPNFINPRTGQWGQRHMSLGALGDSFYEYLLKAWLMSGGAD--EQARIMFDTAMQ 445
Query: 1176 GLLSLTKKTTPSNYYYICE-KNGGSLSDKMDELACFAPGMLALGASGYGPEKSEQIMNLA 1352
L + +PS Y+ E K G + +KMD L+CFA GM AL ++ SE+ M++A
Sbjct: 446 AALDKMLRVSPSGLAYLAELKYGRIIEEKMDHLSCFAGGMFALASTTLDNSMSERYMDVA 505
Query: 1353 KELARTCYNFYQTTPTKLAGENYFFHTGQDMGVGTSWN---ILRPETVESLMYLWRLTGN 1523
K+L TC+ Y + TKL E + F + S +LRPET ES +WRLT
Sbjct: 506 KKLTNTCHESYARSETKLGPEAFRFSNAAEARAQKSNEKVYLLRPETFESYFIMWRLTKQ 565
Query: 1524 KTYQDWGWDIFQAFEKNSRIESGYVGLRDV--NTGEKDNMMQSFFLAETLKYLYLLFSPP 1697
+ Y+DW W+ QA EK+ R+E GY GL +V + D++ QSFFLAETLKYLYL+F
Sbjct: 566 QMYRDWAWEAVQALEKHCRVEGGYTGLVNVYHANPQGDDVQQSFFLAETLKYLYLIFGDD 625
Query: 1698 SVISFDEWVFNTEAHPLRI 1754
S + DEWVFNTEAHP I
Sbjct: 626 SFLPLDEWVFNTEAHPFPI 644
>sptrnew|AAR30196|AAR30196 RE43942p.
Length = 667
Score = 386 bits (992), Expect = e-106
Identities = 206/461 (44%), Positives = 290/461 (62%), Gaps = 11/461 (2%)
Frame = +3
Query: 417 IDPVNNEKREKVKEAMLHAWNSYVKYAWGMDELQPQSK--NGINSFGG--LGATLVDSLD 584
+D E+R+KVKE M HAW++Y YAWG +EL+P S+ + + FG LGAT+VD LD
Sbjct: 193 LDATLEERRQKVKEMMEHAWHNYKLYAWGKNELRPLSQRPHSASIFGSYDLGATIVDGLD 252
Query: 585 TLYIMGLKDEFQKARDWVAESLDFDK-DYDASVFETTIRVVGGLLSSYDLSGDKVFLDKA 761
TLYIMGL+ E+++ RDW+ D + SVFET IR VGG+L+ Y +GD ++ +KA
Sbjct: 253 TLYIMGLEKEYREGRDWIERKFSLDNISAELSVFETNIRFVGGMLTLYAFTGDPLYKEKA 312
Query: 762 KDITDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTNG-DSILADSGTEQLEFIALSQRTG 938
+ + D+LLPA+ T +GIPY +N G A N GW +G SIL++ GT LEF LS TG
Sbjct: 313 QHVADKLLPAFQTPTGIPYALVNTKTGVAKNYGWASGGSSILSEFGTLHLEFAYLSDITG 372
Query: 939 DPKYQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQ 1118
+P Y+++ + + + L++I GL P ++NP +G ++ GA+GDS+YEYLLK W+Q
Sbjct: 373 NPLYRERVQTIRQVLKEIEKPKGLYPNFLNPKTGKWGQLHMSLGALGDSYYEYLLKAWLQ 432
Query: 1119 GNKTEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLA 1298
+T+ + R+M++ +M +L +T+P Y+ + L KMD LACF+ G+ A
Sbjct: 433 SGQTD--EEAREMFDEAMLAILDKMVRTSPGGLTYVSDLKFDRLEHKMDHLACFSGGLFA 490
Query: 1299 LGASGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMGVGTS---WNI 1469
LGA+ + +++ M + K + TC+ Y PT+L E + F + S + I
Sbjct: 491 LGAATRQNDYTDKYMEVGKGITNTCHESYIRAPTQLGPEAFRFSEAVEARALRSQEKYYI 550
Query: 1470 LRPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDVNTGE--KDNMMQ 1643
LRPET ES LWRLT ++ Y+DWGW+ A EK+ R GY GLR+V E KD++ Q
Sbjct: 551 LRPETFESYFVLWRLTHDQKYRDWGWEAVLALEKHCRTAHGYCGLRNVYQQEPQKDDVQQ 610
Query: 1644 SFFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVN 1766
SFFLAETLKYLYLLFS SV+ DEWVFNTEAHPL I N
Sbjct: 611 SFFLAETLKYLYLLFSDDSVLPLDEWVFNTEAHPLPIKGAN 651
>sptr|Q9W2W7|Q9W2W7 Alpha-MAN-I protein.
Length = 667
Score = 386 bits (992), Expect = e-106
Identities = 206/461 (44%), Positives = 290/461 (62%), Gaps = 11/461 (2%)
Frame = +3
Query: 417 IDPVNNEKREKVKEAMLHAWNSYVKYAWGMDELQPQSK--NGINSFGG--LGATLVDSLD 584
+D E+R+KVKE M HAW++Y YAWG +EL+P S+ + + FG LGAT+VD LD
Sbjct: 193 LDATLEERRQKVKEMMEHAWHNYKLYAWGKNELRPLSQRPHSASIFGSYDLGATIVDGLD 252
Query: 585 TLYIMGLKDEFQKARDWVAESLDFDK-DYDASVFETTIRVVGGLLSSYDLSGDKVFLDKA 761
TLYIMGL+ E+++ RDW+ D + SVFET IR VGG+L+ Y +GD ++ +KA
Sbjct: 253 TLYIMGLEKEYREGRDWIERKFSLDNISAELSVFETNIRFVGGMLTLYAFTGDPLYKEKA 312
Query: 762 KDITDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTNG-DSILADSGTEQLEFIALSQRTG 938
+ + D+LLPA+ T +GIPY +N G A N GW +G SIL++ GT LEF LS TG
Sbjct: 313 QHVADKLLPAFQTPTGIPYALVNTKTGVAKNYGWASGGSSILSEFGTLHLEFAYLSDITG 372
Query: 939 DPKYQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQ 1118
+P Y+++ + + + L++I GL P ++NP +G ++ GA+GDS+YEYLLK W+Q
Sbjct: 373 NPLYRERVQTIRQVLKEIEKPKGLYPNFLNPKTGKWGQLHMSLGALGDSYYEYLLKAWLQ 432
Query: 1119 GNKTEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLA 1298
+T+ + R+M++ +M +L +T+P Y+ + L KMD LACF+ G+ A
Sbjct: 433 SGQTD--EEAREMFDEAMLAILDKMVRTSPGGLTYVSDLKFDRLEHKMDHLACFSGGLFA 490
Query: 1299 LGASGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMGVGTS---WNI 1469
LGA+ + +++ M + K + TC+ Y PT+L E + F + S + I
Sbjct: 491 LGAATRQNDYTDKYMEVGKGITNTCHESYIRAPTQLGPEAFRFSEAVEARALRSQEKYYI 550
Query: 1470 LRPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDVNTGE--KDNMMQ 1643
LRPET ES LWRLT ++ Y+DWGW+ A EK+ R GY GLR+V E KD++ Q
Sbjct: 551 LRPETFESYFVLWRLTHDQKYRDWGWEAVLALEKHCRTAHGYCGLRNVYQQEPQKDDVQQ 610
Query: 1644 SFFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVN 1766
SFFLAETLKYLYLLFS SV+ DEWVFNTEAHPL I N
Sbjct: 611 SFFLAETLKYLYLLFSDDSVLPLDEWVFNTEAHPLPIKGAN 651
>sw|P53624|M121_DROME Mannosyl-oligosaccharide alpha-1,2-mannosidase
isoform 1 (EC 3.2.1.113) (Man(9)-alpha-mannosidase).
