BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
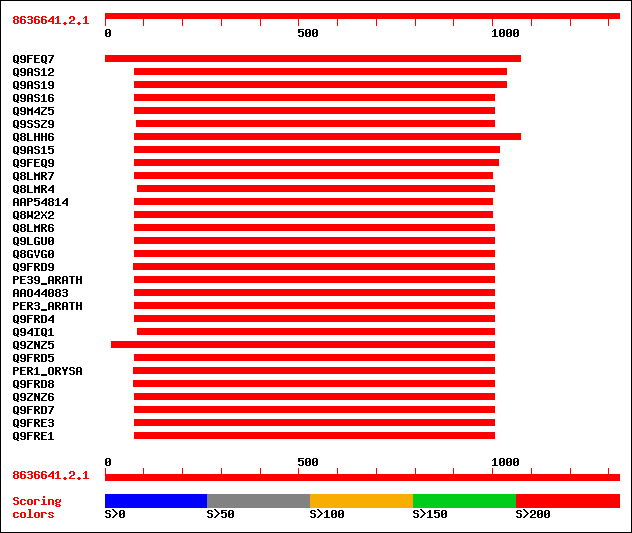
Query= 8636641.2.1
(1326 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q9FEQ7|Q9FEQ7 Peroxidase (Fragment). 676 0.0
sptr|Q9AS12|Q9AS12 Putative peroxidase. 417 e-115
sptr|Q9AS19|Q9AS19 Putative peroxidase. 327 2e-88
sptr|Q9AS16|Q9AS16 Putative peroxidase. 319 5e-86
sptr|Q9M4Z5|Q9M4Z5 Peroxidase prx12 precursor (EC 1.11.1.7). 319 5e-86
sptr|Q9SSZ9|Q9SSZ9 Peroxidase 1. 318 1e-85
sptr|Q8LHH6|Q8LHH6 Putative peroxidase. 315 9e-85
sptr|Q9AS15|Q9AS15 Putative peroxidase. 313 3e-84
sptr|Q9FEQ9|Q9FEQ9 Peroxidase. 313 4e-84
sptr|Q8LMR7|Q8LMR7 Putative peroxidase. 293 3e-78
sptr|Q8LMR4|Q8LMR4 Putative peroxidase. 291 1e-77
sptrnew|AAP54814|AAP54814 Putative peroxidase. 291 1e-77
sptr|Q8W2X2|Q8W2X2 Putative peroxidase. 291 1e-77
sptr|Q8LMR6|Q8LMR6 Putative peroxidase. 286 5e-76
sptr|Q9LGU0|Q9LGU0 Similar to Arabidopsis thaliana peroxidase AT... 277 2e-73
sptr|Q8GVG0|Q8GVG0 Putative peroxidase. 275 1e-72
sptr|Q9FRD9|Q9FRD9 Putative peroxidase. 273 4e-72
sw|Q9SUT2|PE39_ARATH Peroxidase 39 precursor (EC 1.11.1.7) (Atpe... 273 5e-72
sptrnew|AAO44083|AAO44083 At1g05260. 271 2e-71
sw|O23044|PER3_ARATH Peroxidase 3 precursor (EC 1.11.1.7) (Atper... 271 2e-71
sptr|Q9FRD4|Q9FRD4 Putative peroxidase. 269 6e-71
sptr|Q94IQ1|Q94IQ1 Peroxidase. 269 8e-71
sptr|Q9ZNZ5|Q9ZNZ5 Peroxidase precursor (EC 1.11.1.7) (Fragment). 266 5e-70
sptr|Q9FRD5|Q9FRD5 Putative peroxidase. 266 7e-70
sw|P37834|PER1_ORYSA Peroxidase 1 precursor (EC 1.11.1.7). 265 9e-70
sptr|Q9FRD8|Q9FRD8 Putative peroxidase. 265 1e-69
sptr|Q9ZNZ6|Q9ZNZ6 Peroxidase precursor (EC 1.11.1.7) (Fragment). 265 1e-69
sptr|Q9FRD7|Q9FRD7 Putative peroxidase. 265 1e-69
sptr|Q9FRE3|Q9FRE3 Putative peroxidase. 262 9e-69
sptr|Q9FRE1|Q9FRE1 Putative peroxidase. 262 9e-69
>sptr|Q9FEQ7|Q9FEQ7 Peroxidase (Fragment).
Length = 357
Score = 676 bits (1745), Expect = 0.0
Identities = 339/357 (94%), Positives = 339/357 (94%)
Frame = +2
Query: 2 TKRHCLRFXXXXXXXXXXXXXXXXDVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIR 181
TKRHCLRF DVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIR
Sbjct: 1 TKRHCLRFTTVLATTLLSATAACLDVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIR 60
Query: 182 MHFHDCFVRGCDGSVLIDTVGNLTAEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCA 361
MHFHDCFVRGCDGSVLIDTVGNLTAEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCA
Sbjct: 61 MHFHDCFVRGCDGSVLIDTVGNLTAEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCA 120
Query: 362 DVLAFAARDSVVLSGGLGYQVPGGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLT 541
DVLAFAARDSVVLSGGLGYQVPGGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLT
Sbjct: 121 DVLAFAARDSVVLSGGLGYQVPGGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLT 180
Query: 542 IEDLVVLSGAHTIGVSHCSGFAGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICP 721
IEDLVVLSGAHTIGVSHCSGFAGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICP
Sbjct: 181 IEDLVVLSGAHTIGVSHCSGFAGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICP 240
Query: 722 ANTSQFFPNTTVFMDLITPERFDNK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSE 901
ANTSQFFPNTTVFMDLITPERFDNK YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSE
Sbjct: 241 ANTSQFFPNTTVFMDLITPERFDNKYYVGLTNNLGLFKSDVALLTNATMKALVDSFVRSE 300
Query: 902 ATFRTKFARSMIKMGQIEVLTGTQGEIRRNCRVINPVSAPDDVVLARPSGFTGVAAS 1072
ATFRTKFARSMIKMGQIEVLTGTQGEIRRNCRVINPVSA DDVVLARPSGFTGVAAS
Sbjct: 301 ATFRTKFARSMIKMGQIEVLTGTQGEIRRNCRVINPVSATDDVVLARPSGFTGVAAS 357
>sptr|Q9AS12|Q9AS12 Putative peroxidase.
Length = 351
Score = 417 bits (1073), Expect = e-115
Identities = 212/321 (66%), Positives = 254/321 (79%), Gaps = 1/321 (0%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTA 256
VGFY++TCP+AE +VQQ VAAAF+NNSGVAP LIR+HFHDCFVRGCD SVLID GN T
Sbjct: 28 VGFYNKTCPSAERLVQQAVAAAFKNNSGVAPGLIRLHFHDCFVRGCDASVLID--GNDT- 84
Query: 257 EKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGR 436
EK APPNNPSLR F+V+D AKA++EA CP VVSCAD+LAFAARDSV L+G + Y+VP GR
Sbjct: 85 EKTAPPNNPSLRGFEVIDAAKAAVEAACPRVVSCADILAFAARDSVALTGNVTYKVPAGR 144
Query: 437 RDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPT 616
RDG +S +AL+NLPPP FNATEL RFA+K+LT ED+VVLSGAHTIGVSHC F
Sbjct: 145 RDGNVSIAQDALDNLPPPTFNATELVGRFANKSLTAEDMVVLSGAHTIGVSHCDSF---- 200
Query: 617 DLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERFDNK 796
RLYNF+ DP +S AYAFLL+++CP+N+SQFFPNTTV MD+ITP DNK
Sbjct: 201 -----TSRLYNFTGVGDADPAISAAYAFLLRAVCPSNSSQFFPNTTVDMDVITPAALDNK 255
Query: 797 *YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLTG-TQ 973
YVG+ NNLGLF SD ALLTNAT++A VD FV+SE +++KF ++M+KMG IEV TG TQ
Sbjct: 256 YYVGVANNLGLFTSDHALLTNATLRASVDEFVKSETRWKSKFVKAMVKMGGIEVKTGTTQ 315
Query: 974 GEIRRNCRVINPVSAPDDVVL 1036
GE+R NCRV+N SA ++ L
Sbjct: 316 GEVRLNCRVVNKRSANAELEL 336
>sptr|Q9AS19|Q9AS19 Putative peroxidase.
