BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
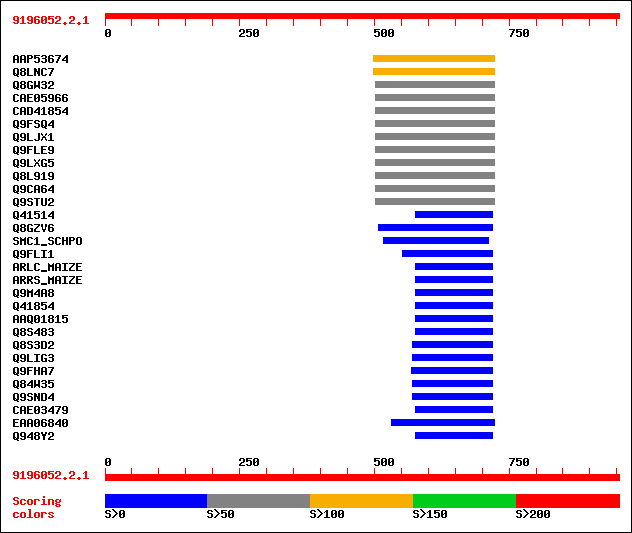
Query= 9196052.2.1
(954 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptrnew|AAP53674|AAP53674 Hypothetical protein. 107 2e-22
sptr|Q8LNC7|Q8LNC7 Hypothetical protein similar to putative DNA ... 107 2e-22
sptr|Q8GW32|Q8GW32 Hypothetical protein (At1g26948). 93 6e-18
sptrnew|CAE05966|CAE05966 OSJNBa0063C18.7 protein. 79 1e-13
sptrnew|CAD41854|CAD41854 OSJNBb0079B02.13 protein. 79 1e-13
sptr|Q9FSQ4|Q9FSQ4 Putative DNA binding protein. 79 1e-13
sptr|Q9LJX1|Q9LJX1 DNA-binding protein-like. 78 2e-13
sptr|Q9FLE9|Q9FLE9 DNA-binding protein-like. 77 4e-13
sptr|Q9LXG5|Q9LXG5 Hypothetical protein (Putative bHLH protein). 76 8e-13
sptr|Q8L919|Q8L919 DNA-binding protein-like. 76 8e-13
sptr|Q9CA64|Q9CA64 Putative DNA-binding protein (At1g74500). 74 3e-12
sptr|Q9STU2|Q9STU2 Hypothetical 8.8 kDa protein. 69 7e-11
sptr|Q41514|Q41514 Myc-like regulatory R gene product (Fragment). 37 0.31
sptr|Q8GZV6|Q8GZV6 Hypothetical protein. 36 0.91
sw|O94383|SMC1_SCHPO Structural maintenance of chromosome 1 (Coh... 35 1.5
sptr|Q9FLI1|Q9FLI1 Genomic DNA, chromosome 5, P1 clone:MIO24. 35 1.5
sw|P13526|ARLC_MAIZE Anthocyanin regulatory Lc protein. 34 2.6
sw|P13027|ARRS_MAIZE Anthocyanin regulatory R-S protein. 34 2.6
sptr|Q9M4A8|Q9M4A8 Transcription factor. 34 2.6
sptr|Q41854|Q41854 SN. 34 2.6
sptrnew|AAQ01815|AAQ01815 BHLH protein. 34 2.6
sptr|Q8S483|Q8S483 Regulatory protein. 34 2.6
sptr|Q8S3D2|Q8S3D2 Putative bHLH transcription factor. 33 4.5
sptr|Q9LIG3|Q9LIG3 Emb|CAB62312.1. 33 4.5
sptr|Q9FHA7|Q9FHA7 Emb|CAB62312.1 (Putative bHLH transcription f... 33 5.9
sptr|Q84W35|Q84W35 Hypothetical protein. 33 7.7
sptr|Q9SND4|Q9SND4 Hypothetical 25.7 kDa protein. 33 7.7
sptrnew|CAE03479|CAE03479 OSJNBa0065O17.4 protein. 33 7.7
sptrnew|EAA06840|EAA06840 AgCP7373 (Fragment). 33 7.7
sptr|Q948Y2|Q948Y2 R-type basic helix-loop-helix protein. 33 7.7
>sptrnew|AAP53674|AAP53674 Hypothetical protein.