Length = 667
Score = 386 bits (991), Expect = e-106
Identities = 206/461 (44%), Positives = 289/461 (62%), Gaps = 11/461 (2%)
Frame = +3
Query: 417 IDPVNNEKREKVKEAMLHAWNSYVKYAWGMDELQPQSK--NGINSFGG--LGATLVDSLD 584
+D E+R+KVKE M HAW++Y YAWG +EL+P S+ + + FG LGAT+VD LD
Sbjct: 193 LDATLEERRQKVKEMMEHAWHNYKLYAWGKNELRPLSQRPHSASIFGSYDLGATIVDGLD 252
Query: 585 TLYIMGLKDEFQKARDWVAESLDFDK-DYDASVFETTIRVVGGLLSSYDLSGDKVFLDKA 761
TLYIMGL+ E+++ RDW+ D + SVFET IR VGG+L+ Y +GD ++ +KA
Sbjct: 253 TLYIMGLEKEYREGRDWIERKFSLDNISAELSVFETNIRFVGGMLTLYAFTGDPLYKEKA 312
Query: 762 KDITDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTNG-DSILADSGTEQLEFIALSQRTG 938
+ + D+LLPA+ T +GIPY +N G A N GW +G SIL++ GT LEF LS TG
Sbjct: 313 QHVADKLLPAFQTPTGIPYALVNTKTGVAKNYGWASGGSSILSEFGTLHLEFAYLSDITG 372
Query: 939 DPKYQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQ 1118
+P Y+++ + + + L++I GL P ++NP +G ++ GA+GDS+YEYLLK W+Q
Sbjct: 373 NPLYRERVQTIRQVLKEIEKPKGLYPNFLNPKTGKWGQLHMSLGALGDSYYEYLLKAWLQ 432
Query: 1119 GNKTEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLA 1298
+T+ + R+M++ +M +L +T+P Y+ + L KMD LACF+ G+ A
Sbjct: 433 SGQTD--EEAREMFDEAMLAILDKMVRTSPGGLTYVSDLKFDRLEHKMDHLACFSGGLFA 490
Query: 1299 LGASGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMGVGTS---WNI 1469
LGA+ + +++ M + K + TC+ Y PT+L E + F + S + I
Sbjct: 491 LGAATRQNDYTDKYMEVGKGITNTCHESYIRAPTQLGPEAFRFSEAVEARALRSQEKYYI 550
Query: 1470 LRPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDVNTGE--KDNMMQ 1643
LRPET ES LWRLT + Y+DWGW+ A EK+ R GY GLR+V E KD++ Q
Sbjct: 551 LRPETFESYFVLWRLTHEQKYRDWGWEAVLALEKHCRTAHGYCGLRNVYQQEPQKDDVQQ 610
Query: 1644 SFFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVN 1766
SFFLAETLKYLYLLFS SV+ DEWVFNTEAHPL I N
Sbjct: 611 SFFLAETLKYLYLLFSDDSVLPLDEWVFNTEAHPLPIKGAN 651
>sw|O02773|M1A1_PIG Mannosyl-oligosaccharide 1,2-alpha-mannosidase IA
(EC 3.2.1.113) (Processing alpha-1,2-mannosidase IA)
(Alpha-1,2-mannosidase IA) (Mannosidase alpha class 1A
member 1) (Man(9)-alpha-mannosidase) (Man9-mannosidase).
