BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
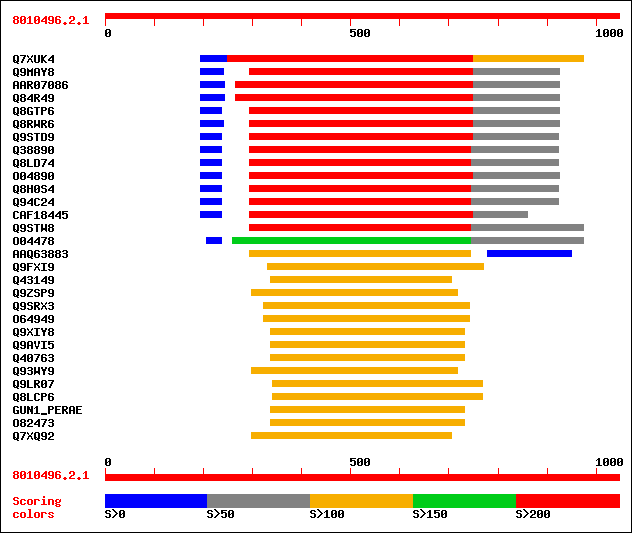
Query= 8010496.2.1
(1046 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q7XUK4|Q7XUK4 OSJNBa0067K08.14 protein. 274 2e-98
sptr|Q9MAY8|Q9MAY8 Endo-1,4-beta-glucanase Cel1. 224 1e-71
sptrnew|AAR07086|AAR07086 Putative endo-1,4-beta-glucanase. 219 6e-70
sptr|Q84R49|Q84R49 Putative endo-1,4-beta-glucanase. 219 6e-70
sptr|Q8GTP6|Q8GTP6 Endo-1,4-beta-D-glucanase (EC 3.2.1.4). 219 1e-69
sptr|Q8RWR6|Q8RWR6 Endo-1,4-beta-glucanase. 216 5e-69
sptr|Q9STD9|Q9STD9 Cellulase (EC 3.2.1.4). 218 2e-68
sptr|Q38890|Q38890 Cellulase precursor (Cellulase homolog OR16PE... 216 3e-68
sptr|Q8LD74|Q8LD74 Cellulase homolog OR16pep. 216 3e-68
sptr|O04890|O04890 Endo-1,4-beta-glucanase (EC 3.2.1.4). 213 9e-68
sptr|Q8H0S4|Q8H0S4 At5g49720/K2I5_8. 216 3e-67
sptr|Q94C24|Q94C24 AT5g49720/K2I5_8. 216 3e-67
sptrnew|CAF18445|CAF18445 Endo-1,4-beta-D-glucanase KORRIGAN (EC... 217 5e-65
sptr|Q9STW8|Q9STW8 Endo-1, 4-beta-glucanase like protein. 203 9e-64
sptr|O04478|O04478 F5I14.14. 186 2e-60
sptrnew|AAQ63883|AAQ63883 Cellulase. 142 2e-36
sptr|Q9FXI9|Q9FXI9 F6F9.1 protein (At1g19940/F6F9_1). 134 2e-30
sptr|Q43149|Q43149 Cellulase (EC 3.2.1.4) (Fragment). 122 7e-27
sptr|Q9ZSP9|Q9ZSP9 Endo-beta-1,4-D-glucanase. 122 7e-27
sptr|Q9SRX3|Q9SRX3 Endo-1,4-beta glucanase. 122 1e-26
sptr|O64949|O64949 Endo-1,4-beta glucanase. 122 1e-26
sptr|Q9XIY8|Q9XIY8 Endo-1,4-beta glucanase precursor. 120 3e-26
sptr|Q9AVI5|Q9AVI5 Endo-1,4-beta glucanase precursor. 120 4e-26
sptr|Q40763|Q40763 Cellulase precursor. 120 4e-26
sptr|Q93WY9|Q93WY9 Endo-beta-1,4-glucanase precursor (EC 3.2.1.4). 120 4e-26
sptr|Q9LR07|Q9LR07 F10A5.13 (Hypothetical protein) (Putative end... 119 6e-26
sptr|Q8LCP6|Q8LCP6 Endo-beta-1,4-glucanase, putative. 119 6e-26
sw|P05522|GUN1_PERAE Endoglucanase 1 precursor (EC 3.2.1.4) (End... 119 8e-26
sptr|O82473|O82473 Endo-1,4-beta-glucanase (EC 3.2.1.4). 119 1e-25
sptr|Q7XQ92|Q7XQ92 OSJNBa0018M05.16 protein. 117 3e-25
>sptr|Q7XUK4|Q7XUK4 OSJNBa0067K08.14 protein.
Length = 623
Score = 274 bits (701), Expect(2) = 2e-98
Identities = 127/166 (76%), Positives = 140/166 (84%)
Frame = -3
Query: 747 KGGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQI 568
KGG+A FNHG+GQPLQY VANSFLAALYADYME+VNVPGWYCGP FMT +DLR FARSQ+
Sbjct: 428 KGGLAQFNHGKGQPLQYTVANSFLAALYADYMESVNVPGWYCGPYFMTVDDLRSFARSQV 487
Query: 567 NYILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGG 388
NYILGDNPKKMSYVVG+G KYPR +HHRGASTPHNG+KYSCTGGYKWRD+K ADPN+L G
Sbjct: 488 NYILGDNPKKMSYVVGYGKKYPRRLHHRGASTPHNGIKYSCTGGYKWRDTKGADPNVLVG 547
Query: 387 AMVGGPDKNDGFKDSRNTSGRTSRRWSGNAGXXXXXXXLTSSGRGA 250
AMVGGPDKND FKD+R T + GNAG LT+SGRGA
Sbjct: 548 AMVGGPDKNDQFKDARLTYAQNEPTLVGNAGLVAALVALTNSGRGA 593
Score = 108 bits (269), Expect(2) = 2e-98
Identities = 51/77 (66%), Positives = 54/77 (70%)
Frame = -2
Query: 973 TDXRLPKXAKXXXNILDFXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSL 794
TD RLPK AK +ILDF VFSW LPGA+ RMFLNPGYPY SL GYHN TS+
Sbjct: 353 TDPRLPKNAKAFYSILDFSVFSWDNKLPGAELLLSRLRMFLNPGYPYEESLIGYHNTTSM 412
Query: 793 NMCMYFPGFAAFNFTXG 743
NMC YFP F AFNFT G
Sbjct: 413 NMCTYFPRFGAFNFTKG 429
Score = 42.0 bits (97), Expect = 0.016
Identities = 22/47 (46%), Positives = 23/47 (48%)
Frame = -2
Query: 334 FGQNEPTLVXXXXXXXXXXXXXXXXXXXSVTAVDKNTMFSAVPHNVP 194
+ QNEPTLV VTAVDKNTMFSAVP P
Sbjct: 566 YAQNEPTLVGNAGLVAALVALTNSGRGAGVTAVDKNTMFSAVPPMFP 612
>sptr|Q9MAY8|Q9MAY8 Endo-1,4-beta-glucanase Cel1.