Length = 88
Score = 107 bits (268), Expect = 2e-22
Identities = 58/77 (75%), Positives = 69/77 (89%), Gaps = 2/77 (2%)
Frame = -1
Query: 723 EEEINELISRLQTLLP-SARRRGG-SQASTTKLLKETCSYIKSLHREVDDLSDRLSDLMA 550
++EINELIS+LQ+LLP S+RRRG S++ TKLLKE CSYIKSLHREVDDLS+RLS+LMA
Sbjct: 12 DDEINELISKLQSLLPESSRRRGATSRSPATKLLKEMCSYIKSLHREVDDLSERLSELMA 71
Query: 549 SMDHNSPGAEIIRSLLR 499
+MD NSP A+IIRSLLR
Sbjct: 72 TMDSNSPQADIIRSLLR 88
>sptr|Q8LNC7|Q8LNC7 Hypothetical protein similar to putative DNA
binding proteins.
Length = 88
Score = 107 bits (268), Expect = 2e-22
Identities = 58/77 (75%), Positives = 69/77 (89%), Gaps = 2/77 (2%)
Frame = -1
Query: 723 EEEINELISRLQTLLP-SARRRGG-SQASTTKLLKETCSYIKSLHREVDDLSDRLSDLMA 550
++EINELIS+LQ+LLP S+RRRG S++ TKLLKE CSYIKSLHREVDDLS+RLS+LMA
Sbjct: 12 DDEINELISKLQSLLPESSRRRGATSRSPATKLLKEMCSYIKSLHREVDDLSERLSELMA 71
Query: 549 SMDHNSPGAEIIRSLLR 499
+MD NSP A+IIRSLLR
Sbjct: 72 TMDSNSPQADIIRSLLR 88
>sptr|Q8GW32|Q8GW32 Hypothetical protein (At1g26948).
Length = 94
Score = 92.8 bits (229), Expect = 6e-18
Identities = 44/74 (59%), Positives = 61/74 (82%)
Frame = -1
Query: 723 EEEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDLSDRLSDLMASM 544
+++I++L+S+LQ L+P RRR + S +K+L+ETC+YI++LHREVDDLSDRLS+L+AS
Sbjct: 19 DDQISDLVSKLQHLIPELRRRRSDKVSASKVLQETCNYIRNLHREVDDLSDRLSELLAST 78
Query: 543 DHNSPGAEIIRSLL 502
D NS A IIRSLL
Sbjct: 79 DDNSAEAAIIRSLL 92
>sptrnew|CAE05966|CAE05966 OSJNBa0063C18.7 protein.
Length = 104
Score = 78.6 bits (192), Expect = 1e-13
Identities = 40/75 (53%), Positives = 58/75 (77%), Gaps = 1/75 (1%)
Frame = -1
Query: 723 EEEINELISRLQTLLPSARRRGGS-QASTTKLLKETCSYIKSLHREVDDLSDRLSDLMAS 547
E++I EL+S+LQ LLP ++ R G+ + S ++L+ETCSYI+SLH+EVD+LS+ L+ L+AS
Sbjct: 29 EDQIAELLSKLQALLPESQARNGAHRGSAARVLQETCSYIRSLHQEVDNLSETLAQLLAS 88
Query: 546 MDHNSPGAEIIRSLL 502
D S A +IRSLL
Sbjct: 89 PDVTSDQAAVIRSLL 103
>sptrnew|CAD41854|CAD41854 OSJNBb0079B02.13 protein.