Length = 347
Score = 327 bits (838), Expect = 2e-88
Identities = 175/320 (54%), Positives = 223/320 (69%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTA 256
VGFY+ TCP+AE +VQQ VAAAF+NNSGVA LIR+HFHDCFVRGCD SVLI+ T
Sbjct: 26 VGFYNETCPSAEALVQQAVAAAFKNNSGVAAGLIRLHFHDCFVRGCDASVLIN---GSTT 82
Query: 257 EKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGR 436
E+ A PN SLR F+V+D AKA++EA CP VSCAD+LAFAARD + L+G + YQVP GR
Sbjct: 83 ERSAGPN-ASLRGFEVIDAAKAAVEAACPSTVSCADILAFAARDGIKLTGNVDYQVPAGR 141
Query: 437 RDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPT 616
RDG +S +AL+NLPPP A EL D+FA+K+LT+ED+VVLSGAHT+G S CS F
Sbjct: 142 RDGNVSIAQDALDNLPPPTATAKELTDKFANKSLTLEDMVVLSGAHTVGRSFCSSF---- 197
Query: 617 DLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERFDNK 796
+DR++N ++ +D LS YA LL+++CP+N + P TT +D+ TP DN
Sbjct: 198 -----LDRIWN-NTTAIVDTGLSPGYAALLRALCPSNANASTPITTA-IDVSTPATLDNN 250
Query: 797 *YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLTGTQG 976
Y L +LGLF SD L NATM ALV F +E ++ +FA +M+KMG IEVLTG G
Sbjct: 251 YYKLLPLDLGLFFSDNQLRVNATMNALVTRFAANETEWKQRFADAMVKMGNIEVLTGGAG 310
Query: 977 EIRRNCRVINPVSAPDDVVL 1036
+IR NC V+NP S+ V+
Sbjct: 311 QIRLNCNVVNPSSSSSSPVV 330
>sptr|Q9AS16|Q9AS16 Putative peroxidase.
Length = 356
Score = 319 bits (818), Expect = 5e-86
Identities = 167/310 (53%), Positives = 215/310 (69%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTA 256
VGFY+ +CP AET+V+Q V AF N+SG+A LIR+HFHDCFVRGCD SVL+ + N TA
Sbjct: 31 VGFYNTSCPNAETLVRQAVTNAFANDSGIAAGLIRLHFHDCFVRGCDASVLLTSPNN-TA 89
Query: 257 EKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGR 436
E+DA PNNPSLR F V+D AKA++E C VSCAD++AFAARDSV L+GG+ YQVP GR
Sbjct: 90 ERDAAPNNPSLRGFQVIDAAKAAVEQSCARTVSCADIVAFAARDSVNLTGGVSYQVPSGR 149
Query: 437 RDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPT 616
RDG +S +A++NLP P F A +L FA+K+LT E++VVLSGAHT+G S CS F
Sbjct: 150 RDGNVSVAQDAIDNLPQPTFTAAQLVASFANKSLTAEEMVVLSGAHTVGRSFCSSF---- 205
Query: 617 DLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERFDNK 796
+ R++N ++P +D LS YA LL+++CP+N S T +D+ TP DN
Sbjct: 206 -----LARIWNNTTPI-VDTGLSPGYAALLRALCPSNAS---ATATTAIDVSTPATLDNN 256
Query: 797 *YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLTGTQG 976
Y L NLGLF SD L NAT+ A V SF +E ++ KF +M+KMG IEVLTG+QG
Sbjct: 257 YYKLLPLNLGLFFSDNQLRVNATLGASVSSFAANETLWKEKFVAAMVKMGSIEVLTGSQG 316
Query: 977 EIRRNCRVIN 1006
E+R NC V+N
Sbjct: 317 EVRLNCSVVN 326
>sptr|Q9M4Z5|Q9M4Z5 Peroxidase prx12 precursor (EC 1.11.1.7).
Length = 331
Score = 319 bits (818), Expect = 5e-86
Identities = 167/310 (53%), Positives = 208/310 (67%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTA 256
VGFY +CP+AE IV++ V F N+ GVAP L+RMHFHDCFVRGCDGSVLID+ + TA
Sbjct: 33 VGFYCESCPSAERIVREEVMKGFMNDKGVAPGLVRMHFHDCFVRGCDGSVLIDSTSSNTA 92
Query: 257 EKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGR 436
EKD+P NNPSLR F+V+D AK LEA+C GVVSCAD+LAFAARDSV ++ G Y VP GR
Sbjct: 93 EKDSPANNPSLRGFEVIDSAKTRLEAECKGVVSCADILAFAARDSVAMTRGQRYDVPSGR 152
Query: 437 RDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPT 616
+DGR+S +E N+P FN T L FA+KNLT E++V LSGAHTIG SHC+ +
Sbjct: 153 KDGRVSLVSEGFQNIPGFTFNVTRLTQSFANKNLTQEEMVTLSGAHTIGRSHCTSVS--- 209
Query: 617 DLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERFDNK 796
+RLYNFS +G DPTL YA L+ CP ++ N V MD ++P D
Sbjct: 210 ------NRLYNFSGTNGADPTLDSKYAGQLQQQCPQGSTN--SNQVVLMDPVSPFITDVN 261
Query: 797 *YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLTGTQG 976
Y + N GLF+SD LLT++ V+ R++ + KFA +M+ MGQIEVLTGT G
Sbjct: 262 YYQDVLANKGLFRSDQTLLTDSNTANEVNQNGRNQFLWMRKFAAAMVNMGQIEVLTGTNG 321
Query: 977 EIRRNCRVIN 1006
EIR NC VIN
Sbjct: 322 EIRTNCSVIN 331
>sptr|Q9SSZ9|Q9SSZ9 Peroxidase 1.
Length = 322
Score = 318 bits (815), Expect = 1e-85
Identities = 171/309 (55%), Positives = 210/309 (67%)
Frame = +2
Query: 80 GFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTAE 259
GFY +C AETIV+Q V AF +SG+A LIR+HFHDCFVRGCDGSVLID+ G+ TAE
Sbjct: 24 GFYQLSCGFAETIVKQEVRNAFFRDSGIAAGLIRLHFHDCFVRGCDGSVLIDSTGSNTAE 83
Query: 260 KDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGRR 439
KD+PPNNPSLR F+VVD K LE CPGVVSCAD+LA+AARDSV ++ GLGY V GRR
Sbjct: 84 KDSPPNNPSLRGFEVVDAIKRRLEVSCPGVVSCADILAYAARDSVEITRGLGYDVLAGRR 143
Query: 440 DGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPTD 619
DGR+S +EAL+NLPPP FN +L FA+K L+ +++V LSGAHT+G SHC+ F
Sbjct: 144 DGRVSLASEALSNLPPPSFNVDQLTRAFANKGLSQDEMVTLSGAHTLGRSHCTSFN---- 199
Query: 620 LNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERFDNK* 799
+RLYNFS+ DPTL AYA LK CP ++ PN V MD TP D
Sbjct: 200 -----NRLYNFSTSSMQDPTLDLAYASQLKQQCPQGSAN--PNLVVPMDPPTPAVSDVSY 252
Query: 800 YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLTGTQGE 979
Y G+ N GLF SD LLT+ +A V +++ + KFA +M+ MG I V+TG GE
Sbjct: 253 YRGVLANRGLFTSDQTLLTSPQTRAQVLQNAQNQFLWWRKFAGAMVSMGNIGVITGGAGE 312
Query: 980 IRRNCRVIN 1006
IRR+CRVIN
Sbjct: 313 IRRDCRVIN 321
>sptr|Q8LHH6|Q8LHH6 Putative peroxidase.