Length = 621
Score = 224 bits (572), Expect(3) = 1e-71
Identities = 101/151 (66%), Positives = 120/151 (79%)
Frame = -3
Query: 747 KGGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQI 568
KGG+ NHG QPLQYVV +FLA+LYADY++ + PGWYCGPNF TT+ LR+FA+SQ+
Sbjct: 428 KGGLIQLNHGGPQPLQYVVNAAFLASLYADYLDTADTPGWYCGPNFYTTDVLRKFAKSQL 487
Query: 567 NYILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGG 388
+YILG NP+KMSYVVGFG KYP+ VHHRGAS PHNGVKY C GG+KWR+SKKA+PN+L G
Sbjct: 488 DYILGKNPQKMSYVVGFGKKYPKRVHHRGASIPHNGVKYGCKGGFKWRESKKANPNILVG 547
Query: 387 AMVGGPDKNDGFKDSRNTSGRTSRRWSGNAG 295
AMV GPDK+DGFKD R T + NAG
Sbjct: 548 AMVAGPDKHDGFKDIRTNYNYTEPTLAANAG 578
Score = 64.3 bits (155), Expect(3) = 1e-71
Identities = 29/61 (47%), Positives = 37/61 (60%)
Frame = -2
Query: 925 DFXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSLNMCMYFPGFAAFNFTX 746
++ VF+W LPG+Q R+FL+PGYPY L +HN T MC Y P F +FNFT
Sbjct: 369 NYGVFTWDDKLPGSQVLLSRLRLFLSPGYPYEEILRTFHNQTDNVMCSYLPVFNSFNFTK 428
Query: 745 G 743
G
Sbjct: 429 G 429
Score = 25.4 bits (54), Expect(3) = 1e-71
Identities = 10/16 (62%), Positives = 13/16 (81%)
Frame = -2
Query: 241 AVDKNTMFSAVPHNVP 194
++DKNT+FSAVP P
Sbjct: 595 SIDKNTIFSAVPPMFP 610
>sptrnew|AAR07086|AAR07086 Putative endo-1,4-beta-glucanase.
Length = 620
Score = 219 bits (558), Expect(3) = 6e-70
Identities = 99/161 (61%), Positives = 121/161 (75%)
Frame = -3
Query: 747 KGGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQI 568
KGGM NHGR QPLQYVV +FLA+LY+DY++A + PGWYCGP F TT LR+FARSQ+
Sbjct: 428 KGGMIQLNHGRPQPLQYVVNAAFLASLYSDYLDAADTPGWYCGPTFYTTEVLRKFARSQL 487
Query: 567 NYILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGG 388
+Y+LG NP KMSYVVGFG+KYP+ HHRGAS PHNGVKY C GG+KWR++KK +PN+L G
Sbjct: 488 DYVLGKNPLKMSYVVGFGNKYPKRAHHRGASIPHNGVKYGCKGGFKWRETKKPNPNILIG 547
Query: 387 AMVGGPDKNDGFKDSRNTSGRTSRRWSGNAGXXXXXXXLTS 265
A+V GPD++DGFKD R T + NAG LT+
Sbjct: 548 ALVAGPDRHDGFKDVRTNYNYTEPTLAANAGLVAALISLTN 588
Score = 64.3 bits (155), Expect(3) = 6e-70
Identities = 29/61 (47%), Positives = 37/61 (60%)
Frame = -2
Query: 925 DFXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSLNMCMYFPGFAAFNFTX 746
++ VF+W LPGAQ R+FL+PGYPY L +HN T MC Y P + +FNFT
Sbjct: 369 NYGVFTWDDKLPGAQVLLSRLRLFLSPGYPYEEILRTFHNQTDNVMCSYLPMYNSFNFTK 428
Query: 745 G 743
G
Sbjct: 429 G 429
Score = 25.4 bits (54), Expect(3) = 6e-70
Identities = 10/17 (58%), Positives = 13/17 (76%)
Frame = -2
Query: 244 TAVDKNTMFSAVPHNVP 194
+ +DKNT+FSAVP P
Sbjct: 593 SGIDKNTIFSAVPPMFP 609
>sptr|Q84R49|Q84R49 Putative endo-1,4-beta-glucanase.
Length = 620
Score = 219 bits (558), Expect(3) = 6e-70
Identities = 99/161 (61%), Positives = 121/161 (75%)
Frame = -3
Query: 747 KGGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQI 568
KGGM NHGR QPLQYVV +FLA+LY+DY++A + PGWYCGP F TT LR+FARSQ+
Sbjct: 428 KGGMIQLNHGRPQPLQYVVNAAFLASLYSDYLDAADTPGWYCGPTFYTTEVLRKFARSQL 487
Query: 567 NYILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGG 388
+Y+LG NP KMSYVVGFG+KYP+ HHRGAS PHNGVKY C GG+KWR++KK +PN+L G
Sbjct: 488 DYVLGKNPLKMSYVVGFGNKYPKRAHHRGASIPHNGVKYGCKGGFKWRETKKPNPNILIG 547
Query: 387 AMVGGPDKNDGFKDSRNTSGRTSRRWSGNAGXXXXXXXLTS 265
A+V GPD++DGFKD R T + NAG LT+
Sbjct: 548 ALVAGPDRHDGFKDVRTNYNYTEPTLAANAGLVAALISLTN 588
Score = 64.3 bits (155), Expect(3) = 6e-70
Identities = 29/61 (47%), Positives = 37/61 (60%)
Frame = -2
Query: 925 DFXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSLNMCMYFPGFAAFNFTX 746
++ VF+W LPGAQ R+FL+PGYPY L +HN T MC Y P + +FNFT
Sbjct: 369 NYGVFTWDDKLPGAQVLLSRLRLFLSPGYPYEEILRTFHNQTDNVMCSYLPMYNSFNFTK 428
Query: 745 G 743
G
Sbjct: 429 G 429
Score = 25.4 bits (54), Expect(3) = 6e-70
Identities = 10/17 (58%), Positives = 13/17 (76%)
Frame = -2
Query: 244 TAVDKNTMFSAVPHNVP 194
+ +DKNT+FSAVP P
Sbjct: 593 SGIDKNTIFSAVPPMFP 609
>sptr|Q8GTP6|Q8GTP6 Endo-1,4-beta-D-glucanase (EC 3.2.1.4).