Length = 104
Score = 78.6 bits (192), Expect = 1e-13
Identities = 40/75 (53%), Positives = 58/75 (77%), Gaps = 1/75 (1%)
Frame = -1
Query: 723 EEEINELISRLQTLLPSARRRGGS-QASTTKLLKETCSYIKSLHREVDDLSDRLSDLMAS 547
E++I EL+S+LQ LLP ++ R G+ + S ++L+ETCSYI+SLH+EVD+LS+ L+ L+AS
Sbjct: 29 EDQIAELLSKLQALLPESQARNGAHRGSAARVLQETCSYIRSLHQEVDNLSETLAQLLAS 88
Query: 546 MDHNSPGAEIIRSLL 502
D S A +IRSLL
Sbjct: 89 PDVTSDQAAVIRSLL 103
>sptr|Q9FSQ4|Q9FSQ4 Putative DNA binding protein.
Length = 104
Score = 78.6 bits (192), Expect = 1e-13
Identities = 40/75 (53%), Positives = 58/75 (77%), Gaps = 1/75 (1%)
Frame = -1
Query: 723 EEEINELISRLQTLLPSARRRGGS-QASTTKLLKETCSYIKSLHREVDDLSDRLSDLMAS 547
E++I EL+S+LQ LLP ++ R G+ + S ++L+ETCSYI+SLH+EVD+LS+ L+ L+AS
Sbjct: 29 EDQIAELLSKLQALLPESQARNGAHRGSAARVLQETCSYIRSLHQEVDNLSETLAQLLAS 88
Query: 546 MDHNSPGAEIIRSLL 502
D S A +IRSLL
Sbjct: 89 PDVTSDQAAVIRSLL 103
>sptr|Q9LJX1|Q9LJX1 DNA-binding protein-like.
Length = 92
Score = 78.2 bits (191), Expect = 2e-13
Identities = 38/75 (50%), Positives = 57/75 (76%), Gaps = 1/75 (1%)
Frame = -1
Query: 723 EEEINELISRLQTLLPSAR-RRGGSQASTTKLLKETCSYIKSLHREVDDLSDRLSDLMAS 547
++++ +L+S+L+ LP RR + S +K+L+ETC+YI+ LHREVD+LSDRLS L+ S
Sbjct: 17 DDQMIDLVSKLRQFLPEIHERRRSDKVSASKVLQETCNYIRKLHREVDNLSDRLSQLLDS 76
Query: 546 MDHNSPGAEIIRSLL 502
+D +SP A +IRSLL
Sbjct: 77 VDEDSPEAAVIRSLL 91
>sptr|Q9FLE9|Q9FLE9 DNA-binding protein-like.
Length = 92
Score = 77.0 bits (188), Expect = 4e-13
Identities = 37/75 (49%), Positives = 60/75 (80%), Gaps = 1/75 (1%)
Frame = -1
Query: 723 EEEINELISRLQTLLPS-ARRRGGSQASTTKLLKETCSYIKSLHREVDDLSDRLSDLMAS 547
+ ++ +L+S+L+ +LP +RR +AS +K+L+ETC+YI++L+REVD+LS+RLS L+ S
Sbjct: 17 DNQMIDLVSKLRQILPEIGQRRRSDKASASKVLQETCNYIRNLNREVDNLSERLSQLLES 76
Query: 546 MDHNSPGAEIIRSLL 502
+D +SP A +IRSLL
Sbjct: 77 VDEDSPEAAVIRSLL 91
>sptr|Q9LXG5|Q9LXG5 Hypothetical protein (Putative bHLH protein).
Length = 94
Score = 75.9 bits (185), Expect = 8e-13
Identities = 36/75 (48%), Positives = 58/75 (77%), Gaps = 1/75 (1%)
Frame = -1
Query: 723 EEEINELISRLQTLLPSARR-RGGSQASTTKLLKETCSYIKSLHREVDDLSDRLSDLMAS 547
+++I +LIS+L+ +P R+ R + S +K+L+ETC+YI++L++E DDLSDRL+ L+ S
Sbjct: 18 DDQITDLISKLRQSIPEIRQNRRSNTVSASKVLQETCNYIRNLNKEADDLSDRLTQLLES 77
Query: 546 MDHNSPGAEIIRSLL 502
+D NSP A +IRSL+
Sbjct: 78 IDPNSPQAAVIRSLI 92
>sptr|Q8L919|Q8L919 DNA-binding protein-like.