Length = 346
Score = 315 bits (807), Expect = 9e-85
Identities = 179/337 (53%), Positives = 225/337 (66%), Gaps = 5/337 (1%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLI--DTVGNL 250
VGFY +CP AE +V+Q VAAAF ++GVA LIR+HFHDCFVRGCD SVL+ + G
Sbjct: 25 VGFYQSSCPNAEALVRQAVAAAFARDAGVAAGLIRLHFHDCFVRGCDASVLLTKNPAGGQ 84
Query: 251 TAEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPG 430
T E+DA PNNPSLR F+V+D AKA++EA CP VSCAD++AFAARDSV L+G + YQVP
Sbjct: 85 T-ERDATPNNPSLRGFEVIDAAKAAVEAACPRTVSCADIIAFAARDSVKLTGNVDYQVPA 143
Query: 431 GRRDGRISNDTEALNNLPPPFFNATELADR-FASKNLTIEDLVVLSGAHTIGVSHCSGFA 607
GRRDG +SN TEAL+NLPPP A +LAD FA+K LT+ED+VVLSGAHT+G S C+ F
Sbjct: 144 GRRDGSVSNGTEALHNLPPPNATAQQLADTFFANKFLTLEDMVVLSGAHTVGRSFCASF- 202
Query: 608 GPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERF 787
+R++N ++P +D L AYA L+++CP + T MD TP
Sbjct: 203 --------FNRVWNGNTPI-VDAGLDPAYAAQLRALCPTRDTL----ATTPMDPDTPATL 249
Query: 788 DNK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLTG 967
DN Y L GLF SD L NATM ALV F +EA ++ +FA +M+KMG IEV TG
Sbjct: 250 DNNYYKLLPQGKGLFFSDNQLRVNATMNALVTRFAANEAEWKQRFADAMVKMGHIEVQTG 309
Query: 968 TQGEIRRNCRVINPVSAPDDVVLARPSGFTG--VAAS 1072
G+IR NC V+NP ++ +V LA TG VAAS
Sbjct: 310 RCGQIRVNCNVVNPSTSSPEVELAGEDQETGGAVAAS 346
>sptr|Q9AS15|Q9AS15 Putative peroxidase.
Length = 353
Score = 313 bits (803), Expect = 3e-84
Identities = 172/314 (54%), Positives = 214/314 (68%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTA 256
VGFY+ +CPTAE +V+Q V AA NNSG+A LIR+HFHDCFVRGCD SVLI + N TA
Sbjct: 32 VGFYNTSCPTAEALVRQAVVAAVANNSGLAAGLIRLHFHDCFVRGCDASVLIFSP-NGTA 90
Query: 257 EKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGR 436
E+DA PNNPSLR F+V+D AKA++EA CP VSCAD+LAFAARDSV L+G YQVP GR
Sbjct: 91 ERDAAPNNPSLRGFEVIDAAKAAVEAACPRTVSCADILAFAARDSVNLTGNSFYQVPAGR 150
Query: 437 RDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPT 616
RDG +S DT+A LP P AT+L D F +NLT E++V+LSG+HTIG SHC+ F
Sbjct: 151 RDGNVSIDTDAF-TLPGPNLTATQLVDGFKLRNLTAEEMVILSGSHTIGRSHCASF---- 205
Query: 617 DLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERFDNK 796
L +RL N T+S AY LL+++CP T +F P TT +D+ TP DN
Sbjct: 206 -LFKNRERLAN--------GTISPAYQALLEALCPPTTGRFTPITTE-IDVSTPATLDNN 255
Query: 797 *YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLTGTQG 976
Y L NLGL SD L+ NAT+ VD+F +E ++ KF +MIKMG I+VLTG +G
Sbjct: 256 YYKLLPLNLGLHFSDDQLIRNATLLPFVDAFAANETLWKEKFVAAMIKMGNIDVLTGARG 315
Query: 977 EIRRNCRVINPVSA 1018
EIR NC +NP S+
Sbjct: 316 EIRLNCSAVNPSSS 329
>sptr|Q9FEQ9|Q9FEQ9 Peroxidase.
Length = 344
Score = 313 bits (802), Expect = 4e-84
Identities = 168/317 (52%), Positives = 214/317 (67%), Gaps = 4/317 (1%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDT-VGNLT 253
VGFYD +CP AE +V+Q VAAAF ++G+A LIR+HFHDCFVRGCDGSVL+ G
Sbjct: 14 VGFYDTSCPNAEALVRQAVAAAFAKDAGIAAGLIRLHFHDCFVRGCDGSVLLTVNPGGGQ 73
Query: 254 AEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGG 433
E+DA PNNPSLR FDV+D AK ++E CP VSCAD++AFAARDS+ L+G + YQVP G
Sbjct: 74 TERDALPNNPSLRGFDVIDAAKTAVEQSCPRTVSCADIVAFAARDSISLTGSVSYQVPAG 133
Query: 434 RRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGP 613
RRDGR+SN TE + +LPPP A L D F +K L++ED+VVLSGAHT+G S C+ F
Sbjct: 134 RRDGRVSNATETV-DLPPPTSTAQSLTDLFKAKELSVEDMVVLSGAHTVGRSFCASF--- 189
Query: 614 TDLNGPVDRLYNFSSPDG---IDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPER 784
R++N S+ +D LS +YA LL+++CP+NT+Q P TT MD TP
Sbjct: 190 ------FKRVWNTSTNPATAIVDAGLSPSYAQLLRALCPSNTTQTTPITTA-MDPGTPNV 242
Query: 785 FDNK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLT 964
DN Y + GLF SD L N M ALV SF +E ++ KFA +M+KMG+I+V T
Sbjct: 243 LDNNYYKLPASRHGLFFSDNPLRVNPQMAALVSSFASNETLWKEKFAAAMVKMGRIQVQT 302
Query: 965 GTQGEIRRNCRVINPVS 1015
GT GE+R NC V+NP S
Sbjct: 303 GTCGEVRLNCGVVNPSS 319
>sptr|Q8LMR7|Q8LMR7 Putative peroxidase.
Length = 311
Score = 293 bits (751), Expect = 3e-78
Identities = 158/308 (51%), Positives = 192/308 (62%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTA 256
VG+YD CP AE IVQ+ V+ A N G+A L+R+HFHDCFVRGCD SVL+D+ A
Sbjct: 13 VGYYDTLCPAAEIIVQEEVSKAVSGNPGMAAGLVRLHFHDCFVRGCDASVLLDSTQGNRA 72
Query: 257 EKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGR 436
EKDAPPN SLR F+V+D AK+ LE C GVVSCADVLAFAARD++ L GG YQVPGGR
Sbjct: 73 EKDAPPNT-SLRGFEVIDSAKSRLETACFGVVSCADVLAFAARDALALVGGNAYQVPGGR 131
Query: 437 RDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPT 616
RDG +S E NLPPP N +L F +K LT ++V LSGAHTIGVSHCS F+
Sbjct: 132 RDGNVSVAQETNGNLPPPSANVAQLNQMFGAKGLTQAEMVALSGAHTIGVSHCSSFS--- 188
Query: 617 DLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERFDNK 796
+RLY+ G DP++ +Y L + CP Q V MD +TP FD
Sbjct: 189 ------NRLYSSGPNAGQDPSMDPSYVAALTTQCPQQQGQPAAG-MVPMDAVTPNAFDTN 241
Query: 797 *YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLTGTQG 976
Y + N GL SD ALL + T A V + + +F+T FA +M+KMG I VLTG G
Sbjct: 242 YYAAIVANRGLLSSDQALLADQTTAAQVVGYTNNPDSFQTDFAAAMVKMGSIGVLTGNAG 301
Query: 977 EIRRNCRV 1000
IR NCRV
Sbjct: 302 TIRTNCRV 309
>sptr|Q8LMR4|Q8LMR4 Putative peroxidase.