Length = 621
Score = 219 bits (559), Expect(3) = 1e-69
Identities = 97/151 (64%), Positives = 121/151 (80%)
Frame = -3
Query: 747 KGGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQI 568
KGG+ NHGR QPLQYVV +FLA LY++Y++A + PGWYCGPNF +T+ LREFA++QI
Sbjct: 430 KGGLIQLNHGRPQPLQYVVNAAFLATLYSEYLDAADTPGWYCGPNFYSTDVLREFAKTQI 489
Query: 567 NYILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGG 388
+YILG NP+KMSYVVGFG+ YP+HVHHRGAS P N +KYSC GG+KWRD+ KA+PN + G
Sbjct: 490 DYILGKNPRKMSYVVGFGNHYPKHVHHRGASIPKNKIKYSCKGGWKWRDTPKANPNTIDG 549
Query: 387 AMVGGPDKNDGFKDSRNTSGRTSRRWSGNAG 295
AMV GPDK+DGF+D R+ T +GNAG
Sbjct: 550 AMVAGPDKHDGFRDVRSNYNYTEPTLAGNAG 580
Score = 63.5 bits (153), Expect(3) = 1e-69
Identities = 29/61 (47%), Positives = 34/61 (55%)
Frame = -2
Query: 925 DFXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSLNMCMYFPGFAAFNFTX 746
D+ V SW L GAQ +FL+PGYPY L +HN TS+ MC Y P F FN T
Sbjct: 371 DYGVLSWDNKLAGAQVLLSRLXLFLSPGYPYEEILRTFHNQTSIIMCSYLPVFTTFNRTK 430
Query: 745 G 743
G
Sbjct: 431 G 431
Score = 25.0 bits (53), Expect(3) = 1e-69
Identities = 10/15 (66%), Positives = 12/15 (80%)
Frame = -2
Query: 238 VDKNTMFSAVPHNVP 194
+DKNT+FSAVP P
Sbjct: 596 IDKNTIFSAVPPMFP 610
>sptr|Q8RWR6|Q8RWR6 Endo-1,4-beta-glucanase.
Length = 620
Score = 216 bits (550), Expect(3) = 5e-69
Identities = 98/151 (64%), Positives = 117/151 (77%)
Frame = -3
Query: 747 KGGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQI 568
KGG+ NHG QPLQYVV +FLA+LYADY++ + P WYC PNF TT+ LR+FA+SQ+
Sbjct: 428 KGGLIQLNHGGPQPLQYVVNAAFLASLYADYLDTADTPRWYCRPNFYTTDVLRKFAKSQL 487
Query: 567 NYILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGG 388
+YILG NP+KMSYVVGFG KYP+ VHHRGAS PHNGVKY C GG+KWR+ KKA+PN+L G
Sbjct: 488 DYILGKNPQKMSYVVGFGKKYPKRVHHRGASIPHNGVKYGCKGGFKWREFKKANPNILVG 547
Query: 387 AMVGGPDKNDGFKDSRNTSGRTSRRWSGNAG 295
AMV GPDK+DGFKD R T + NAG
Sbjct: 548 AMVAGPDKHDGFKDIRTNYNYTEPTLAANAG 578
Score = 64.3 bits (155), Expect(3) = 5e-69
Identities = 29/61 (47%), Positives = 37/61 (60%)
Frame = -2
Query: 925 DFXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSLNMCMYFPGFAAFNFTX 746
++ VF+W LPG+Q R+FL+PGYPY L +HN T MC Y P F +FNFT
Sbjct: 369 NYGVFTWDDKLPGSQVLLSRLRLFLSPGYPYEEILRTFHNQTDNVMCSYLPVFNSFNFTK 428
Query: 745 G 743
G
Sbjct: 429 G 429
Score = 25.4 bits (54), Expect(3) = 5e-69
Identities = 10/16 (62%), Positives = 13/16 (81%)
Frame = -2
Query: 241 AVDKNTMFSAVPHNVP 194
++DKNT+FSAVP P
Sbjct: 595 SIDKNTIFSAVPPMFP 610
>sptr|Q9STD9|Q9STD9 Cellulase (EC 3.2.1.4).
Length = 621
Score = 218 bits (554), Expect(3) = 2e-68
Identities = 97/151 (64%), Positives = 117/151 (77%)
Frame = -3
Query: 747 KGGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQI 568
+GG+ NHG QPLQY +FLA LY+DY++A + PGWYCGPNF +TN LREFAR+QI
Sbjct: 428 RGGLIELNHGDPQPLQYAANAAFLATLYSDYLDAADTPGWYCGPNFYSTNVLREFARTQI 487
Query: 567 NYILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGG 388
+YILG NP+KMSY+VGFG+KYP+HVHHRGAS P N VKY+C GG+KWRDSKK +PN + G
Sbjct: 488 DYILGKNPRKMSYLVGFGTKYPKHVHHRGASIPKNKVKYNCKGGWKWRDSKKPNPNTIEG 547
Query: 387 AMVGGPDKNDGFKDSRNTSGRTSRRWSGNAG 295
AMV GPDK DGF+D R T +GNAG
Sbjct: 548 AMVAGPDKRDGFRDVRTNYNYTEPTLAGNAG 578
Score = 61.2 bits (147), Expect(3) = 2e-68
Identities = 29/60 (48%), Positives = 35/60 (58%)
Frame = -2
Query: 922 FXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSLNMCMYFPGFAAFNFTXG 743
+ VFSW L GAQ R+FL+PGYPY + +HN TS+ MC Y P F FN T G
Sbjct: 370 YGVFSWDNKLAGAQLLLSRLRLFLSPGYPYEEIVRTFHNQTSIVMCSYLPYFNKFNRTRG 429
Score = 24.6 bits (52), Expect(3) = 2e-68
Identities = 10/15 (66%), Positives = 12/15 (80%)
Frame = -2
Query: 238 VDKNTMFSAVPHNVP 194
+DKNT+FSAVP P
Sbjct: 596 IDKNTIFSAVPPLFP 610
>sptr|Q38890|Q38890 Cellulase precursor (Cellulase homolog OR16PEP
precursor).