Length = 659
Score = 382 bits (982), Expect = e-104
Identities = 205/462 (44%), Positives = 290/462 (62%), Gaps = 10/462 (2%)
Frame = +3
Query: 420 DPVNNEKREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGINS--FGGL-GATLVDSLDTL 590
D EKR K+KE M HAWN+Y YAWG +EL+P SK G +S FG + GAT+VD+LDTL
Sbjct: 195 DAAVREKRAKIKEMMKHAWNNYKLYAWGKNELKPVSKGGHSSSLFGNIKGATIVDALDTL 254
Query: 591 YIMGLKDEFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDI 770
+IM +K+EF++A+ WV E L+F+ + + SVFE IR +GGL+S+Y LSG+++F KA ++
Sbjct: 255 FIMKMKNEFEEAKAWVEEHLNFNVNAEVSVFEVNIRFIGGLISAYYLSGEEIFRKKAVEL 314
Query: 771 TDRLLPAWDTTSGIPYNRINLAHGRAHNPGW-TNGDSILADSGTEQLEFIALSQRTGDPK 947
+LLPA+ T SGIP+ +N+ G N W + G SILA+ GT LEFI LS +G+P
Sbjct: 315 GVKLLPAFYTPSGIPWALLNIKSGIGRNWPWASGGSSILAEFGTLHLEFIHLSYLSGNPF 374
Query: 948 YQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQGNK 1127
+ +K N+ K L + GL P Y+NP+SG ++ G +GDSFYEYLLK W+ +K
Sbjct: 375 FAEKVMNIRKVLNNLEKPQGLYPNYLNPNSGQWGQYHVSVGGLGDSFYEYLLKAWLMSDK 434
Query: 1128 TEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALGA 1307
T+ ++M+ +++ + + + + + YI E GG L KM L CFA GM ALGA
Sbjct: 435 TD--LEAKKMYFDAIKAIETHLIRKSRNGLTYIAEWKGGLLEHKMGHLTCFAGGMFALGA 492
Query: 1308 SGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMGVGTSWN----ILR 1475
++ + L E+ARTC+ Y T KL E + F G + + T N ILR
Sbjct: 493 DDAPDGLTQHYLQLGAEIARTCHESYSRTFVKLGPEAFRFDGGVE-AIATRQNEKYYILR 551
Query: 1476 PETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDVNTGEK--DNMMQSF 1649
PE VE+ +Y+WRLT + Y+ W W+ +A EK+ R+ GY GLRDV + D++ QSF
Sbjct: 552 PEVVETYLYMWRLTHDPKYRKWAWEAVEALEKHCRVNGGYSGLRDVYVSAQTYDDVQQSF 611
Query: 1650 FLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVNDNK 1775
FLAETLKYLYL+FS ++ + W+FNTEAHPL ++ N K
Sbjct: 612 FLAETLKYLYLIFSDDDLLPLEHWIFNTEAHPLPVLSRNIKK 653
>sw|Q9NR34|M1C1_HUMAN Mannosyl-oligosaccharide 1,2-alpha-mannosidase
IC (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IC)
(Alpha-1,2-mannosidase IC) (Mannosidase alpha class 1C
member 1) (HMIC).
Length = 630
Score = 382 bits (980), Expect = e-104
Identities = 209/467 (44%), Positives = 296/467 (63%), Gaps = 10/467 (2%)
Frame = +3
Query: 438 KREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGI--NSFGGL-GATLVDSLDTLYIMGLK 608
+REK+KE M AW SY +YA G +EL+P +K+G N FGGL GAT++DSLDTLY+M LK
Sbjct: 174 QREKIKEMMQFAWQSYKRYAMGKNELRPLTKDGYEGNMFGGLSGATVIDSLDTLYLMELK 233
Query: 609 DEFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDITDRLLP 788
+EFQ+A+ WV ES + +AS+FE IR +GGLLS++ L+G++VF KA + ++LLP
Sbjct: 234 EEFQEAKAWVGESFHLNVSGEASLFEVNIRYIGGLLSAFYLTGEEVFRIKAIRLGEKLLP 293
Query: 789 AWDTTSGIPYNRINLAHGRAHNPGW--TNGDSILADSGTEQLEFIALSQRTGDPKYQQKA 962
A++T +GIP ++ G N GW SILA+ G+ LEF+ L++ +G+ + +K
Sbjct: 294 AFNTPTGIPKGVVSFKSG---NWGWATAGSSSILAEFGSLHLEFLHLTELSGNQVFAEKV 350
Query: 963 ENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQGNKTEHVK 1142
N+ K L+KI GL P +++P SG ++ G +GDSFYEYL+K W+ KT+
Sbjct: 351 RNIRKVLRKIEKPFGLYPNFLSPVSGNWVQHHVSVGGLGDSFYEYLIKSWLMSGKTD--M 408
Query: 1143 HYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALGASGYGP 1322
+ M+ ++E + + +P YI E GG L KM LACF+ GM+ALGA
Sbjct: 409 EAKNMYYEALEAIETYLLNVSPGGLTYIAEWRGGILDHKMGHLACFSGGMIALGAEDAKE 468
Query: 1323 EKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQD---MGVGTSWNILRPETVES 1493
EK LA ++ +TC+ Y + TKL E ++F++G++ + S+ ILRPE VES
Sbjct: 469 EKRAHYRELAAQITKTCHESYARSDTKLGPEAFWFNSGREAVATQLSESYYILRPEVVES 528
Query: 1494 LMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDV--NTGEKDNMMQSFFLAETL 1667
MYLWR T N Y++WGW++ A EK R E+G+ G++DV +T DN QSFFLAETL
Sbjct: 529 YMYLWRQTHNPIYREWGWEVVLALEKYCRTEAGFSGIQDVYSSTPNHDNKQQSFFLAETL 588
Query: 1668 KYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVNDNKGIGTPVRQFGR 1808
KYLYLLFS ++S ++WVFNTEAHPL PVN + G R +GR
Sbjct: 589 KYLYLLFSEDDLLSLEDWVFNTEAHPL---PVNHSDSSG---RAWGR 629
>sptr|Q9W2W6|Q9W2W6 CG32684 protein.
Length = 643
Score = 380 bits (976), Expect = e-104
Identities = 203/458 (44%), Positives = 288/458 (62%), Gaps = 11/458 (2%)
Frame = +3
Query: 426 VNNEKREKVKEAMLHAWNSYVKYAWGMDELQPQSK--NGINSFGG--LGATLVDSLDTLY 593
++ EKR +V + M HAW++Y YAWG +EL+P S+ + + FG LGAT+VD LDTLY
Sbjct: 172 IDYEKRNQVVKMMEHAWHNYKLYAWGKNELRPLSQRPHSASIFGSYDLGATIVDGLDTLY 231
Query: 594 IMGLKDEFQKARDWVAESLDFDK-DYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDI 770
IMGL+ E+++ RDW+ D + SVFET IR VGG+L+ Y +GD ++ +KA+ +
Sbjct: 232 IMGLEKEYREGRDWIERKFSLDNISAELSVFETNIRFVGGMLTLYAFTGDPLYKEKAQHV 291
Query: 771 TDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTNG-DSILADSGTEQLEFIALSQRTGDPK 947
D+LLPA+ T +GIPY +N G A N GW +G SIL++ GT LEF LS TG+P
Sbjct: 292 ADKLLPAFQTPTGIPYALVNTKTGVAKNYGWASGGSSILSEFGTLHLEFAYLSDITGNPL 351
Query: 948 YQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQGNK 1127
Y+++ + + + L++I GL P ++NP +G ++ GA+GDS+YEYLLK W+Q +
Sbjct: 352 YRERVQTIRQVLKEIEKPKGLYPNFLNPKTGKWGQLHMSLGALGDSYYEYLLKAWLQSGQ 411
Query: 1128 TEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALGA 1307
T+ + R+M++ +M +L +T+P Y+ + L KMD LACF+ G+ ALGA
Sbjct: 412 TD--EEAREMFDEAMLAILDKMVRTSPGGLTYVSDLKFDRLEHKMDHLACFSGGLFALGA 469
Query: 1308 SGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMGVGTS---WNILRP 1478
+ + +++ M + K + TC+ Y PT+L E + F + S + ILRP
Sbjct: 470 ATRQNDYTDKYMEVGKGITNTCHESYIRAPTQLGPEAFRFSEAVEARALRSQEKYYILRP 529
Query: 1479 ETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDVNTGE--KDNMMQSFF 1652
ET ES LWRLT ++ Y+DWGW+ A EK+ R GY GLR+V E KD++ QSFF
Sbjct: 530 ETFESYFVLWRLTHDQKYRDWGWEAVLALEKHCRTAHGYCGLRNVYQQEPQKDDVQQSFF 589
Query: 1653 LAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVN 1766
LAETLKYLYLLFS SV+ DEWVFNTEAHPL I N
Sbjct: 590 LAETLKYLYLLFSDDSVLPLDEWVFNTEAHPLPIKGAN 627
>sw|P53625|M122_DROME Mannosyl-oligosaccharide alpha-1,2-mannosidase
isoform 2 (EC 3.2.1.113) (Man(9)-alpha-mannosidase).