Length = 319
Score = 291 bits (746), Expect = 1e-77
Identities = 157/310 (50%), Positives = 193/310 (62%), Gaps = 2/310 (0%)
Frame = +2
Query: 83 FYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTAEK 262
FY TCP AETIV+Q V A N G A L+RMHFHDCFVRGCDGSVL+++ + AE+
Sbjct: 19 FYAATCPQAETIVRQEVTRALYTNIGFAAGLVRMHFHDCFVRGCDGSVLLESTSDNVAER 78
Query: 263 DAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGRRD 442
D+P NNPSLR F+V+D AKA LEA CPGVVSCADVLA+AARD V L+GG Y VPGGRRD
Sbjct: 79 DSPINNPSLRGFEVIDAAKARLEAACPGVVSCADVLAYAARDGVALTGGPRYDVPGGRRD 138
Query: 443 GRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPTDL 622
G S + E +N+P P F +L FA+K LT E++V LSGAHT+G +HC+ F+
Sbjct: 139 GTASLEPEVADNIPAPTFTLDQLTQSFAAKGLTQEEMVTLSGAHTVGRAHCTSFS----- 193
Query: 623 NGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPA--NTSQFFPNTTVFMDLITPERFDNK 796
DRLYNFS+ DP++ A L+ CPA V M+ TP FD
Sbjct: 194 ----DRLYNFSATGAADPSVDPALLPQLRRACPAAGPDGAVDAGLVVPMEPRTPNGFDAL 249
Query: 797 *YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLTGTQG 976
Y + N LF SD ALL++ A V ++ KFA +M+KMGQIEVLTG G
Sbjct: 250 YYWAVLRNRALFTSDQALLSSPPTAAQVRQTAYGGYPWKLKFAAAMVKMGQIEVLTGGSG 309
Query: 977 EIRRNCRVIN 1006
EIR C +N
Sbjct: 310 EIRTKCSAVN 319
>sptrnew|AAP54814|AAP54814 Putative peroxidase.
Length = 338
Score = 291 bits (745), Expect = 1e-77
Identities = 156/313 (49%), Positives = 195/313 (62%), Gaps = 5/313 (1%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTA 256
VGFYD +CP AE IVQQ V+ A N G+A L+R+HFHDCFVRGCD SVLID+ A
Sbjct: 35 VGFYDNSCPAAEIIVQQEVSKAVSANPGLAAGLVRLHFHDCFVRGCDASVLIDSTKGNQA 94
Query: 257 EKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGR 436
EKDA PN SLR F+VVDR KA +E C GVVSCAD+LAFAARDSV L+GG YQVP GR
Sbjct: 95 EKDAGPNT-SLRGFEVVDRIKARVEQACFGVVSCADILAFAARDSVALTGGNAYQVPAGR 153
Query: 437 RDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPT 616
RDG +S ++ NLPPP + ++L FA+K L+ ++V LSGAHTIG SHCS F+
Sbjct: 154 RDGSVSRSSDTGGNLPPPTASVSQLTQMFAAKGLSQREMVALSGAHTIGASHCSSFS--- 210
Query: 617 DLNGPVDRLYNFSSP-----DGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPE 781
RLY + G DPT+ AY L CP + V MD +TP
Sbjct: 211 ------SRLYRAGTTAGGAGGGQDPTMDPAYVAQLAQQCPQSGGAAGGGALVPMDAVTPN 264
Query: 782 RFDNK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVL 961
FD + G+ NN GL SD ALL + V ++ +TF++ FA +M+KMG + VL
Sbjct: 265 AFDEGFFKGVMNNRGLLSSDQALLGDKNTAVQVVAYANDASTFQSDFAAAMVKMGAVGVL 324
Query: 962 TGTQGEIRRNCRV 1000
TG+ G++R NCRV
Sbjct: 325 TGSSGKVRANCRV 337
>sptr|Q8W2X2|Q8W2X2 Putative peroxidase.
Length = 338
Score = 291 bits (745), Expect = 1e-77
Identities = 156/313 (49%), Positives = 195/313 (62%), Gaps = 5/313 (1%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTA 256
VGFYD +CP AE IVQQ V+ A N G+A L+R+HFHDCFVRGCD SVLID+ A
Sbjct: 35 VGFYDNSCPAAEIIVQQEVSKAVSANPGLAAGLVRLHFHDCFVRGCDASVLIDSTKGNQA 94
Query: 257 EKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGR 436
EKDA PN SLR F+VVDR KA +E C GVVSCAD+LAFAARDSV L+GG YQVP GR
Sbjct: 95 EKDAGPNT-SLRGFEVVDRIKARVEQACFGVVSCADILAFAARDSVALTGGNAYQVPAGR 153
Query: 437 RDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPT 616
RDG +S ++ NLPPP + ++L FA+K L+ ++V LSGAHTIG SHCS F+
Sbjct: 154 RDGSVSRSSDTGGNLPPPTASVSQLTQMFAAKGLSQREMVALSGAHTIGASHCSSFS--- 210
Query: 617 DLNGPVDRLYNFSSP-----DGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPE 781
RLY + G DPT+ AY L CP + V MD +TP
Sbjct: 211 ------SRLYRAGTTAGGAGGGQDPTMDPAYVAQLAQQCPQSGGAAGGGALVPMDAVTPN 264
Query: 782 RFDNK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVL 961
FD + G+ NN GL SD ALL + V ++ +TF++ FA +M+KMG + VL
Sbjct: 265 AFDEGFFKGVMNNRGLLSSDQALLGDKNTAVQVVAYANDASTFQSDFAAAMVKMGAVGVL 324
Query: 962 TGTQGEIRRNCRV 1000
TG+ G++R NCRV
Sbjct: 325 TGSSGKVRANCRV 337
>sptr|Q8LMR6|Q8LMR6 Putative peroxidase.
Length = 322
Score = 286 bits (732), Expect = 5e-76
Identities = 156/310 (50%), Positives = 196/310 (63%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTA 256
VGFYD++CP AE IV+ V A N G+A L+RMHFHDCFV+GCD SVL+D+ N TA
Sbjct: 28 VGFYDQSCPQAEVIVRDEVGKAVSANVGLAAGLVRMHFHDCFVKGCDASVLLDSTANSTA 87
Query: 257 EKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGR 436
EKDA PN SLR F+VVD AK LE+ C GVVSCAD+LAFAARDSVVL+GG Y+VP GR
Sbjct: 88 EKDAIPNK-SLRGFEVVDSAKRRLESACKGVVSCADILAFAARDSVVLAGGTPYRVPAGR 146
Query: 437 RDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPT 616
RDG S ++A+ NLP P + +L FA+ L+ +D+V+LSGAHTIGV+HCS F+
Sbjct: 147 RDGNTSVASDAMANLPRPTSDVAQLTQSFATHGLSQDDMVILSGAHTIGVAHCSSFS--- 203
Query: 617 DLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERFDNK 796
RLY ++S G DP L+ A A L CP ++ TV MD + FD
Sbjct: 204 ------SRLYGYNSSTGQDPALNAAMASRLSRSCPQGSA-----NTVAMDDGSENTFDTS 252
Query: 797 *YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLTGTQG 976
Y L G+ SD L + ALV + F TKF ++M+KMG I+VLTG+ G
Sbjct: 253 YYQNLLAGRGVLASDQTLTADNATAALVAQNAYNMYLFATKFGQAMVKMGAIQVLTGSDG 312
Query: 977 EIRRNCRVIN 1006
+IR NCRV N
Sbjct: 313 QIRTNCRVAN 322
>sptr|Q9LGU0|Q9LGU0 Similar to Arabidopsis thaliana peroxidase ATP19a
(Putative peroxidase).