Length = 92
Score = 75.9 bits (185), Expect = 8e-13
Identities = 37/75 (49%), Positives = 56/75 (74%), Gaps = 1/75 (1%)
Frame = -1
Query: 723 EEEINELISRLQTLLPSAR-RRGGSQASTTKLLKETCSYIKSLHREVDDLSDRLSDLMAS 547
++++ +L+S+L+ LP RR + S + +L+ETC+YI+ LHREVD+LSDRLS L+ S
Sbjct: 17 DDQMIDLVSKLRQFLPEIHERRRSDKVSASXVLQETCNYIRKLHREVDNLSDRLSQLLDS 76
Query: 546 MDHNSPGAEIIRSLL 502
+D +SP A +IRSLL
Sbjct: 77 VDEDSPEAAVIRSLL 91
>sptr|Q9CA64|Q9CA64 Putative DNA-binding protein (At1g74500).
Length = 93
Score = 73.9 bits (180), Expect = 3e-12
Identities = 39/75 (52%), Positives = 57/75 (76%), Gaps = 1/75 (1%)
Frame = -1
Query: 723 EEEINELISRLQTLLPSAR-RRGGSQASTTKLLKETCSYIKSLHREVDDLSDRLSDLMAS 547
E++IN+LI +LQ LLP R R + S ++L++TC+YI++LHREVDDLS+RLS+L+A+
Sbjct: 19 EDQINDLIIKLQQLLPELRDSRRSDKVSAARVLQDTCNYIRNLHREVDDLSERLSELLAN 78
Query: 546 MDHNSPGAEIIRSLL 502
D + A +IRSLL
Sbjct: 79 SD--TAQAALIRSLL 91
>sptr|Q9STU2|Q9STU2 Hypothetical 8.8 kDa protein.
Length = 78
Score = 69.3 bits (168), Expect = 7e-11
Identities = 38/75 (50%), Positives = 54/75 (72%), Gaps = 1/75 (1%)
Frame = -1
Query: 723 EEEINELISRLQTLLPS-ARRRGGSQASTTKLLKETCSYIKSLHREVDDLSDRLSDLMAS 547
+E+IN+L+ +L LLP A R + S +++L+ETCSYI++L +EVDDLS+RLS L+ S
Sbjct: 4 DEQINDLVLQLHRLLPELANNRRSGKVSASRVLQETCSYIRNLSKEVDDLSERLSQLLES 63
Query: 546 MDHNSPGAEIIRSLL 502
D S A +IRSLL
Sbjct: 64 TD--SAQAALIRSLL 76
>sptr|Q41514|Q41514 Myc-like regulatory R gene product (Fragment).
Length = 146
Score = 37.4 bits (85), Expect = 0.31
Identities = 22/48 (45%), Positives = 31/48 (64%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDL 577
E++NE+ L++LLPS R G QAS +L ET +Y+K L R V +L
Sbjct: 12 EKLNEMFLVLKSLLPSIHR--GEQAS---ILAETIAYLKELQRRVQEL 54
>sptr|Q8GZV6|Q8GZV6 Hypothetical protein.
Length = 776
Score = 35.8 bits (81), Expect = 0.91
Identities = 22/71 (30%), Positives = 39/71 (54%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDLSDRLSDLMASMD 541
E+INE + LQ L+P + G +L E +Y++SL R+V+ LS +LS + ++
Sbjct: 648 EKINERMKLLQDLVPGCNKITGK----AMMLDEIINYVQSLQRQVEFLSMKLSTISPELN 703
Query: 540 HNSPGAEIIRS 508
+ +I+ S
Sbjct: 704 SDLDLQDILCS 714
>sw|O94383|SMC1_SCHPO Structural maintenance of chromosome 1
(Cohesin complex subunit psm1) (Chromosome segregation
protein smc1).