Length = 643
Score = 380 bits (975), Expect = e-104
Identities = 203/458 (44%), Positives = 287/458 (62%), Gaps = 11/458 (2%)
Frame = +3
Query: 426 VNNEKREKVKEAMLHAWNSYVKYAWGMDELQPQSK--NGINSFGG--LGATLVDSLDTLY 593
++ EKR +V + M HAW++Y YAWG +EL+P S+ + + FG LGAT+VD LDTLY
Sbjct: 172 IDYEKRNQVVKMMEHAWHNYKLYAWGKNELRPLSQRPHSASIFGSYDLGATIVDGLDTLY 231
Query: 594 IMGLKDEFQKARDWVAESLDFDK-DYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDI 770
IMGL+ E+++ RDW+ D + SVFET IR VGG+L+ Y +GD ++ +KA+ +
Sbjct: 232 IMGLEKEYREGRDWIERKFSLDNISAELSVFETNIRFVGGMLTLYAFTGDPLYKEKAQHV 291
Query: 771 TDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTNG-DSILADSGTEQLEFIALSQRTGDPK 947
D+LLPA+ T +GIPY +N G A N GW +G SIL++ GT LEF LS TG+P
Sbjct: 292 ADKLLPAFQTPTGIPYALVNTKTGVAKNYGWASGGSSILSEFGTLHLEFAYLSDITGNPL 351
Query: 948 YQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQGNK 1127
Y+++ + + + L++I GL P ++NP +G ++ GA+GDS+YEYLLK W+Q +
Sbjct: 352 YRERVQTIRQVLKEIEKPKGLYPNFLNPKTGKWGQLHMSLGALGDSYYEYLLKAWLQSGQ 411
Query: 1128 TEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALGA 1307
T+ + R+M++ +M +L +T+P Y+ + L KMD LACF+ G+ ALGA
Sbjct: 412 TD--EEAREMFDEAMLAILDKMVRTSPGGLTYVSDLKFDRLEHKMDHLACFSGGLFALGA 469
Query: 1308 SGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMGVGTS---WNILRP 1478
+ + +++ M + K + TC+ Y PT+L E + F + S + ILRP
Sbjct: 470 ATRQNDYTDKYMEVGKGITNTCHESYIRAPTQLGPEAFRFSEAVEARALRSQEKYYILRP 529
Query: 1479 ETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDVNTGE--KDNMMQSFF 1652
ET ES LWRLT + Y+DWGW+ A EK+ R GY GLR+V E KD++ QSFF
Sbjct: 530 ETFESYFVLWRLTHEQKYRDWGWEAVLALEKHCRTAHGYCGLRNVYQQEPQKDDVQQSFF 589
Query: 1653 LAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVPVN 1766
LAETLKYLYLLFS SV+ DEWVFNTEAHPL I N
Sbjct: 590 LAETLKYLYLLFSDDSVLPLDEWVFNTEAHPLPIKGAN 627
>sw|P33908|M1A1_HUMAN Mannosyl-oligosaccharide 1,2-alpha-mannosidase
IA (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IA)
(Alpha-1,2-mannosidase IA) (Mannosidase alpha class 1A
member 1) (Man(9)-alpha-mannosidase) (Man9-mannosidase).
Length = 653
Score = 378 bits (970), Expect = e-103
Identities = 204/457 (44%), Positives = 286/457 (62%), Gaps = 10/457 (2%)
Frame = +3
Query: 420 DPVNNEKREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGINS--FGGL-GATLVDSLDTL 590
D EKR K+KE M HAWN+Y YAWG++EL+P SK G +S FG + GAT+VD+LDTL
Sbjct: 189 DAAIREKRAKIKEMMKHAWNNYKGYAWGLNELKPISKGGHSSSLFGNIKGATIVDALDTL 248
Query: 591 YIMGLKDEFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDI 770
+IM +K EF++A+ WV E+LDF+ + + SVFE IR VGGLLS+Y LSG+++F KA ++
Sbjct: 249 FIMEMKHEFEEAKSWVEENLDFNVNAEISVFEVNIRFVGGLLSAYYLSGEEIFRKKAVEL 308
Query: 771 TDRLLPAWDTTSGIPYNRINLAHGRAHNPGW-TNGDSILADSGTEQLEFIALSQRTGDPK 947
+LLPA+ T SGIP+ +N+ G N W + G SILA+ GT LEF+ LS +G+P
Sbjct: 309 GVKLLPAFHTPSGIPWALLNMKSGIGRNWPWASGGSSILAEFGTLHLEFMHLSHLSGNPI 368
Query: 948 YQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQGNK 1127
+ +K N+ L K+ GL P Y+NP SG ++ G +GDSFYEYLLK W+ +K
Sbjct: 369 FAEKVMNIRTVLNKLEKPQGLYPNYLNPSSGQWGQHHVSVGGLGDSFYEYLLKAWLMSDK 428
Query: 1128 TEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALGA 1307
T+ ++M+ +++ + + + + S YI E G L KM L CFA GM ALGA
Sbjct: 429 TD--LEAKKMYFDAVQAIETHLIRKSSSGLTYIAEWKRGLLEHKMGHLTCFAGGMFALGA 486
Query: 1308 SGYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMGVGTSWN----ILR 1475
++ + L E+ARTC+ Y T KL E + F G + + T N ILR
Sbjct: 487 DAAPEGMAQHYLELGAEIARTCHESYNRTFMKLGPEAFRFDGGVE-AIATRQNEKYYILR 545
Query: 1476 PETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDVNTGEK--DNMMQSF 1649
PE +E+ MY+WRLT + Y+ W W+ +A E + R+ GY GLRDV + D++ QSF
Sbjct: 546 PEVMETYMYMWRLTHDPKYRKWAWEAVEALENHCRVNGGYSGLRDVYLLHESYDDVQQSF 605
Query: 1650 FLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIVP 1760
FLAETLKYLYL+FS ++ + W+FN+EAH L I+P
Sbjct: 606 FLAETLKYLYLIFSDDDLLPLEHWIFNSEAHLLPILP 642
>sptr|Q8BTW7|Q8BTW7 Similar to alpha 1 (Fragment).