Length = 621
Score = 216 bits (551), Expect(3) = 3e-68
Identities = 98/150 (65%), Positives = 117/150 (78%)
Frame = -3
Query: 744 GGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQIN 565
GG+ NHG QPLQY V +FLA LY+DY++A + PGWYCGPNF +T+ LR+FARSQI+
Sbjct: 429 GGLIELNHGAPQPLQYSVNAAFLATLYSDYLDAADTPGWYCGPNFYSTSVLRDFARSQID 488
Query: 564 YILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGGA 385
YILG NP+KMSYVVGFG+KYPRHVHHRGAS P N VKY+C GG+KWRDSKK +PN + GA
Sbjct: 489 YILGKNPRKMSYVVGFGTKYPRHVHHRGASIPKNKVKYNCKGGWKWRDSKKPNPNTIEGA 548
Query: 384 MVGGPDKNDGFKDSRNTSGRTSRRWSGNAG 295
MV GPDK DG++D R T +GNAG
Sbjct: 549 MVAGPDKRDGYRDVRMNYNYTEPTLAGNAG 578
Score = 62.0 bits (149), Expect(3) = 3e-68
Identities = 30/60 (50%), Positives = 35/60 (58%)
Frame = -2
Query: 922 FXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSLNMCMYFPGFAAFNFTXG 743
+ VFSW L GAQ R+FL+PGYPY L +HN TS+ MC Y P F FN T G
Sbjct: 370 YGVFSWDNKLAGAQLLLSRLRLFLSPGYPYEEILRTFHNQTSIVMCSYLPIFNKFNRTNG 429
Score = 24.6 bits (52), Expect(3) = 3e-68
Identities = 10/15 (66%), Positives = 12/15 (80%)
Frame = -2
Query: 238 VDKNTMFSAVPHNVP 194
+DKNT+FSAVP P
Sbjct: 596 IDKNTIFSAVPPLFP 610
>sptr|Q8LD74|Q8LD74 Cellulase homolog OR16pep.
Length = 621
Score = 216 bits (551), Expect(3) = 3e-68
Identities = 98/150 (65%), Positives = 117/150 (78%)
Frame = -3
Query: 744 GGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQIN 565
GG+ NHG QPLQY V +FLA LY+DY++A + PGWYCGPNF +T+ LR+FARSQI+
Sbjct: 429 GGLIELNHGAPQPLQYSVNAAFLATLYSDYLDAADTPGWYCGPNFYSTSVLRDFARSQID 488
Query: 564 YILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGGA 385
YILG NP+KMSYVVGFG+KYPRHVHHRGAS P N VKY+C GG+KWRDSKK +PN + GA
Sbjct: 489 YILGKNPRKMSYVVGFGTKYPRHVHHRGASIPKNKVKYNCKGGWKWRDSKKPNPNTIEGA 548
Query: 384 MVGGPDKNDGFKDSRNTSGRTSRRWSGNAG 295
MV GPDK DG++D R T +GNAG
Sbjct: 549 MVAGPDKRDGYRDVRMNYNYTEPTLAGNAG 578
Score = 62.0 bits (149), Expect(3) = 3e-68
Identities = 30/60 (50%), Positives = 35/60 (58%)
Frame = -2
Query: 922 FXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSLNMCMYFPGFAAFNFTXG 743
+ VFSW L GAQ R+FL+PGYPY L +HN TS+ MC Y P F FN T G
Sbjct: 370 YGVFSWDNKLAGAQLLLSRLRLFLSPGYPYEEILRTFHNQTSIVMCSYLPIFNKFNRTNG 429
Score = 24.6 bits (52), Expect(3) = 3e-68
Identities = 10/15 (66%), Positives = 12/15 (80%)
Frame = -2
Query: 238 VDKNTMFSAVPHNVP 194
+DKNT+FSAVP P
Sbjct: 596 IDKNTIFSAVPPLFP 610
>sptr|O04890|O04890 Endo-1,4-beta-glucanase (EC 3.2.1.4).
Length = 617
Score = 213 bits (541), Expect(3) = 9e-68
Identities = 96/151 (63%), Positives = 119/151 (78%)
Frame = -3
Query: 747 KGGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQI 568
KGG+ NHGR QPLQYVV +FLA L++DY+ A + PGWYCGPNF +T+ LR+FA +QI
Sbjct: 426 KGGLIQLNHGRPQPLQYVVNAAFLATLFSDYLAAADTPGWYCGPNFYSTDVLRKFAETQI 485
Query: 567 NYILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGG 388
+YILG NP+KMSYVVGFG+ YP+HVHHRGAS P N VKY+C GG+K+RDS KA+PN + G
Sbjct: 486 DYILGKNPRKMSYVVGFGNHYPKHVHHRGASIPKNKVKYNCKGGWKYRDSSKANPNTIVG 545
Query: 387 AMVGGPDKNDGFKDSRNTSGRTSRRWSGNAG 295
AMV GPDK+DGF+D R+ T +GNAG
Sbjct: 546 AMVAGPDKHDGFRDVRSNYNYTEPTLAGNAG 576
Score = 63.5 bits (153), Expect(3) = 9e-68
Identities = 30/61 (49%), Positives = 36/61 (59%)
Frame = -2
Query: 925 DFXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSLNMCMYFPGFAAFNFTX 746
D+ V SW L GAQ R+FL+PGYPY L +HN TS+ MC Y P F +FN T
Sbjct: 367 DYGVLSWDNKLTGAQVLLSRMRLFLSPGYPYEEILRTFHNQTSIIMCSYLPIFTSFNRTK 426
Query: 745 G 743
G
Sbjct: 427 G 427
Score = 25.4 bits (54), Expect(3) = 9e-68
Identities = 10/15 (66%), Positives = 12/15 (80%)
Frame = -2
Query: 238 VDKNTMFSAVPHNVP 194
+DKNT+FSAVP P
Sbjct: 592 IDKNTLFSAVPPMFP 606
>sptr|Q8H0S4|Q8H0S4 At5g49720/K2I5_8.
Length = 621
Score = 216 bits (551), Expect(3) = 3e-67
Identities = 98/150 (65%), Positives = 117/150 (78%)
Frame = -3
Query: 744 GGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQIN 565
GG+ NHG QPLQY V +FLA LY+DY++A + PGWYCGPNF +T+ LR+FARSQI+
Sbjct: 429 GGLIELNHGAPQPLQYSVNAAFLATLYSDYLDAADTPGWYCGPNFYSTSVLRDFARSQID 488
Query: 564 YILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGGA 385
YILG NP+KMSYVVGFG+KYPRHVHHRGAS P N VKY+C GG+KWRDSKK +PN + GA
Sbjct: 489 YILGKNPRKMSYVVGFGTKYPRHVHHRGASIPKNKVKYNCKGGWKWRDSKKPNPNTIEGA 548
Query: 384 MVGGPDKNDGFKDSRNTSGRTSRRWSGNAG 295
MV GPDK DG++D R T +GNAG
Sbjct: 549 MVAGPDKRDGYRDVRMNYNYTEPTLAGNAG 578
Score = 58.5 bits (140), Expect(3) = 3e-67
Identities = 29/60 (48%), Positives = 34/60 (56%)
Frame = -2
Query: 922 FXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSLNMCMYFPGFAAFNFTXG 743
+ VFSW L GAQ R+FL+PG PY L +HN TS+ MC Y P F FN T G
Sbjct: 370 YGVFSWDNKLAGAQLLLSRLRLFLSPGCPYEEILRTFHNQTSIVMCSYLPIFNKFNRTNG 429
Score = 24.6 bits (52), Expect(3) = 3e-67
Identities = 10/15 (66%), Positives = 12/15 (80%)
Frame = -2
Query: 238 VDKNTMFSAVPHNVP 194
+DKNT+FSAVP P
Sbjct: 596 IDKNTIFSAVPPLFP 610
>sptr|Q94C24|Q94C24 AT5g49720/K2I5_8.