Length = 1233
Score = 35.0 bits (79), Expect = 1.5
Identities = 20/65 (30%), Positives = 34/65 (52%)
Frame = -1
Query: 711 NELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDLSDRLSDLMASMDHNS 532
+ L+ +LQTL + + S+ S T ++ + S IKSLH V L +DL+A ++
Sbjct: 395 SNLLFKLQTLNRNIKVTSQSKDSLTSIVGDLESKIKSLHESVSSLDTERADLLAKINEKI 454
Query: 531 PGAEI 517
E+
Sbjct: 455 ESLEL 459
>sptr|Q9FLI1|Q9FLI1 Genomic DNA, chromosome 5, P1 clone:MIO24.
Length = 174
Score = 35.0 bits (79), Expect = 1.5
Identities = 21/56 (37%), Positives = 34/56 (60%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDLSDRLSDLM 553
+E+ L + L++LLP +G + ST+ + E +YIK L R++ +LS R DLM
Sbjct: 15 QEMASLYASLRSLLPLHFIKG--KRSTSDQVNEAVNYIKYLQRKIKELSVRRDDLM 68
>sw|P13526|ARLC_MAIZE Anthocyanin regulatory Lc protein.
Length = 610
Score = 34.3 bits (77), Expect = 2.6
Identities = 18/48 (37%), Positives = 28/48 (58%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDL 577
E++NE+ L++LLPS R + +L ET +Y+K L R V +L
Sbjct: 426 EKLNEMFLVLKSLLPSIHR-----VNKASILAETIAYLKELQRRVQEL 468
>sw|P13027|ARRS_MAIZE Anthocyanin regulatory R-S protein.
Length = 612
Score = 34.3 bits (77), Expect = 2.6
Identities = 18/48 (37%), Positives = 28/48 (58%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDL 577
E++NE+ L++LLPS R + +L ET +Y+K L R V +L
Sbjct: 428 EKLNEMFLVLKSLLPSIHR-----VNKASILAETIAYLKELQRRVQEL 470
>sptr|Q9M4A8|Q9M4A8 Transcription factor.
Length = 611
Score = 34.3 bits (77), Expect = 2.6
Identities = 18/48 (37%), Positives = 28/48 (58%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDL 577
E++NE+ L++LLPS R + +L ET +Y+K L R V +L
Sbjct: 428 EKLNEMFLVLKSLLPSIHR-----VNKASILAETIAYLKELQRRVQEL 470
>sptr|Q41854|Q41854 SN.
Length = 616
Score = 34.3 bits (77), Expect = 2.6
Identities = 18/48 (37%), Positives = 28/48 (58%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDL 577
E++NE+ L++LLPS R + +L ET +Y+K L R V +L
Sbjct: 432 EKLNEMFLVLKSLLPSIHR-----VNKASILAETIAYLKELQRRVQEL 474
>sptrnew|AAQ01815|AAQ01815 BHLH protein.
Length = 609
Score = 34.3 bits (77), Expect = 2.6
Identities = 18/48 (37%), Positives = 28/48 (58%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDL 577
E++NE+ L++LLPS R + +L ET +Y+K L R V +L
Sbjct: 425 EKLNEMFLVLKSLLPSIHR-----VNKASILAETIAYLKELQRRVQEL 467
>sptr|Q8S483|Q8S483 Regulatory protein.
Length = 585
Score = 34.3 bits (77), Expect = 2.6
Identities = 18/48 (37%), Positives = 28/48 (58%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDL 577
E++NE+ L++LLPS R + +L ET +Y+K L R V +L
Sbjct: 401 EKLNEMFLVLKSLLPSIHR-----VNKASILAETIAYLKELQRRVQEL 443
>sptr|Q8S3D2|Q8S3D2 Putative bHLH transcription factor.