Length = 459
Score = 376 bits (966), Expect = e-103
Identities = 212/456 (46%), Positives = 291/456 (63%), Gaps = 15/456 (3%)
Frame = +3
Query: 432 NEKREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGINSFGGLGATLVDSLDTLYIMGLKD 611
NE+++ V EA LHAW Y K+AWG DEL+P SK + + GLG TL+D+LDT++I+GLK
Sbjct: 7 NERQKGVIEAFLHAWKGYQKFAWGHDELKPVSKT-FSEWFGLGLTLIDALDTMWILGLKQ 65
Query: 612 EFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDITDRLLPA 791
EF++AR WV+E+LDF K+ D ++FE+TIR++GGLLS+Y LSGD +FL KA+D RL+PA
Sbjct: 66 EFKQARKWVSENLDFQKNVDVNLFESTIRILGGLLSTYHLSGDSLFLTKAEDFGKRLMPA 125
Query: 792 WDTTSGIPYNRINLAHGRAHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQKAENV 971
+ T S IPY+ +N+ G AH+P WT+ DS +A+ + QLEF LS+ TG K+Q+ E V
Sbjct: 126 FTTPSKIPYSDVNIGTGFAHSPQWTS-DSTVAEVTSIQLEFRELSRLTGIKKFQEAVEEV 184
Query: 972 IKQLQKIY-PSDGLLPIYINPHSGTASY-STITFGAMGDSFYEYLLKVWIQGNKTEHVKH 1145
K + + DGL+P++IN +SG ++ T GA DS+YEYLLK WIQG K E
Sbjct: 185 TKHIHSLSGKKDGLVPMFINTNSGLFTHPGVFTLGARADSYYEYLLKQWIQGGKKE--TQ 242
Query: 1146 YRQMWETSMEGLLS-LTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALGASGYGP 1322
+ + ++EG+ + L +++ P ++ E G S KMD L CF PG LALG P
Sbjct: 243 LLEDYVKAIEGIKAHLLRQSQPRKLTFVGELAHGRFSAKMDHLVCFLPGTLALGVHHGLP 302
Query: 1323 EKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFF-------HTGQDMGVGTSWNILRPE 1481
M+LA+ L TCY Q T L+ E F H ++ N+LRPE
Sbjct: 303 ADH---MDLARALMETCYQMNQQMETGLSPEIAHFNMYPRADHKDVEVKPADRHNLLRPE 359
Query: 1482 TVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIES-GYVGLRDVNTGEKD---NMMQSF 1649
TVESL YL+R+T ++ YQDWGW+I Q+F K +R+ S GY + +V K + M+SF
Sbjct: 360 TVESLFYLYRVTRDRKYQDWGWEILQSFNKYTRVPSGGYSSINNVQNSHKPEPRDKMESF 419
Query: 1650 FLAETLKYLYLLFSPP-SVISFDEWVFNTEAHPLRI 1754
F+ ETLKYLYLLFS ++S D VFNTEAHPL I
Sbjct: 420 FVGETLKYLYLLFSDDLELLSLDSCVFNTEAHPLPI 455
>sptr|Q923C1|Q923C1 Similar to alpha 1,2-mannosidase (Fragment).
Length = 569
Score = 376 bits (966), Expect = e-103
Identities = 212/456 (46%), Positives = 291/456 (63%), Gaps = 15/456 (3%)
Frame = +3
Query: 432 NEKREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGINSFGGLGATLVDSLDTLYIMGLKD 611
NE+++ V EA LHAW Y K+AWG DEL+P SK + + GLG TL+D+LDT++I+GLK
Sbjct: 117 NERQKGVIEAFLHAWKGYQKFAWGHDELKPVSKT-FSEWFGLGLTLIDALDTMWILGLKQ 175
Query: 612 EFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDITDRLLPA 791
EF++AR WV+E+LDF K+ D ++FE+TIR++GGLLS+Y LSGD +FL KA+D RL+PA
Sbjct: 176 EFKQARKWVSENLDFQKNVDVNLFESTIRILGGLLSTYHLSGDSLFLTKAEDFGKRLMPA 235
Query: 792 WDTTSGIPYNRINLAHGRAHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQKAENV 971
+ T S IPY+ +N+ G AH+P WT+ DS +A+ + QLEF LS+ TG K+Q+ E V
Sbjct: 236 FTTPSKIPYSDVNIGTGFAHSPQWTS-DSTVAEVTSIQLEFRELSRLTGIKKFQEAVEEV 294
Query: 972 IKQLQKIY-PSDGLLPIYINPHSGTASY-STITFGAMGDSFYEYLLKVWIQGNKTEHVKH 1145
K + + DGL+P++IN +SG ++ T GA DS+YEYLLK WIQG K E
Sbjct: 295 TKHIHSLSGKKDGLVPMFINTNSGLFTHPGVFTLGARADSYYEYLLKQWIQGGKKE--TQ 352
Query: 1146 YRQMWETSMEGLLS-LTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALGASGYGP 1322
+ + ++EG+ + L +++ P ++ E G S KMD L CF PG LALG P
Sbjct: 353 LLEDYVKAIEGIKAHLLRQSQPRKLTFVGELAHGRFSAKMDHLVCFLPGTLALGVHHGLP 412
Query: 1323 EKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFF-------HTGQDMGVGTSWNILRPE 1481
M+LA+ L TCY Q T L+ E F H ++ N+LRPE
Sbjct: 413 ADH---MDLARALMETCYQMNQQMETGLSPEIAHFNMYPRADHKDVEVKPADRHNLLRPE 469
Query: 1482 TVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIES-GYVGLRDVNTGEKD---NMMQSF 1649
TVESL YL+R+T ++ YQDWGW+I Q+F K +R+ S GY + +V K + M+SF
Sbjct: 470 TVESLFYLYRVTRDRKYQDWGWEILQSFNKYTRVPSGGYSSINNVQNSHKPEPRDKMESF 529
Query: 1650 FLAETLKYLYLLFSPP-SVISFDEWVFNTEAHPLRI 1754
F+ ETLKYLYLLFS ++S D VFNTEAHPL I
Sbjct: 530 FVGETLKYLYLLFSDDLELLSLDSCVFNTEAHPLPI 565
>sw|Q9UKM7|M1B1_HUMAN Endoplasmic reticulum mannosyl-oligosaccharide
1,2-alpha-mannosidase (EC 3.2.1.113) (ER
alpha-1,2-mannosidase) (Mannosidase alpha class 1B member
1) (Man9GlcNAc2-specific processing alpha-mannosidase)
(UNQ747/PRO1477).