Length = 621
Score = 216 bits (551), Expect(3) = 3e-67
Identities = 98/150 (65%), Positives = 117/150 (78%)
Frame = -3
Query: 744 GGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQIN 565
GG+ NHG QPLQY V +FLA LY+DY++A + PGWYCGPNF +T+ LR+FARSQI+
Sbjct: 429 GGLIELNHGAPQPLQYSVNAAFLATLYSDYLDAADTPGWYCGPNFYSTSVLRDFARSQID 488
Query: 564 YILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGGA 385
YILG NP+KMSYVVGFG+KYPRHVHHRGAS P N VKY+C GG+KWRDSKK +PN + GA
Sbjct: 489 YILGKNPRKMSYVVGFGTKYPRHVHHRGASIPKNKVKYNCKGGWKWRDSKKPNPNTIEGA 548
Query: 384 MVGGPDKNDGFKDSRNTSGRTSRRWSGNAG 295
MV GPDK DG++D R T +GNAG
Sbjct: 549 MVAGPDKRDGYRDVRMNYNYTEPTLAGNAG 578
Score = 58.5 bits (140), Expect(3) = 3e-67
Identities = 29/60 (48%), Positives = 34/60 (56%)
Frame = -2
Query: 922 FXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSLNMCMYFPGFAAFNFTXG 743
+ VFSW L GAQ R+FL+PG PY L +HN TS+ MC Y P F FN T G
Sbjct: 370 YGVFSWDNKLAGAQLLLSRLRLFLSPGCPYEEILRTFHNQTSIVMCSYLPIFNKFNRTNG 429
Score = 24.6 bits (52), Expect(3) = 3e-67
Identities = 10/15 (66%), Positives = 12/15 (80%)
Frame = -2
Query: 238 VDKNTMFSAVPHNVP 194
+DKNT+FSAVP P
Sbjct: 596 IDKNTIFSAVPPLFP 610
>sptrnew|CAF18445|CAF18445 Endo-1,4-beta-D-glucanase KORRIGAN (EC
3.2.1.4) (Fragment).
Length = 229
Score = 217 bits (552), Expect(3) = 5e-65
Identities = 96/151 (63%), Positives = 119/151 (78%)
Frame = -3
Query: 747 KGGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQI 568
KGG+ NHGR QPLQYVV +FLA LY+DY++A + PGWYCGPNF +T+ LREFAR+QI
Sbjct: 45 KGGLIQLNHGRPQPLQYVVNAAFLATLYSDYLDAADTPGWYCGPNFYSTDKLREFARTQI 104
Query: 567 NYILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGG 388
+YILG NP+KMSY+VGFG+ YP+HVHHRGAS P VKY+C GG+KWRD+ K +PN+L G
Sbjct: 105 DYILGKNPRKMSYIVGFGNHYPKHVHHRGASIPKTKVKYNCKGGWKWRDTSKPNPNILVG 164
Query: 387 AMVGGPDKNDGFKDSRNTSGRTSRRWSGNAG 295
AMV GPD++DGF D R+ T +GNAG
Sbjct: 165 AMVAGPDRHDGFHDVRSNYNYTEPTLAGNAG 195
Score = 50.1 bits (118), Expect(3) = 5e-65
Identities = 21/39 (53%), Positives = 26/39 (66%)
Frame = -2
Query: 859 MFLNPGYPYXXSLAGYHNATSLNMCMYFPGFAAFNFTXG 743
+FL+PGYPY L +HN TS+ MC Y P F +FN T G
Sbjct: 8 LFLSPGYPYEEILRTFHNQTSIIMCSYLPVFTSFNRTKG 46
Score = 25.4 bits (54), Expect(3) = 5e-65
Identities = 10/15 (66%), Positives = 12/15 (80%)
Frame = -2
Query: 238 VDKNTMFSAVPHNVP 194
+DKNT+FSAVP P
Sbjct: 211 IDKNTLFSAVPPMFP 225
>sptr|Q9STW8|Q9STW8 Endo-1, 4-beta-glucanase like protein.
Length = 620
Score = 203 bits (516), Expect(2) = 9e-64
Identities = 93/150 (62%), Positives = 113/150 (75%)
Frame = -3
Query: 744 GGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQIN 565
GG+ NHG QPLQYV +FLAAL++DY+EA + PGWYCGPNF TT LR F+RSQI+
Sbjct: 430 GGLIQLNHGAPQPLQYVANAAFLAALFSDYLEAADTPGWYCGPNFYTTEFLRNFSRSQID 489
Query: 564 YILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGGA 385
YILG NP+KMSYVVG+G +YP+ VHHRGAS P N +K +CTGG+KW+ SKK +PN + GA
Sbjct: 490 YILGKNPRKMSYVVGYGQRYPKQVHHRGASIPKN-MKETCTGGFKWKKSKKNNPNAINGA 548
Query: 384 MVGGPDKNDGFKDSRNTSGRTSRRWSGNAG 295
MV GPDK+DGF D R T +GNAG
Sbjct: 549 MVAGPDKHDGFHDIRTNYNYTEPTLAGNAG 578
Score = 63.9 bits (154), Expect(2) = 9e-64
Identities = 32/77 (41%), Positives = 40/77 (51%)
Frame = -2
Query: 973 TDXRLPKXAKXXXNILDFXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSL 794
T + + A N + VFSW LPGAQ R+FL+PGYPY L+ +HN T
Sbjct: 354 TSHHMAEKAGAFGNSPYYGVFSWDNKLPGAQLLLTRMRLFLSPGYPYEDMLSEFHNQTGR 413
Query: 793 NMCMYFPGFAAFNFTXG 743
MC Y P + FN T G
Sbjct: 414 VMCSYLPYYKKFNRTNG 430
>sptr|O04478|O04478 F5I14.14.