Length = 373
Score = 33.5 bits (75), Expect = 4.5
Identities = 19/50 (38%), Positives = 28/50 (56%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDLSD 571
E I+E I LQTL+P GG++ T +L E +Y+K L +V L +
Sbjct: 289 ERISEKIRVLQTLVP-----GGTKMDTASMLDEAANYLKFLRAQVKALEN 333
>sptr|Q9LIG3|Q9LIG3 Emb|CAB62312.1.
Length = 402
Score = 33.5 bits (75), Expect = 4.5
Identities = 19/50 (38%), Positives = 28/50 (56%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDLSD 571
E I+E I LQTL+P GG++ T +L E +Y+K L +V L +
Sbjct: 318 ERISEKIRVLQTLVP-----GGTKMDTASMLDEAANYLKFLRAQVKALEN 362
>sptr|Q9FHA7|Q9FHA7 Emb|CAB62312.1 (Putative bHLH transcription
factor).
Length = 241
Score = 33.1 bits (74), Expect = 5.9
Identities = 18/51 (35%), Positives = 28/51 (54%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDLSDR 568
E I+E I LQ L+P GG++ T +L E Y+K L ++V L ++
Sbjct: 142 ERISERIRILQRLVP-----GGTKMDTASMLDEAIHYVKFLKKQVQSLEEQ 187
>sptr|Q84W35|Q84W35 Hypothetical protein.
Length = 373
Score = 32.7 bits (73), Expect = 7.7
Identities = 19/50 (38%), Positives = 27/50 (54%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDLSD 571
E I+E I LQTL+P GG++ T +L E +Y K L +V L +
Sbjct: 289 ERISEKIRVLQTLVP-----GGTKMDTASMLDEAANYFKFLRAQVKALEN 333
>sptr|Q9SND4|Q9SND4 Hypothetical 25.7 kDa protein.
Length = 231
Score = 32.7 bits (73), Expect = 7.7
Identities = 18/50 (36%), Positives = 27/50 (54%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDLSD 571
E I+E I LQ L+P GG++ T +L E Y+K L ++V L +
Sbjct: 139 ERISERIRILQRLVP-----GGTKMDTASMLDEAIHYVKFLKKQVQSLEE 183
>sptrnew|CAE03479|CAE03479 OSJNBa0065O17.4 protein.
Length = 265
Score = 32.7 bits (73), Expect = 7.7
Identities = 17/48 (35%), Positives = 28/48 (58%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDL 577
E++NE+ L++LLPS R+ +L ET +Y+K L + V +L
Sbjct: 202 EKLNEMFLILKSLLPSVRK-----VDKASILAETITYLKVLEKRVKEL 244
>sptrnew|EAA06840|EAA06840 AgCP7373 (Fragment).
Length = 734
Score = 32.7 bits (73), Expect = 7.7
Identities = 22/65 (33%), Positives = 35/65 (53%), Gaps = 1/65 (1%)
Frame = -1
Query: 723 EEEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHR-EVDDLSDRLSDLMAS 547
EEE EL +R LL + G T+ L+ + +K +++ ++DDL R +L+AS
Sbjct: 664 EEEHTELQNRYDALL----QMYGEAVEKTEELQLDLADVKDMYKIQIDDLLQRQRELIAS 719
Query: 546 MDHNS 532
M H S
Sbjct: 720 MSHQS 724
>sptr|Q948Y2|Q948Y2 R-type basic helix-loop-helix protein.
Length = 451
Score = 32.7 bits (73), Expect = 7.7
Identities = 17/48 (35%), Positives = 28/48 (58%)
Frame = -1
Query: 720 EEINELISRLQTLLPSARRRGGSQASTTKLLKETCSYIKSLHREVDDL 577
E++NE+ L++LLPS R+ +L ET +Y+K L + V +L
Sbjct: 388 EKLNEMFLILKSLLPSVRK-----VDKASILAETITYLKVLEKRVKEL 430
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 458,521,033
Number of Sequences: 1272877
Number of extensions: 7340725
Number of successful extensions: 20157
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 19588
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 20145
length of database: 406,886,216
effective HSP length: 122
effective length of database: 251,595,222
effective search space used: 49061068290
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