Length = 336
Score = 277 bits (709), Expect = 2e-73
Identities = 149/315 (47%), Positives = 201/315 (63%), Gaps = 5/315 (1%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTV--GNL 250
VG+Y+ TCP AE +V+ V AA + G P L+R+ FHDCFVRGCD SVL+D V N
Sbjct: 36 VGYYNYTCPRAEDLVRNVVRAAILRDPGNGPGLVRLFFHDCFVRGCDASVLLDAVPGSNA 95
Query: 251 TAEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPG 430
EK + NNPSLR F V+DRAK LE +C G VSCAD++AFAARD+ + GG+ + VP
Sbjct: 96 RVEKMSQANNPSLRGFAVIDRAKRVLERRCRGTVSCADIVAFAARDACGIMGGIDFAVPS 155
Query: 431 GRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAG 610
GRRDG +S +++ LNNLPPPFFNAT+L FA+KNLT +D+VVLSGAH+ G SHCS F+
Sbjct: 156 GRRDGAVSAESDVLNNLPPPFFNATQLVAGFAAKNLTADDMVVLSGAHSFGRSHCSAFS- 214
Query: 611 PTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICP---ANTSQFFPNTTVFMDLITPE 781
RLY +PD + AYA L++ CP A + + V +D +T
Sbjct: 215 --------FRLYPQVAPD-----MDAAYAAQLRARCPPPAAPPATGRRDRVVDLDPVTKL 261
Query: 782 RFDNK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVL 961
DN+ Y + LF SD L++ + ALVD + R+ + ++FA +M+KMG ++VL
Sbjct: 262 VLDNQYYKNIQRGEVLFTSDATLVSQSDTAALVDLYARNRKLWASRFAAAMVKMGNLDVL 321
Query: 962 TGTQGEIRRNCRVIN 1006
TG+QGEIR+ C +N
Sbjct: 322 TGSQGEIRKFCNRVN 336
>sptr|Q8GVG0|Q8GVG0 Putative peroxidase.
Length = 322
Score = 275 bits (703), Expect = 1e-72
Identities = 153/313 (48%), Positives = 195/313 (62%), Gaps = 3/313 (0%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLID-TVGNLT 253
VG+Y R C AE +V+ V A R N GV ++RM FHDCFV+GCD SVL+D T N
Sbjct: 26 VGYYKRKCAPAEYVVRAVVGNAVRQNPGVGAGIVRMFFHDCFVQGCDASVLLDPTAANPQ 85
Query: 254 AEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDS--VVLSGGLGYQVP 427
EK PPN PSLR F+V+D AKA++E CPGVVSCAD++AFAARD+ + GG+ Y++P
Sbjct: 86 PEKLGPPNFPSLRGFEVIDAAKAAVEKACPGVVSCADIIAFAARDASFFLSGGGISYRIP 145
Query: 428 GGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFA 607
GR DGR+S E L LPPP FN T+L F +K L +D+V LSGAHTIG SHCS FA
Sbjct: 146 AGRLDGRVSLANETLAFLPPPVFNLTQLVASFQAKGLDADDMVTLSGAHTIGRSHCSSFA 205
Query: 608 GPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERF 787
DRL S P +DP L+ A L+S CPA+ + F + TV D +TP+R
Sbjct: 206 ---------DRL---SPPSDMDPGLAAA----LRSKCPASPN-FTDDPTVAQDAVTPDRM 248
Query: 788 DNK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLTG 967
D + Y + + LF SD ALL + A+V + + +FAR+M+KMG IEV T
Sbjct: 249 DRQYYRNVLDRKVLFDSDAALLASRPTAAMVARNAAARGRWERRFARAMVKMGGIEVKTA 308
Query: 968 TQGEIRRNCRVIN 1006
GEIRR CRV+N
Sbjct: 309 ANGEIRRMCRVVN 321
>sptr|Q9FRD9|Q9FRD9 Putative peroxidase.
Length = 323
Score = 273 bits (698), Expect = 4e-72
Identities = 149/314 (47%), Positives = 196/314 (62%), Gaps = 3/314 (0%)
Frame = +2
Query: 74 DVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLID-TVGNL 250
+VG+Y ++CP ETIV++ V N+G+ LIR+ FHDCFV GCDGSVL+D T N
Sbjct: 26 EVGYYKKSCPRVETIVREEVKKFVYKNAGIGAGLIRLLFHDCFVEGCDGSVLLDPTPANP 85
Query: 251 TAEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQV-- 424
EK +PPN PSLR F+V+D AK ++E CPGVVSCAD++AFAARD+ + ++
Sbjct: 86 APEKLSPPNFPSLRGFEVIDAAKDAVEKACPGVVSCADIVAFAARDAAYFLSRMRVKINM 145
Query: 425 PGGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGF 604
P GR DGR SN ++AL+NLPPPFFN TEL D FA+K L ED+VVLSGAHT+G SHCS F
Sbjct: 146 PAGRFDGRHSNSSDALDNLPPPFFNVTELVDIFATKGLDAEDMVVLSGAHTVGRSHCSSF 205
Query: 605 AGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPER 784
DRL S DG +A LL+ CPAN + + TV D++TP
Sbjct: 206 V--------PDRLAVASDIDG-------GFAGLLRRRCPANPTTAH-DPTVNQDVVTPNA 249
Query: 785 FDNK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLT 964
FDN+ Y + + LF SD ALLT+ +V + +F ++ +KM ++V
Sbjct: 250 FDNQYYKNVIAHKVLFTSDAALLTSPATAKMVSDNANIPGWWEDRFKKAFVKMAAVDVKN 309
Query: 965 GTQGEIRRNCRVIN 1006
G QGEIR+NCRV+N
Sbjct: 310 GYQGEIRKNCRVVN 323
>sw|Q9SUT2|PE39_ARATH Peroxidase 39 precursor (EC 1.11.1.7) (Atperox
P39) (ATP19a).
Length = 326
Score = 273 bits (697), Expect = 5e-72
Identities = 157/313 (50%), Positives = 195/313 (62%), Gaps = 3/313 (0%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLID-TVGNLT 253
+GFYD+TCP AE IVQ V N +A LIRMHFHDCFVRGCDGS+LI+ T N
Sbjct: 27 MGFYDQTCPYAEKIVQDVVNQHINNAPSLAAGLIRMHFHDCFVRGCDGSILINATSSNQQ 86
Query: 254 AEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGG 433
EK APP N ++R FD +D+ K++LE++CPG+VSCAD++ A RDS+V GG + VP G
Sbjct: 87 VEKLAPP-NLTVRGFDFIDKVKSALESKCPGIVSCADIITLATRDSIVAIGGPTWNVPTG 145
Query: 434 RRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGP 613
RRDGRISN EA+NN+PPPF N T L F ++ L ++DLV+LSGAHTIGVSHCS F+
Sbjct: 146 RRDGRISNFAEAMNNIPPPFGNFTTLITLFGNQGLDVKDLVLLSGAHTIGVSHCSSFS-- 203
Query: 614 TDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTT-VFMDLITPERFD 790
+RL+NF+ DP+L YA LKS NTT V MD + FD
Sbjct: 204 -------NRLFNFTGVGDQDPSLDSEYADNLKS---RRCLSIADNTTKVEMDPGSRNTFD 253
Query: 791 NK*YVGLTNNLGLFKSDVALLTNATMKALVDSFV-RSEATFRTKFARSMIKMGQIEVLTG 967
Y + GLF+SD AL N A V F SE F +F+ SM KMG+I V TG
Sbjct: 254 LSYYRLVLKRRGLFESDAALTMNPAALAQVKRFAGGSEQEFFAEFSNSMEKMGRIGVKTG 313
Query: 968 TQGEIRRNCRVIN 1006
+ GEIRR C +N
Sbjct: 314 SDGEIRRTCAFVN 326
>sptrnew|AAO44083|AAO44083 At1g05260.