Length = 623
Score = 186 bits (473), Expect(3) = 2e-60
Identities = 89/162 (54%), Positives = 114/162 (70%)
Frame = -3
Query: 744 GGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQIN 565
GG+ N G+ +PL+YV SFLA+L+ADY+ + VPGWYCGP F+ + L++FA+SQI+
Sbjct: 433 GGLMQLNLGKPRPLEYVAHASFLASLFADYLNSTGVPGWYCGPTFVENHVLKDFAQSQID 492
Query: 564 YILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGGA 385
YILGDNP KMSYVVGFG K+PR VHHRGA+ P++ + SC G K+RD+K +PN + GA
Sbjct: 493 YILGDNPLKMSYVVGFGKKFPRRVHHRGATIPNDKKRRSCREGLKYRDTKNPNPNNITGA 552
Query: 384 MVGGPDKNDGFKDSRNTSGRTSRRWSGNAGXXXXXXXLTSSG 259
MVGGP+K D F D RN + SGNAG LTSSG
Sbjct: 553 MVGGPNKFDEFHDLRNNYNASEPTLSGNAGLVAALVSLTSSG 594
Score = 66.6 bits (161), Expect(3) = 2e-60
Identities = 33/77 (42%), Positives = 40/77 (51%)
Frame = -2
Query: 973 TDXRLPKXAKXXXNILDFXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSL 794
T +P+ AK N + V SW LPGA R+FLNPG+PY L YHNAT +
Sbjct: 357 TTPSVPQTAKAFANRPELMVPSWNNKLPGAMLLMTRYRLFLNPGFPYENMLNRYHNATGI 416
Query: 793 NMCMYFPGFAAFNFTXG 743
MC Y + FN T G
Sbjct: 417 TMCAYLKQYNVFNRTSG 433
Score = 23.5 bits (49), Expect(3) = 2e-60
Identities = 8/11 (72%), Positives = 11/11 (100%)
Frame = -2
Query: 238 VDKNTMFSAVP 206
+DKNTMF++VP
Sbjct: 598 IDKNTMFNSVP 608
>sptrnew|AAQ63883|AAQ63883 Cellulase.
Length = 601
Score = 142 bits (358), Expect(2) = 2e-36
Identities = 70/150 (46%), Positives = 93/150 (62%)
Frame = -3
Query: 744 GGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQIN 565
GG+ + G LQY V SFL+ LY+DY++ + + G C + + + LR+F+ SQ+N
Sbjct: 426 GGLVIPKPDNGPLLQYAVTASFLSKLYSDYIDHLKISGASCETDTFSVSMLRDFSSSQVN 485
Query: 564 YILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKADPNLLGGA 385
YILG NP KMSY+VG+G K+P VHHR AS P + Y+C G W +SK +P +L GA
Sbjct: 486 YILGQNPMKMSYLVGYGDKFPVQVHHRSASIPWDKRLYNCDDGKTWLNSKNPNPQVLLGA 545
Query: 384 MVGGPDKNDGFKDSRNTSGRTSRRWSGNAG 295
MVGGPD ND F D R+ T S NAG
Sbjct: 546 MVGGPDTNDHFTDQRSNKRFTEPTISSNAG 575
Score = 33.1 bits (74), Expect(2) = 2e-36
Identities = 20/57 (35%), Positives = 23/57 (40%)
Frame = -2
Query: 949 AKXXXNILDFXVFSWXXXLPGAQXXXXXXRMFLNPGYPYXXSLAGYHNATSLNMCMY 779
AK LD VF W L R F +PG+PY L N+T MC Y
Sbjct: 359 AKKDETNLDKGVFYWNNKLSAVAVLLTGIRYFRDPGFPYEDVLKFSSNSTHSLMCSY 415
>sptr|Q9FXI9|Q9FXI9 F6F9.1 protein (At1g19940/F6F9_1).
Length = 515
Score = 134 bits (338), Expect = 2e-30
Identities = 69/147 (46%), Positives = 92/147 (62%)
Frame = -3
Query: 771 ASPPSTSPKGGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDL 592
+SP +TS + L LQ+ V+++FLA LY+DYM V C +DL
Sbjct: 340 SSPTATSSRTDGGLIWVSEWNALQHPVSSAFLATLYSDYMLTSGVKELSCSDQSFKPSDL 399
Query: 591 REFARSQINYILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKK 412
R+FARSQ +Y+LG NP+KMSY+VG+G KYP VHHRGAS P + C G+KW +S +
Sbjct: 400 RKFARSQADYMLGKNPEKMSYLVGYGEKYPEFVHHRGASIPADATT-GCKDGFKWLNSDE 458
Query: 411 ADPNLLGGAMVGGPDKNDGFKDSRNTS 331
+PN+ GA+VGGP ND F D+RN S
Sbjct: 459 PNPNVAYGALVGGPFLNDTFIDARNNS 485
>sptr|Q43149|Q43149 Cellulase (EC 3.2.1.4) (Fragment).
Length = 508
Score = 122 bits (307), Expect = 7e-27
Identities = 60/127 (47%), Positives = 82/127 (64%), Gaps = 4/127 (3%)
Frame = -3
Query: 705 LQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQINYILGDNPKKMSYV 526
LQYV + S + +Y+ + A N+ G CG T+ ++ FA+SQ++YILG+NP KMSY+
Sbjct: 337 LQYVTSASMVFLIYSKILNAANIKGVQCGSVNFPTSQIKAFAKSQVDYILGNNPMKMSYM 396
Query: 525 VGFGSKYPRHVHHRGASTPH---NGVKYSCTGGYK-WRDSKKADPNLLGGAMVGGPDKND 358
VGFGSKYP +HHRGAS P + + C GY W S K++PN GA+VGGP+ ND
Sbjct: 397 VGFGSKYPLQLHHRGASIPSMQIHPARVGCNEGYSAWYSSSKSNPNTHVGAIVGGPNSND 456
Query: 357 GFKDSRN 337
F D R+
Sbjct: 457 QFNDVRS 463
>sptr|Q9ZSP9|Q9ZSP9 Endo-beta-1,4-D-glucanase.
Length = 625
Score = 122 bits (307), Expect = 7e-27
Identities = 66/146 (45%), Positives = 88/146 (60%), Gaps = 6/146 (4%)
Frame = -3
Query: 717 RGQPLQYVVANSFLAALYADYMEAVNVPGWY--CGPNFMTTNDLREFARSQINYILGDNP 544
R +Q+V + +FLA Y+DY+ + G Y C F++ N+L FA+SQ++YILGDNP
Sbjct: 336 RWNNMQFVTSAAFLATTYSDYLASA---GKYLKCSSGFVSPNELLSFAKSQVDYILGDNP 392
Query: 543 KKMSYVVGFGSKYPRHVHHRGASTPH---NGVKYSCTGGY-KWRDSKKADPNLLGGAMVG 376
+ SY+VG+G+ YPR VHHR +S N SC GGY W K +DPNLL GA+VG
Sbjct: 393 RATSYMVGYGNNYPRQVHHRASSIVSFKVNPSFVSCRGGYATWYSRKASDPNLLTGALVG 452
Query: 375 GPDKNDGFKDSRNTSGRTSRRWSGNA 298
GPD D F D R+ +T NA
Sbjct: 453 GPDAYDNFADQRDNYEQTEPATYNNA 478
>sptr|Q9SRX3|Q9SRX3 Endo-1,4-beta glucanase.