Length = 699
Score = 374 bits (961), Expect = e-102
Identities = 212/460 (46%), Positives = 296/460 (64%), Gaps = 16/460 (3%)
Frame = +3
Query: 423 PVNNEKREK-VKEAMLHAWNSYVKYAWGMDELQPQSKNGINSFGGLGATLVDSLDTLYIM 599
PV+ R+K V + LHAW Y K+AWG DEL+P S++ + + GLG TL+D+LDT++I+
Sbjct: 243 PVHLNYRQKGVIDVFLHAWKGYRKFAWGHDELKPVSRS-FSEWFGLGLTLIDALDTMWIL 301
Query: 600 GLKDEFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDITDR 779
GL+ EF++AR WV++ L F+KD D ++FE+TIR++GGLLS+Y LSGD +FL KA+D +R
Sbjct: 302 GLRKEFEEARKWVSKKLHFEKDVDVNLFESTIRILGGLLSAYHLSGDSLFLRKAEDFGNR 361
Query: 780 LLPAWDTTSGIPYNRINLAHGRAHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQK 959
L+PA+ T S IPY+ +N+ G AH P WT+ DS +A+ + QLEF LS+ TGD K+Q+
Sbjct: 362 LMPAFRTPSKIPYSDVNIGTGVAHPPRWTS-DSTVAEVTSIQLEFRELSRLTGDKKFQEA 420
Query: 960 AENVIKQLQKIY-PSDGLLPIYINPHSGTASY-STITFGAMGDSFYEYLLKVWIQGNKTE 1133
E V + + + DGL+P++IN HSG ++ T GA DS+YEYLLK WIQG K E
Sbjct: 421 VEKVTQHIHGLSGKKDGLVPMFINTHSGLFTHLGVFTLGARADSYYEYLLKQWIQGGKQE 480
Query: 1134 HVKHYRQMWETSMEGLLS-LTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALGAS 1310
+ + ++EG+ + L + + PS ++ E G S KMD L CF PG LALG
Sbjct: 481 --TQLLEDYVEAIEGVRTHLLRHSEPSKLTFVGELAHGRFSAKMDHLVCFLPGTLALGVY 538
Query: 1311 GYGPEKSEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMG-------VGTSWNI 1469
P M LA+EL TCY + T L+ E F+ G N+
Sbjct: 539 HGLPASH---MELAQELMETCYQMNRQMETGLSPEIVHFNLYPQPGRRDVEVKPADRHNL 595
Query: 1470 LRPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIES-GYVGLRDVNTGEKD---NM 1637
LRPETVESL YL+R+TG++ YQDWGW+I Q+F + +R+ S GY + +V +K +
Sbjct: 596 LRPETVESLFYLYRVTGDRKYQDWGWEILQSFSRFTRVPSGGYSSINNVQDPQKPEPRDK 655
Query: 1638 MQSFFLAETLKYLYLLFS-PPSVISFDEWVFNTEAHPLRI 1754
M+SFFL ETLKYL+LLFS P+++S D +VFNTEAHPL I
Sbjct: 656 MESFFLGETLKYLFLLFSDDPNLLSLDAYVFNTEAHPLPI 695
>sptr|Q18788|Q18788 C52E4.5 protein.
Length = 590
Score = 370 bits (949), Expect = e-101
Identities = 201/457 (43%), Positives = 290/457 (63%), Gaps = 11/457 (2%)
Frame = +3
Query: 420 DPVNNEKREKVKEAMLHAWNSYVKYAWGMDELQPQSK--NGINSFGG--LGATLVDSLDT 587
D N+ +R+KVKE M+HAW Y Y+WG +EL+P SK N N FGG + AT+VD+ DT
Sbjct: 133 DEENDLRRQKVKEMMIHAWEGYKNYSWGANELRPMSKKPNSQNIFGGSQMPATIVDAADT 192
Query: 588 LYIMGLKDEFQKARDWVAESLDFDKDYDA-SVFETTIRVVGGLLSSYDLSGDKVFLDKAK 764
L+IM LKD++++ARD++ + K SVFETTIR +GGLLS Y L+ + +++KA+
Sbjct: 193 LFIMDLKDKYKEARDYIENNFSMAKSTSTLSVFETTIRFLGGLLSLYALTQESFYIEKAR 252
Query: 765 DITDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTNG-DSILADSGTEQLEFIALSQRTGD 941
++ + LLPA++T SGIP + +++A A N GW NG SIL++ G+ LEF+ LS+ +
Sbjct: 253 EVGEALLPAFNTPSGIPKSNLDVASKHASNYGWANGGQSILSEIGSLHLEFLYLSRISNA 312
Query: 942 PKYQQKAENVIKQLQKIYPSDGLLPIYINPHSGTASYSTITFGAMGDSFYEYLLKVWIQG 1121
P +++K + V L+K +GL YINP +G + S ++ GA+GDSFYEYL+K ++Q
Sbjct: 313 PIFEKKVKKVRDALEKAEKPNGLYSNYINPDTGKFTGSHMSLGALGDSFYEYLIKSYVQS 372
Query: 1122 NKTEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLAL 1301
N T+ + W+ S + K + SN Y E N G KM LACF PGM AL
Sbjct: 373 NYTD-TQAKNMYWDVSDAIQKHMIKVSKQSNLTYTVELNNGQAQHKMGHLACFVPGMFAL 431
Query: 1302 GASGYGPEKSE-QIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMGVGTSWN--IL 1472
A E+ + +IM LA+ELA+TC+ Y + T + E ++F+ + S N I
Sbjct: 432 QAINEDTEEEKLRIMTLAEELAKTCHESYIRSETHIGPEMFYFNERDEATSKHSENGYIQ 491
Query: 1473 RPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDVNTGE--KDNMMQS 1646
RPE +E YLWRLTG Y+DW WD QA EK R++SG+ GL++V + ++++MQS
Sbjct: 492 RPEVIEGWFYLWRLTGKTMYRDWVWDAVQAIEKYCRVDSGFTGLQNVYNPKAGREDVMQS 551
Query: 1647 FFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRIV 1757
FFLAE LKY YL F+ S+IS D+WVFNTEAHP+ ++
Sbjct: 552 FFLAEFLKYAYLTFADESLISLDKWVFNTEAHPVPVL 588
>sptr|Q9VAP8|Q9VAP8 CG11874 protein (SD05769P).