Length = 326
Score = 271 bits (693), Expect = 2e-71
Identities = 159/313 (50%), Positives = 200/313 (63%), Gaps = 3/313 (0%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLID-TVGNLT 253
+ FY +CP AE IVQ V+ N +A ALIRMHFHDCFVRGCDGSVLI+ T GN
Sbjct: 28 MNFYANSCPNAEKIVQDFVSNHVSNAPSLAAALIRMHFHDCFVRGCDGSVLINSTSGN-- 85
Query: 254 AEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGG 433
AE+DA PN ++R F +D K+ LEAQCPG+VSCAD++A A+RD+VV +GG + VP G
Sbjct: 86 AERDATPNL-TVRGFGFIDAIKSVLEAQCPGIVSCADIIALASRDAVVFTGGPNWSVPTG 144
Query: 434 RRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGP 613
RRDGRISN EAL N+PPP N T L FA++ L ++DLV+LSGAHTIGVSHCS F
Sbjct: 145 RRDGRISNAAEALANIPPPTSNITNLQTLFANQGLDLKDLVLLSGAHTIGVSHCSSF--- 201
Query: 614 TDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKS-ICPANTSQFFPNTTVFMDLITPERFD 790
+RLYNF+ G DP L YA LKS CP+ T V MD + + FD
Sbjct: 202 ------TNRLYNFTGRGGQDPALDSEYAANLKSRKCPSLNDN---KTIVEMDPGSRKTFD 252
Query: 791 NK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVR-SEATFRTKFARSMIKMGQIEVLTG 967
Y + GLF+SD AL TN T + ++ + S +F ++FA+SM KMG+I V TG
Sbjct: 253 LSYYQLVLKRRGLFQSDSALTTNPTTLSNINRILTGSVGSFFSEFAKSMEKMGRINVKTG 312
Query: 968 TQGEIRRNCRVIN 1006
+ G +RR C V N
Sbjct: 313 SAGVVRRQCSVAN 325
>sw|O23044|PER3_ARATH Peroxidase 3 precursor (EC 1.11.1.7) (Atperox
P3) (Rare cold inducible protein) (RCI3A) (ATPRC).
Length = 326
Score = 271 bits (693), Expect = 2e-71
Identities = 159/313 (50%), Positives = 200/313 (63%), Gaps = 3/313 (0%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLID-TVGNLT 253
+ FY +CP AE IVQ V+ N +A ALIRMHFHDCFVRGCDGSVLI+ T GN
Sbjct: 28 MNFYANSCPNAEKIVQDFVSNHVSNAPSLAAALIRMHFHDCFVRGCDGSVLINSTSGN-- 85
Query: 254 AEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGG 433
AE+DA PN ++R F +D K+ LEAQCPG+VSCAD++A A+RD+VV +GG + VP G
Sbjct: 86 AERDATPNL-TVRGFGFIDAIKSVLEAQCPGIVSCADIIALASRDAVVFTGGPNWSVPTG 144
Query: 434 RRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGP 613
RRDGRISN EAL N+PPP N T L FA++ L ++DLV+LSGAHTIGVSHCS F
Sbjct: 145 RRDGRISNAAEALANIPPPTSNITNLQTLFANQGLDLKDLVLLSGAHTIGVSHCSSF--- 201
Query: 614 TDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKS-ICPANTSQFFPNTTVFMDLITPERFD 790
+RLYNF+ G DP L YA LKS CP+ T V MD + + FD
Sbjct: 202 ------TNRLYNFTGRGGQDPALDSEYAANLKSRKCPSLNDN---KTIVEMDPGSRKTFD 252
Query: 791 NK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVR-SEATFRTKFARSMIKMGQIEVLTG 967
Y + GLF+SD AL TN T + ++ + S +F ++FA+SM KMG+I V TG
Sbjct: 253 LSYYQLVLKRRGLFQSDSALTTNPTTLSNINRILTGSVGSFFSEFAKSMEKMGRINVKTG 312
Query: 968 TQGEIRRNCRVIN 1006
+ G +RR C V N
Sbjct: 313 SAGVVRRQCSVAN 325
>sptr|Q9FRD4|Q9FRD4 Putative peroxidase.
Length = 339
Score = 269 bits (688), Expect = 6e-71
Identities = 154/314 (49%), Positives = 202/314 (64%), Gaps = 4/314 (1%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLID-TVGNLT 253
+G+Y CP AE IV+ VAAA + GV LIRM FHDCFV GCD SVL+D T N
Sbjct: 43 IGYYHDKCPHAEAIVKGVVAAALHRDPGVGAGLIRMLFHDCFVEGCDASVLLDPTPANPQ 102
Query: 254 AEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARD-SVVLSGG-LGYQVP 427
EK APPNNPSLR F+V+D AK ++EA CPGVVSCAD++AFAARD S LS + + +P
Sbjct: 103 PEKLAPPNNPSLRGFEVIDAAKDAVEAACPGVVSCADIVAFAARDASFFLSDSRVSFDIP 162
Query: 428 GGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFA 607
GR DGR SN + AL+ LPPP FN +L FA+K L++ED+VVLSGAHTIG+SHCS F
Sbjct: 163 SGRLDGRYSNASRALDFLPPPTFNLGQLVANFAAKGLSVEDMVVLSGAHTIGLSHCSSFV 222
Query: 608 GPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERF 787
DRL + IDP ++A +L++ CPA+ S + TV D++TP +
Sbjct: 223 S--------DRL---AVASDIDP----SFAAVLRAQCPASPSS-SNDPTVVQDVVTPNKL 266
Query: 788 DNK*YVGLTNNLGLFKSDVALLTN-ATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLT 964
DN+ Y + + LF SD +LL + AT K +VD+ + +F +M+KM +EV T
Sbjct: 267 DNQYYKNVLAHRALFTSDASLLASPATAKMVVDN-ANIPGWWEDRFKTAMVKMAAVEVKT 325
Query: 965 GTQGEIRRNCRVIN 1006
G+ GEIRR+CR +N
Sbjct: 326 GSNGEIRRHCRAVN 339
>sptr|Q94IQ1|Q94IQ1 Peroxidase.
Length = 354
Score = 269 bits (687), Expect = 8e-71
Identities = 148/310 (47%), Positives = 191/310 (61%), Gaps = 2/310 (0%)
Frame = +2
Query: 83 FYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTAEK 262
FYD CP AE+I++ + FR + G A L+R+HFHDCFV+GCDGSVL+D + +EK
Sbjct: 40 FYDSICPNAESIIRSRLQQVFRQDIGQAAGLLRLHFHDCFVQGCDGSVLLDGSASGPSEK 99
Query: 263 DAPPN-NPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGRR 439
DAPPN + F +++ + + C VVSCAD+ A AARDSV LSGG Y +P GRR
Sbjct: 100 DAPPNLTLRQQAFRIIEDLRRRVHRDCGRVVSCADITAIAARDSVFLSGGPDYDLPLGRR 159
Query: 440 DG-RISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPT 616
DG + E L NLPPP FNA+ + A+KN T D+V LSG HTIG+ HC+ F
Sbjct: 160 DGLNFATRNETLANLPPPSFNASAILTSLATKNFTPTDVVALSGGHTIGIGHCTSFT--- 216
Query: 617 DLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERFDNK 796
+RLY DP++ K +A LK+ CP + S NTTV +D+ +P +FDNK
Sbjct: 217 ------ERLY-----PNQDPSMDKTFANNLKNTCPTSNST---NTTV-LDIRSPNKFDNK 261
Query: 797 *YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLTGTQG 976
YV L N GLF SD L T+ + +V SF +E+ F +F SMIKMGQ+ VLTGTQG
Sbjct: 262 YYVDLMNRQGLFTSDQDLYTDRRTRGIVTSFAINESLFFEEFVNSMIKMGQLNVLTGTQG 321
Query: 977 EIRRNCRVIN 1006
EIR NC V N
Sbjct: 322 EIRANCSVRN 331
>sptr|Q9ZNZ5|Q9ZNZ5 Peroxidase precursor (EC 1.11.1.7) (Fragment).