Length = 501
Score = 122 bits (305), Expect = 1e-26
Identities = 64/143 (44%), Positives = 86/143 (60%), Gaps = 3/143 (2%)
Frame = -3
Query: 741 GMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQINY 562
G LF G +QYV + SFL YA Y+ + YCG + +T LR A+ Q++Y
Sbjct: 341 GGLLFKMGESN-MQYVTSTSFLLLTYAKYLTSARTVA-YCGGSVVTPARLRSIAKKQVDY 398
Query: 561 ILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGV---KYSCTGGYKWRDSKKADPNLLG 391
+LG NP KMSY+VG+G KYPR +HHRG+S P V + C G+ S+ +PN L
Sbjct: 399 LLGGNPLKMSYMVGYGLKYPRRIHHRGSSLPSVAVHPTRIQCHDGFSLFTSQSPNPNDLV 458
Query: 390 GAMVGGPDKNDGFKDSRNTSGRT 322
GA+VGGPD+ND F D R+ GR+
Sbjct: 459 GAVVGGPDQNDQFPDERSDYGRS 481
>sptr|O64949|O64949 Endo-1,4-beta glucanase.
Length = 501
Score = 122 bits (305), Expect = 1e-26
Identities = 64/143 (44%), Positives = 86/143 (60%), Gaps = 3/143 (2%)
Frame = -3
Query: 741 GMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQINY 562
G LF G +QYV + SFL YA Y+ + YCG + +T LR A+ Q++Y
Sbjct: 341 GGLLFKMGESN-MQYVTSTSFLLLTYAKYLTSARTVA-YCGGSVVTPARLRSIAKKQVDY 398
Query: 561 ILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGV---KYSCTGGYKWRDSKKADPNLLG 391
+LG NP KMSY+VG+G KYPR +HHRG+S P V + C G+ S+ +PN L
Sbjct: 399 LLGGNPLKMSYMVGYGLKYPRRIHHRGSSLPSVAVHPTRIQCHDGFSLFTSQSPNPNDLV 458
Query: 390 GAMVGGPDKNDGFKDSRNTSGRT 322
GA+VGGPD+ND F D R+ GR+
Sbjct: 459 GAVVGGPDQNDQFPDERSDYGRS 481
>sptr|Q9XIY8|Q9XIY8 Endo-1,4-beta glucanase precursor.
Length = 494
Score = 120 bits (301), Expect = 3e-26
Identities = 63/135 (46%), Positives = 84/135 (62%), Gaps = 3/135 (2%)
Frame = -3
Query: 732 LFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQINYILG 553
LF LQYV + +FL YA Y+ + N CG + +T L A+ Q++YILG
Sbjct: 335 LFYKATESNLQYVTSTTFLLLTYAKYLGS-NGGVAKCGGSTVTAESLIAQAKKQVDYILG 393
Query: 552 DNPKKMSYVVGFGSKYPRHVHHRGASTPH---NGVKYSCTGGYKWRDSKKADPNLLGGAM 382
DNP KMSY+VGFG+KYP+HVHHRG+S P + + SC G+++ S +PN+L GA+
Sbjct: 394 DNPAKMSYMVGFGNKYPQHVHHRGSSVPSIHAHPNRISCNDGFQYLYSSSPNPNVLVGAI 453
Query: 381 VGGPDKNDGFKDSRN 337
VGGPD D F D RN
Sbjct: 454 VGGPDNRDHFADDRN 468
>sptr|Q9AVI5|Q9AVI5 Endo-1,4-beta glucanase precursor.
Length = 494
Score = 120 bits (300), Expect = 4e-26
Identities = 61/135 (45%), Positives = 85/135 (62%), Gaps = 3/135 (2%)
Frame = -3
Query: 732 LFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQINYILG 553
LF LQYV + +FL YA Y+ + N CG + +TT L A+ Q++YILG
Sbjct: 335 LFYKASESNLQYVTSTTFLLLTYAKYLGS-NGGVARCGGSTVTTESLIAQAKKQVDYILG 393
Query: 552 DNPKKMSYVVGFGSKYPRHVHHRGASTPH---NGVKYSCTGGYKWRDSKKADPNLLGGAM 382
DNP +MSY+VGFG++YP+HVHHRG+S P + + SC G+++ S +PN+L GA+
Sbjct: 394 DNPARMSYMVGFGNRYPQHVHHRGSSVPSIHAHPNRISCNDGFQFLYSSSPNPNVLVGAI 453
Query: 381 VGGPDKNDGFKDSRN 337
+GGPD D F D RN
Sbjct: 454 IGGPDNRDNFADDRN 468
>sptr|Q40763|Q40763 Cellulase precursor.
Length = 494
Score = 120 bits (300), Expect = 4e-26
Identities = 61/135 (45%), Positives = 84/135 (62%), Gaps = 3/135 (2%)
Frame = -3
Query: 732 LFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQINYILG 553
LF LQYV + +FL YA Y+ + N CG + +T L A+ Q++YILG
Sbjct: 335 LFYKASESNLQYVTSTTFLLLTYAKYLGS-NGGVARCGGSTVTAESLIAQAKKQVDYILG 393
Query: 552 DNPKKMSYVVGFGSKYPRHVHHRGASTP---HNGVKYSCTGGYKWRDSKKADPNLLGGAM 382
DNP +MSY+VGFG++YP+HVHHRG+S P + SC G+++ S +PN+L GA+
Sbjct: 394 DNPARMSYMVGFGNRYPQHVHHRGSSVPSIHDTRIGISCNDGFQFLYSSSPNPNVLVGAI 453
Query: 381 VGGPDKNDGFKDSRN 337
+GGPD D F DSRN
Sbjct: 454 IGGPDNRDNFADSRN 468
>sptr|Q93WY9|Q93WY9 Endo-beta-1,4-glucanase precursor (EC 3.2.1.4).
Length = 624
Score = 120 bits (300), Expect = 4e-26
Identities = 63/144 (43%), Positives = 86/144 (59%), Gaps = 4/144 (2%)
Frame = -3
Query: 717 RGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQINYILGDNPKK 538
R +Q+V + SFLA +Y+DY+ + C + ++L FA+SQ++YILGDNP+
Sbjct: 344 RWNNMQFVTSASFLATVYSDYLASAR-KSLKCSSGTVLPSELLSFAKSQVDYILGDNPRA 402
Query: 537 MSYVVGFGSKYPRHVHHRGA---STPHNGVKYSCTGGY-KWRDSKKADPNLLGGAMVGGP 370
SY+VG+G+ YPR VHHRG+ S + SC GGY W K +DPNLL GA+VGGP
Sbjct: 403 TSYMVGYGNNYPRQVHHRGSSIVSVKKDPSFVSCRGGYATWFSRKASDPNLLAGAIVGGP 462
Query: 369 DKNDGFKDSRNTSGRTSRRWSGNA 298
D D F D R+ +T NA
Sbjct: 463 DAYDNFADQRDNYEQTEPATYNNA 486
>sptr|Q9LR07|Q9LR07 F10A5.13 (Hypothetical protein) (Putative
endo-beta-1,4-glucanase).