Length = 685
Score = 369 bits (948), Expect = e-101
Identities = 213/492 (43%), Positives = 304/492 (61%), Gaps = 26/492 (5%)
Frame = +3
Query: 366 ENPVVSNKNSRKDLVVEIDPV----------NNEKREKVKEAMLHAWNSYVKYAWGMDEL 515
+NP+ S K+ + +I P+ NE++ V A H+W Y KYAWG D L
Sbjct: 201 KNPLTLGGVSFKEHLKKIKPILAKGQHFHGATNERQSAVVAAFKHSWAGYKKYAWGHDNL 260
Query: 516 QPQSKNGINSFGGLGATLVDSLDTLYIMGLKDEFQKARDWVAESLDFDKDYDASVFETTI 695
+P S+ FG LG T+VDSLDT+YIMGL DEF++ RDWV +SL FD D ++FE TI
Sbjct: 261 KPISQYSHEWFG-LGLTIVDSLDTMYIMGLDDEFKEGRDWVEQSLRFDTKRDVNLFEVTI 319
Query: 696 RVVGGLLSSYDLSGDKVFLDKAKDITDRLLPAWDTTSGIPYNRINLAHGRAHNPGWTNGD 875
RV+GGLLS+Y LSGD +FL KA ++ +RLLPA+ + S IPY+ +NL AH+P W + D
Sbjct: 320 RVLGGLLSAYHLSGDTMFLAKAAELGNRLLPAFQSPSNIPYSDVNLGDLSAHSPKW-SPD 378
Query: 876 SILADSGTEQLEFIALSQRTGDPKYQQKAENVIKQLQKIYPSDGLLPIYINPHSGT-ASY 1052
S ++ T QLEF LS+ T Y+Q A V +++ + + GL+PI+IN ++GT +Y
Sbjct: 379 SSTSEVTTIQLEFRDLSRSTNISIYEQVAHKVNEKVHDLEKNHGLVPIFINANTGTFRNY 438
Query: 1053 STITFGAMGDSFYEYLLKVWIQGNKTEH---VKHYRQMWETSMEGLLSLTKKTTPSNYY- 1220
+TI+ GA GDS+YEYLLK WIQ + ++ + Y Q +++G+L+ + TP ++
Sbjct: 439 ATISLGARGDSYYEYLLKQWIQTGRKDNDNLILDYMQ----AVDGVLTQLMRRTPREHWV 494
Query: 1221 YICEK-NGGSLSDKMDELACFAPGMLALGASGYGPEKSEQIMNLAKELARTCYNFYQTTP 1397
YI E NG KMD L C+ PG L LG P+ + LA++L TCY Y P
Sbjct: 495 YIGELINGKDFKPKMDHLTCYLPGTLILGHQNGMPDSH---LILARDLLDTCYQTYMMNP 551
Query: 1398 TKLAGE-NYFFHTGQD-----MGVGTSWNILRPETVESLMYLWRLTGNKTYQDWGWDIFQ 1559
T LA E +YF T +D + + N+LRPE VESL Y + +TGN+TYQD GW IFQ
Sbjct: 552 THLAAEISYFALTEKDDQDIYVKPNDAHNLLRPEFVESLYYFYSITGNRTYQDMGWKIFQ 611
Query: 1560 AFEKNSRIESGYVGLRDVNTGEKD---NMMQSFFLAETLKYLYLLFSPP-SVISFDEWVF 1727
AFE ++++ +GY + +V + ++M+SF+++ETLKY YLLFS I ++WVF
Sbjct: 612 AFETHAKVNAGYTSMGNVKNTQSTRLRDLMESFWMSETLKYFYLLFSDDRKEIDLEQWVF 671
Query: 1728 NTEAHPLRIVPV 1763
N+E HPL + P+
Sbjct: 672 NSEGHPLPVYPL 683
>sptr|Q7QK05|Q7QK05 AgCP2314 (Fragment).