Length = 351
Score = 266 bits (680), Expect = 5e-70
Identities = 154/332 (46%), Positives = 206/332 (62%), Gaps = 2/332 (0%)
Frame = +2
Query: 17 LRFXXXXXXXXXXXXXXXXDVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHD 196
LRF +GFY +CP AE IV + V N +A ALIRMHFHD
Sbjct: 32 LRFLSLCLLALIASTHAQLQLGFYANSCPKAEQIVLKFVHDHIHNAPSLAAALIRMHFHD 91
Query: 197 CFVRGCDGSVLIDTVGNLTAEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAF 376
CFVRGCD SVL+++ N AEK+APPN ++R FD +DR K+ +EA+CPGVVSCAD+L
Sbjct: 92 CFVRGCDASVLLNSTTN-QAEKNAPPNL-TVRGFDFIDRIKSLVEAECPGVVSCADILTL 149
Query: 377 AARDSVVLSGGLGYQVPGGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLV 556
AARD++V +GG ++VP GRRDG +SN TEA NN+P P N T L FA++ L ++DLV
Sbjct: 150 AARDTIVATGGPFWKVPTGRRDGVVSNLTEARNNIPAPSSNFTTLQTLFANQGLDLKDLV 209
Query: 557 VLSGAHTIGVSHCSGFAGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQ 736
+LSGAHTIG++HCS + +RL+NF+ DP+L YA LK+ + ++
Sbjct: 210 LLSGAHTIGIAHCSSLS---------NRLFNFTGKGDQDPSLDSEYAANLKAFKCTDLNK 260
Query: 737 FFPNTT-VFMDLITPERFDNK*YVGLTNNLGLFKSDVALLTNATMKA-LVDSFVRSEATF 910
NTT + MD + + FD Y + GLF+SD ALLTN+ KA ++ S F
Sbjct: 261 L--NTTKIEMDPGSRKTFDLSYYSHVIKRRGLFESDAALLTNSVTKAQIIQLLEGSVENF 318
Query: 911 RTKFARSMIKMGQIEVLTGTQGEIRRNCRVIN 1006
+FA S+ KMG+I V TGT+GEIR++C IN
Sbjct: 319 FAEFATSIEKMGRINVKTGTEGEIRKHCAFIN 350
>sptr|Q9FRD5|Q9FRD5 Putative peroxidase.
Length = 332
Score = 266 bits (679), Expect = 7e-70
Identities = 152/314 (48%), Positives = 200/314 (63%), Gaps = 4/314 (1%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLID-TVGNLT 253
VG+Y CP AE IV+ V AA + GV LIRM FHDCFV GCD SVL+D T N
Sbjct: 35 VGYYHDKCPHAEAIVRGAVGAAILRDPGVGAGLIRMLFHDCFVEGCDASVLLDPTPANPQ 94
Query: 254 AEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARD-SVVLSGG-LGYQVP 427
EK APPNNPSLR F+V+D AK ++EA CPGVVSCAD++AFAARD S LS + + +P
Sbjct: 95 PEKLAPPNNPSLRGFEVIDAAKTAVEAACPGVVSCADIVAFAARDASFFLSNSRVSFDMP 154
Query: 428 GGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFA 607
GR DGR SN + L+ LPPP FN +L FA+K L++ED+VVL+G+HT+G SHCS F
Sbjct: 155 SGRLDGRYSNASRTLDFLPPPKFNLGQLVANFAAKGLSVEDMVVLAGSHTVGRSHCSSFV 214
Query: 608 GPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERF 787
DRL + P IDP ++A L+ CPA+ S + TV D+ TP +
Sbjct: 215 --------PDRL---AVPSDIDP----SFAATLRGQCPASPSS-GNDPTVVQDVETPNKL 258
Query: 788 DNK*YVGLTNNLGLFKSDVALLTN-ATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLT 964
DN+ Y + + GLF SD +LLT+ ATMK ++D+ + +F ++M+K+ +EV T
Sbjct: 259 DNQYYKNVLAHKGLFTSDASLLTSPATMKMVLDN-ANIPGWWEDRFQKAMVKLAAVEVKT 317
Query: 965 GTQGEIRRNCRVIN 1006
G GE+RRNCR +N
Sbjct: 318 GGNGEVRRNCRAVN 331
>sw|P37834|PER1_ORYSA Peroxidase 1 precursor (EC 1.11.1.7).
Length = 326
Score = 265 bits (678), Expect = 9e-70
Identities = 157/316 (49%), Positives = 191/316 (60%), Gaps = 5/316 (1%)
Frame = +2
Query: 74 DVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLT 253
D FY +CP+ E +V++ + A +A L+RMHFHDCFVRGCDGSVL+D+ GN T
Sbjct: 25 DEKFYSNSCPSVEAVVRKEMVRALGRAPSLAGPLLRMHFHDCFVRGCDGSVLLDSAGNST 84
Query: 254 AEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGG 433
AEKDA PN +LR F V+R KA++E CPG VSCADVLA ARD+V LS G + VP G
Sbjct: 85 AEKDATPNQ-TLRGFGFVERVKAAVEKACPGTVSCADVLALMARDAVWLSKGPFWAVPLG 143
Query: 434 RRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGP 613
RRDGR+S E + LPPP N TEL FA+KNL ++DLVVLS HTIG SHC F
Sbjct: 144 RRDGRVSIANET-DQLPPPTANFTELTQMFAAKNLDLKDLVVLSAGHTIGTSHCFSF--- 199
Query: 614 TDLNGPVDRLYNFSSPDG---IDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPER 784
DRLYNF+ D IDPTL Y L+S C TS T V MD + +
Sbjct: 200 ------TDRLYNFTGLDNAHDIDPTLELQYMARLRSKC---TSLQDNTTLVEMDPGSFKT 250
Query: 785 FDNK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVRS--EATFRTKFARSMIKMGQIEV 958
FD + + GLF SD LLTN +A V + F FA SM+KMG +EV
Sbjct: 251 FDLGYFKNVAKRRGLFHSDGELLTNGFTRAYVQRHAGGGYKDEFFADFAASMVKMGGVEV 310
Query: 959 LTGTQGEIRRNCRVIN 1006
LTG+QGEIR+ C V+N
Sbjct: 311 LTGSQGEIRKKCNVVN 326
>sptr|Q9FRD8|Q9FRD8 Putative peroxidase.
Length = 332
Score = 265 bits (677), Expect = 1e-69
Identities = 150/315 (47%), Positives = 200/315 (63%), Gaps = 4/315 (1%)
Frame = +2
Query: 74 DVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLID-TVGNL 250
++ +Y CP AE +V+ V A R N G A+IRM FHDCFV GCD S+L+D T N
Sbjct: 31 ELAYYRDKCPQAEAVVKAVVGEAVRQNPGNGAAVIRMLFHDCFVEGCDASILLDPTPFNP 90
Query: 251 TAEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVV-LSGGLGY-QV 424
T EK + PNNPS+R FD++D K ++EA CPGVVSCAD++AFAARD+ LSGG Y +
Sbjct: 91 TPEKLSAPNNPSMRGFDLIDAIKHAVEAACPGVVSCADIIAFAARDATYFLSGGKVYFDM 150
Query: 425 PGGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGF 604
P GRRDG SND+ ++ LPPP N ++L FA K L++ED+VVLSGAHT+G SHCS F
Sbjct: 151 PSGRRDGTFSNDSGPIDFLPPPTSNLSDLVSSFAVKGLSVEDMVVLSGAHTVGRSHCSSF 210
Query: 605 AGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPER 784
P LN V FS DG +A+ L+S CP + + + TV +D +TP
Sbjct: 211 V-PDRLNASV-----FSDIDG-------GFAWFLRSQCPLDATPGGNDPTVMLDFVTPNT 257
Query: 785 FDNK*YVGLTNNLGLFKSDVALLTN-ATMKALVDSFVRSEATFRTKFARSMIKMGQIEVL 961
DN+ Y + ++ LF SD ALLT+ T K +VD+ V + +F +M+K+ I+V
Sbjct: 258 LDNQYYKNVLDHKVLFTSDAALLTSPETAKMVVDNAV-IPGWWEDRFKAAMVKLASIQVK 316
Query: 962 TGTQGEIRRNCRVIN 1006
TG QG+IR+NCRVIN
Sbjct: 317 TGYQGQIRKNCRVIN 331
>sptr|Q9ZNZ6|Q9ZNZ6 Peroxidase precursor (EC 1.11.1.7) (Fragment).