Length = 525
Score = 119 bits (299), Expect = 6e-26
Identities = 61/143 (42%), Positives = 86/143 (60%)
Frame = -3
Query: 768 SPPSTSPKGGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLR 589
SP ST+ + L +Q V+++FLA+L++DYM + C +LR
Sbjct: 350 SPTSTASRTNGGLIWVSEWNSMQQSVSSAFLASLFSDYMLTSRIHKISCDGKIFKATELR 409
Query: 588 EFARSQINYILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKA 409
+FA+SQ +Y+LG NP S+VVG+G KYP+ VHHRGAS P + C G+KW +S K
Sbjct: 410 DFAKSQADYMLGKNPLGTSFVVGYGDKYPQFVHHRGASIPADATT-GCLDGFKWFNSTKP 468
Query: 408 DPNLLGGAMVGGPDKNDGFKDSR 340
+PN+ GA+VGGP N+ F DSR
Sbjct: 469 NPNIAYGALVGGPFFNETFTDSR 491
>sptr|Q8LCP6|Q8LCP6 Endo-beta-1,4-glucanase, putative.
Length = 525
Score = 119 bits (299), Expect = 6e-26
Identities = 61/143 (42%), Positives = 86/143 (60%)
Frame = -3
Query: 768 SPPSTSPKGGMALFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLR 589
SP ST+ + L +Q V+++FLA+L++DYM + C +LR
Sbjct: 350 SPTSTASRTNGGLIWVSEWNSMQQSVSSAFLASLFSDYMLTSRIHKISCDGKIFKATELR 409
Query: 588 EFARSQINYILGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGVKYSCTGGYKWRDSKKA 409
+FA+SQ +Y+LG NP S+VVG+G KYP+ VHHRGAS P + C G+KW +S K
Sbjct: 410 DFAKSQADYMLGKNPLGTSFVVGYGDKYPQFVHHRGASIPADATT-GCLDGFKWFNSTKP 468
Query: 408 DPNLLGGAMVGGPDKNDGFKDSR 340
+PN+ GA+VGGP N+ F DSR
Sbjct: 469 NPNIAYGALVGGPFFNETFTDSR 491
>sw|P05522|GUN1_PERAE Endoglucanase 1 precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (Abscission cellulase 1).
Length = 494
Score = 119 bits (298), Expect = 8e-26
Identities = 61/137 (44%), Positives = 82/137 (59%), Gaps = 5/137 (3%)
Frame = -3
Query: 732 LFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWY--CGPNFMTTNDLREFARSQINYI 559
L G LQYV + +FL YA+Y+ N G + CG +T +L A+ Q++YI
Sbjct: 332 LLYKGSASNLQYVTSTAFLLLTYANYL---NSSGGHASCGTTTVTAKNLISLAKKQVDYI 388
Query: 558 LGDNPKKMSYVVGFGSKYPRHVHHRGASTPHNGV---KYSCTGGYKWRDSKKADPNLLGG 388
LG NP KMSY+VGFG +YP+HVHHRG+S P V C G+++ S +PN+L G
Sbjct: 389 LGQNPAKMSYMVGFGERYPQHVHHRGSSLPSVQVHPNSIPCNAGFQYLYSSPPNPNILVG 448
Query: 387 AMVGGPDKNDGFKDSRN 337
A++GGPD D F D RN
Sbjct: 449 AILGGPDNRDSFSDDRN 465
>sptr|O82473|O82473 Endo-1,4-beta-glucanase (EC 3.2.1.4).
Length = 497
Score = 119 bits (297), Expect = 1e-25
Identities = 64/136 (47%), Positives = 82/136 (60%), Gaps = 4/136 (2%)
Frame = -3
Query: 732 LFNHGRGQPLQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQINYILG 553
L G G LQYV ++SFL YA Y+ + N CG + L E A+ Q++YILG
Sbjct: 338 LLYKGSGSNLQYVTSSSFLLLTYAKYLRS-NGGDVSCGSSRFPAERLVELAKKQVDYILG 396
Query: 552 DNPKKMSYVVGFGSKYPRHVHHRGASTP----HNGVKYSCTGGYKWRDSKKADPNLLGGA 385
DNP K+SY+VGFG KYP VHHRG+S P H G C G++ +S +PN+L GA
Sbjct: 397 DNPAKISYMVGFGDKYPLRVHHRGSSLPSVHAHPG-HIGCNDGFQSLNSGSPNPNVLIGA 455
Query: 384 MVGGPDKNDGFKDSRN 337
+VGGPD D F+D RN
Sbjct: 456 IVGGPDSRDNFEDDRN 471
>sptr|Q7XQ92|Q7XQ92 OSJNBa0018M05.16 protein.
Length = 625
Score = 117 bits (293), Expect = 3e-25
Identities = 63/140 (45%), Positives = 85/140 (60%), Gaps = 4/140 (2%)
Frame = -3
Query: 705 LQYVVANSFLAALYADYMEAVNVPGWYCGPNFMTTNDLREFARSQINYILGDNPKKMSYV 526
+QYV +++FL +YADY+ A + C + ++ FARSQ++Y+LG NPK MSY+
Sbjct: 357 MQYVSSSAFLLTVYADYL-AESRGTLRCPDGEVKPAEILRFARSQVDYVLGKNPKGMSYM 415
Query: 525 VGFGSKYPRHVHHRGASTPH---NGVKYSCTGGY-KWRDSKKADPNLLGGAMVGGPDKND 358
VG+GS YP HVHHRGAS P C + K+ +SK ADPN+L GA+VGGPD ND
Sbjct: 416 VGYGSYYPTHVHHRGASIPSIYAMNATVGCMESFDKYYNSKNADPNVLHGALVGGPDAND 475
Query: 357 GFKDSRNTSGRTSRRWSGNA 298
+ D R +GNA
Sbjct: 476 AYDDDRCNYQHAEPTLAGNA 495
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 613,824,776
Number of Sequences: 1395590
Number of extensions: 11640400
Number of successful extensions: 31689
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 30105
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 31432
length of database: 442,889,342
effective HSP length: 124
effective length of database: 269,836,182
effective search space used: 60443304768
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