Length = 582
Score = 362 bits (928), Expect = 2e-98
Identities = 207/462 (44%), Positives = 305/462 (66%), Gaps = 15/462 (3%)
Frame = +3
Query: 432 NEKREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGINSFGGLGATLVDSLDTLYIMGLKD 611
N+++ V +A+LH+W Y +YAWG D L+P S G + + GLG TLVDSLDTLYIM L+D
Sbjct: 132 NDRQRAVVDAVLHSWKGYKEYAWGHDNLKPISM-GYSDWFGLGLTLVDSLDTLYIMDLQD 190
Query: 612 EFQKARDWVAESLDFDKDYDASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDITDRLLPA 791
E+ +AR WV + L FD + + ++FE TIRVVGGLLS+Y LSGD++FLDKA D+ +RLLP
Sbjct: 191 EYDEARAWVEKYLKFDINREVNLFEVTIRVVGGLLSAYHLSGDRMFLDKAIDLGNRLLPC 250
Query: 792 WDTTSGIPYNRINLAHGRAHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQKAENV 971
+D+ SGIP++ +N+ AH P W + DS ++ T QLEF LS+ + +P +++ A V
Sbjct: 251 FDSPSGIPFSDVNIGSLAAHAPKW-SPDSSTSEVTTIQLEFRDLSRASQNPVFEKVASRV 309
Query: 972 IKQLQKIYPSDGLLPIYINPHSGT-ASYSTITFGAMGDSFYEYLLKVWIQ-GNKTEHVKH 1145
++ ++ ++GL+PI+IN ++G +++T++ GA DS+YEYLLK W+Q G K + +
Sbjct: 310 NLKIHELDKNEGLVPIFINANTGQFRNFATVSLGARADSYYEYLLKQWLQTGKKAD--DY 367
Query: 1146 YRQMWETSMEGLLSLTKKTTPS-NYYYICE-KNGGSLSDKMDELACFAPGMLALGASGYG 1319
+ ++ SM G+L+ +TTP+ + YI E NG KMD L C+ PG L LG
Sbjct: 368 LIEDYQKSMRGVLNQLVRTTPNEKHVYIGELLNGKDFKPKMDHLTCYLPGTLLLGYKNGM 427
Query: 1320 PEKSEQIMNLAKELARTCYNFYQTTPTKLAGE-NYFFHTGQ---DMGVGT--SWNILRPE 1481
P+ + LA +L TCY Y PT+LA E +YF G+ D+ V T + N+LRPE
Sbjct: 428 PKTH---LRLATDLLETCYQTYMKQPTQLAPEISYFNVNGESETDIYVKTNDAHNLLRPE 484
Query: 1482 TVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDV----NTGEKDNMMQSF 1649
VESL Y + +TGN+TYQD GW IF+AF + +++++GY + +V NT +D MM+SF
Sbjct: 485 FVESLYYFYAITGNRTYQDMGWTIFEAFNRYTKVKNGYTSIGNVKNPLNTRPRD-MMESF 543
Query: 1650 FLAETLKYLYLLFSPP-SVISFDEWVFNTEAHPLRIVPVNDN 1772
+L ETLKY YLLFS + I D++VFN+EAH ++P+ D+
Sbjct: 544 WLGETLKYFYLLFSDDRNEIDLDKYVFNSEAH---LLPIRDD 582
>sptr|P90787|P90787 D2030.1 protein.
Length = 540
Score = 339 bits (869), Expect = 1e-91
Identities = 191/456 (41%), Positives = 278/456 (60%), Gaps = 17/456 (3%)
Frame = +3
Query: 438 KREKVKEAMLHAWNSYVKYAWGMDELQPQSKNGINS----FGGLGATLVDSLDTLYIMGL 605
+R VK+ + AW+ Y KYAWG +EL+P S++G +S +G GAT++D++DTLY++GL
Sbjct: 89 RRIFVKQMIKFAWDGYRKYAWGENELRPNSRSGHSSSIFGYGKTGATIIDAIDTLYLVGL 148
Query: 606 KDEFQKARDWVAESLDFDKDY--DASVFETTIRVVGGLLSSYDLSGDKVFLDKAKDITDR 779
K+E+++ARDW+A+ DF D SVFET IR GGLLS++ L+GDK+FL KA+D+
Sbjct: 149 KEEYKEARDWIAD-FDFKTSAKGDLSVFETNIRFTGGLLSAFALTGDKMFLKKAEDVATI 207
Query: 780 LLPAWDTTSGIPYNRINLAHGRAHNPGWTNGDSILADSGTEQLEFIALSQRTGDPKYQQK 959
LLPA++T SGIP + I+ GR+ W +G +IL++ G+ QLEF LS TG+P + QK
Sbjct: 208 LLPAFETPSGIPNSLIDAQTGRSKTYSWASGKAILSEYGSIQLEFDYLSNLTGNPVFAQK 267
Query: 960 AENVIKQLQKIYPSDGLLPIYIN----PHSGTASYSTITFGAMGDSFYEYLLKVWI-QGN 1124
A+ + L + +GL PIYI P G +S GAM DS+YEYLLK WI G
Sbjct: 268 ADKIRDVLTAMEKPEGLYPIYITMDNPPRWGQHLFS---MGAMADSWYEYLLKQWIATGK 324
Query: 1125 KTEHVKHYRQMWETSMEGLLSLTKKTTPSNYYYICEKNGGSLSDKMDELACFAPGMLALG 1304
K + K + +ME + K+ SN +Y + NG + + LACF+ GM+ L
Sbjct: 325 KDDRTKREYEEAIFAMEKRMLF--KSEQSNLWYFAKMNGNRMEHSFEHLACFSGGMVVLH 382
Query: 1305 ASGYGPEK-SEQIMNLAKELARTCYNFYQTTPTKLAGENYFFHTGQDMGV---GTSWNIL 1472
A + S+ M L KE+ TC+ Y + T + E++ F + + S+ IL
Sbjct: 383 AMNEKNKTISDHYMTLGKEIGHTCHESYARSTTGIGPESFQFTSSVEAKTERRQDSYYIL 442
Query: 1473 RPETVESLMYLWRLTGNKTYQDWGWDIFQAFEKNSRIESGYVGLRDV--NTGEKDNMMQS 1646
RPE VE+ YLWR T ++ Y+ W WD Q E+ + +GY G+R+V ++ E+D++ QS
Sbjct: 443 RPEVVETWFYLWRATKDEKYRQWAWDHVQNLEEYCKGTAGYSGIRNVYESSPEQDDVQQS 502
Query: 1647 FFLAETLKYLYLLFSPPSVISFDEWVFNTEAHPLRI 1754
F AE KYLYL+FS +++ D+WVFNTEAHP RI
Sbjct: 503 FLFAELFKYLYLIFSEDNILPLDQWVFNTEAHPFRI 538
Database: swall
Posted date: Feb 26, 2004 3:00 PM
Number of letters in database: 439,479,560
Number of sequences in database: 1,381,838
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,830,595,587
Number of Sequences: 1381838
Number of extensions: 41985695
Number of successful extensions: 115844
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 104960
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 115248
length of database: 439,479,560
effective HSP length: 131
effective length of database: 258,458,782
effective search space used: 159727527276
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