Length = 352
Score = 265 bits (676), Expect = 1e-69
Identities = 150/312 (48%), Positives = 205/312 (65%), Gaps = 2/312 (0%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLIDTVGNLTA 256
+GFY ++CP AE IV + V N +A ALIRMHFHDCFVRGCD SVL+++ N A
Sbjct: 53 LGFYAKSCPNAEQIVLKFVHDHIHNAPSLAAALIRMHFHDCFVRGCDASVLLNSTTN-QA 111
Query: 257 EKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQVPGGR 436
EK+APPN ++R FD +DR K+ +EA+CPGVVSCAD+L +ARD++V +GG ++VP GR
Sbjct: 112 EKNAPPNL-TVRGFDFIDRIKSLVEAECPGVVSCADILTLSARDTIVATGGPFWKVPTGR 170
Query: 437 RDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFAGPT 616
RDG ISN TEA +N+P P N T L FA++ L ++DLV+LSGAHTIG++HCS +
Sbjct: 171 RDGVISNLTEARDNIPAPSSNFTTLQTLFANQGLDLKDLVLLSGAHTIGIAHCSSLS--- 227
Query: 617 DLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTT-VFMDLITPERFDN 793
+RL+NF+ DP+L YA LK+ + ++ NTT + MD + + FD
Sbjct: 228 ------NRLFNFTGKGDQDPSLDSEYAANLKAFKCTDLNKL--NTTKIEMDPGSRKTFDL 279
Query: 794 K*YVGLTNNLGLFKSDVALLTNATMKA-LVDSFVRSEATFRTKFARSMIKMGQIEVLTGT 970
Y + GLF+SD ALLTN+ KA +++ S F +FA SM KMG+I V TGT
Sbjct: 280 SYYSHVIKRRGLFESDAALLTNSVTKAQIIELLEGSVENFFAEFATSMEKMGRINVKTGT 339
Query: 971 QGEIRRNCRVIN 1006
+GEIR++C +N
Sbjct: 340 EGEIRKHCAFLN 351
>sptr|Q9FRD7|Q9FRD7 Putative peroxidase.
Length = 340
Score = 265 bits (676), Expect = 1e-69
Identities = 155/314 (49%), Positives = 200/314 (63%), Gaps = 4/314 (1%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLID-TVGNLT 253
VG+Y CP AE IV+ V AA +N GV LIRM FHDCFV GCD SVL+D T N
Sbjct: 43 VGYYYAKCPHAEEIVKNVVGAAILHNPGVGAGLIRMLFHDCFVEGCDASVLLDPTPANPQ 102
Query: 254 AEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARD-SVVLSGG-LGYQVP 427
EK +PPN PSLR ++V+D AKA++EA CPGVVSCAD++AFAARD S LS + +Q+P
Sbjct: 103 PEKLSPPNMPSLRGYEVIDAAKAAVEAACPGVVSCADIVAFAARDASFFLSNSRVAFQMP 162
Query: 428 GGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGFA 607
GR DGR SN + AL+ LPPP FN +L FA+K L +ED+VVLSGAHT+G SHCS F
Sbjct: 163 AGRLDGRYSNASRALDFLPPPKFNLGQLVANFATKGLGMEDMVVLSGAHTVGDSHCSSFV 222
Query: 608 GPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPNTTVFMDLITPERF 787
DRL + P ++P L A +L++ CPA S + TV D++TP +
Sbjct: 223 --------PDRL---AVPSDMEPPL----AAMLRTQCPAKPSS-GNDPTVVQDVVTPNKL 266
Query: 788 DNK*YVGLTNNLGLFKSDVALLTN-ATMKALVDSFVRSEATFRTKFARSMIKMGQIEVLT 964
DN+ Y + + LF SD +LL + AT K +VD+ + +F ++M+KM IEV T
Sbjct: 267 DNQYYKNVLAHRVLFTSDASLLASPATAKMVVDN-ANIPGWWEDRFTKAMVKMASIEVKT 325
Query: 965 GTQGEIRRNCRVIN 1006
G GEIRRNCR +N
Sbjct: 326 GGNGEIRRNCRAVN 339
>sptr|Q9FRE3|Q9FRE3 Putative peroxidase.
Length = 323
Score = 262 bits (669), Expect = 9e-69
Identities = 149/319 (46%), Positives = 190/319 (59%), Gaps = 9/319 (2%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLID-TVGNLT 253
+G+Y ++CP E IV+ V ++G+ LIR+ FHDCFV GCDGSVL+D T N
Sbjct: 27 LGYYKQSCPRVEAIVRDEVKKFVYKDAGIGAGLIRLVFHDCFVEGCDGSVLLDPTPANPK 86
Query: 254 AEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQ--VP 427
EK +PPN PSLR F+V+D AK ++E CPGVVSCAD++AFAARD+ + VP
Sbjct: 87 PEKLSPPNMPSLRGFEVIDAAKDAVEKVCPGVVSCADIVAFAARDAAYFLSRFRVKINVP 146
Query: 428 GGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGF- 604
GGR DGR S D++ALNNLPPP FN +L FA+K L ED+VVLSGAHT+G SHCS F
Sbjct: 147 GGRLDGRRSLDSDALNNLPPPNFNVNQLIGAFAAKGLDAEDMVVLSGAHTVGRSHCSSFV 206
Query: 605 ----AGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPAN-TSQFFPNTTVFMDL 769
A P+D+NG +A LK CPAN TS P TV D
Sbjct: 207 SDRVAAPSDING--------------------GFANFLKQRCPANPTSSNDP--TVNQDA 244
Query: 770 ITPERFDNK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQ 949
+TP FDN+ Y + + LF SD ALLT+ +V + KFA++ +KM
Sbjct: 245 VTPNAFDNQYYKNVVAHKVLFASDAALLTSPATAKMVSDNANIPGWWEDKFAKAFVKMAS 304
Query: 950 IEVLTGTQGEIRRNCRVIN 1006
+ V TG GEIRR+CRV+N
Sbjct: 305 VGVKTGYPGEIRRHCRVVN 323
>sptr|Q9FRE1|Q9FRE1 Putative peroxidase.
Length = 323
Score = 262 bits (669), Expect = 9e-69
Identities = 149/319 (46%), Positives = 190/319 (59%), Gaps = 9/319 (2%)
Frame = +2
Query: 77 VGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGCDGSVLID-TVGNLT 253
+G+Y ++CP E IV+ V ++G+ LIR+ FHDCFV GCDGSVL+D T N
Sbjct: 27 LGYYKQSCPRVEAIVRDEVKKFVYKDAGIGAGLIRLVFHDCFVEGCDGSVLLDPTPANPK 86
Query: 254 AEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSVVLSGGLGYQ--VP 427
EK +PPN PSLR F+V+D AK ++E CPGVVSCAD++AFAARD+ + VP
Sbjct: 87 PEKLSPPNMPSLRGFEVIDAAKDAVEKVCPGVVSCADIVAFAARDAAYFLSRFRVKINVP 146
Query: 428 GGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSGAHTIGVSHCSGF- 604
GGR DGR S D++ALNNLPPP FN +L FA+K L ED+VVLSGAHT+G SHCS F
Sbjct: 147 GGRLDGRRSLDSDALNNLPPPNFNVNQLIGAFAAKGLDAEDMVVLSGAHTVGRSHCSSFV 206
Query: 605 ----AGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPAN-TSQFFPNTTVFMDL 769
A P+D+NG +A LK CPAN TS P TV D
Sbjct: 207 SDRVAAPSDING--------------------GFANFLKQRCPANPTSSNDP--TVNQDA 244
Query: 770 ITPERFDNK*YVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFARSMIKMGQ 949
+TP FDN+ Y + + LF SD ALLT+ +V + KFA++ +KM
Sbjct: 245 VTPNAFDNQYYKNVVAHKVLFASDAALLTSPATAKMVSDNANIPGWWEDKFAKAFVKMAS 304
Query: 950 IEVLTGTQGEIRRNCRVIN 1006
+ V TG GEIRR+CRV+N
Sbjct: 305 VGVKTGYPGEIRRHCRVVN 323
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 831,137,537
Number of Sequences: 1272877
Number of extensions: 17875505
Number of successful extensions: 72368
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 62140
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 70141
length of database: 406,886,216
effective HSP length: 126
effective length of database: 246,503,714
effective search space used: 77648669910
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

