BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
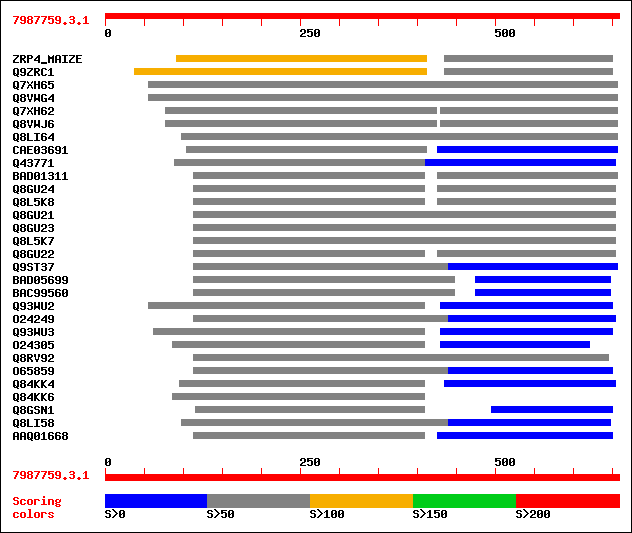
Query= 7987759.3.1
(658 letters)
Database: swall
1,381,838 sequences; 439,479,560 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sw|P47917|ZRP4_MAIZE O-methyltransferase ZRP4 (EC 2.1.1.-) (OMT). 123 1e-43
sptr|Q9ZRC1|Q9ZRC1 O-methyltransferase. 107 5e-33
sptr|Q7XH65|Q7XH65 Putative o-methyltransferase ZRP4. 96 7e-29
sptr|Q8VWG4|Q8VWG4 Putative to o-methyltransferase ZRP4 (Putativ... 96 7e-29
sptr|Q7XH62|Q7XH62 Putative o-methyltransferase ZRP4. 95 1e-28
sptr|Q8VWJ6|Q8VWJ6 Putative to o-methyltransferase ZRP4 (Putativ... 95 1e-28
sptr|Q8LI64|Q8LI64 Putative o-methyltransferase. 94 2e-27
sptrnew|CAE03691|CAE03691 OSJNBb0026E15.9 protein. 96 6e-27
sptr|Q43771|Q43771 Flavonoid 7-O-methyltransferase (EC 2.1.1.6). 92 3e-26
sptrnew|BAD01311|BAD01311 Putative catechol O-methyltransferase. 87 4e-26
sptr|Q8GU24|Q8GU24 Orcinol O-methyltransferase. 82 1e-25
sptr|Q8L5K8|Q8L5K8 Orcinol O-methyltransferase. 82 1e-25
sptr|Q8GU21|Q8GU21 Orcinol O-methyltransferase (Fragment). 81 1e-25
sptr|Q8GU23|Q8GU23 Orcinol O-methyltransferase. 79 5e-25
sptr|Q8L5K7|Q8L5K7 Orcinol O-methyltransferase. 79 5e-25
sptr|Q8GU22|Q8GU22 Orcinol O-methyltransferase (Fragment). 80 9e-25
sptr|Q9ST37|Q9ST37 O-methyltransferase. 83 1e-23
sptrnew|BAD05699|BAD05699 Putative flavonoid 7-O-methyltransferase. 87 3e-23
sptrnew|BAC99560|BAC99560 Putative flavonoid 7-O-methyltransferase. 87 3e-23
sptr|Q93WU2|Q93WU2 Eugenol O-methyltransferase. 82 2e-22
sptr|O24249|O24249 Methyltransferase. 76 8e-22
sptr|Q93WU3|Q93WU3 Chavicol O-methyltransferase. 75 3e-20
sptr|O24305|O24305 6a-hydroxymaackiain methyltransferase. 92 4e-20
sptr|Q8RV92|Q8RV92 Putative o-methyltransferase. 99 6e-20
sptr|O65859|O65859 O-methyltransferase. 82 1e-19
sptr|Q84KK4|Q84KK4 S-adenosyl-L-methionine: 2,7,4'-trihydroxyiso... 89 2e-19
sptr|Q84KK6|Q84KK6 S-adenosyl-L-methionine: 2,7,4'-trihydroxyiso... 91 1e-17
sptr|Q8GSN1|Q8GSN1 Flavonoid O-methyltransferase. 71 9e-17
sptr|Q8LI58|Q8LI58 OJ1634_H04.28 protein. 87 2e-16
sptrnew|AAQ01668|AAQ01668 (R,S)-reticuline 7-O-methyltransferase. 69 1e-14
>sw|P47917|ZRP4_MAIZE O-methyltransferase ZRP4 (EC 2.1.1.-) (OMT).
Length = 364
Score = 123 bits (309), Expect(2) = 1e-43
Identities = 63/107 (58%), Positives = 80/107 (74%)
Frame = +2
Query: 92 SSEQQVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPS 271
+S Q LDA+L+LW+ TF ++KSMALK+A+ L I DAIH HGG+A++ QI++++ LHPS
Sbjct: 7 NSTDQSLLDAQLELWHTTFAFMKSMALKSAIHLRIADAIHLHGGAASLSQILSKVHLHPS 66
Query: 272 KTPCLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVG 412
+ LRRLMRVLT T VFGTQ P G D E VYTLTP SRLL+G
Sbjct: 67 RVSSLRRLMRVLTTTNVFGTQ-PLGGGSDDDSEPVYTLTPVSRLLIG 112
Score = 75.1 bits (183), Expect(2) = 1e-43
Identities = 34/72 (47%), Positives = 46/72 (63%)
Frame = +3
Query: 435 PLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAGMVA 614
PL ++L P VS F +L WFQHE+PDP F THGR IWEL DA F L + G+ +
Sbjct: 122 PLAAMVLDPTIVSPFSELGAWFQHELPDPCIFKHTHGRGIWELTKDDATFDALVNDGLAS 181
Query: 615 DSGFIMNVVVRE 650
DS I++V +++
Sbjct: 182 DSQLIVDVAIKQ 193
>sptr|Q9ZRC1|Q9ZRC1 O-methyltransferase.
Length = 373
Score = 107 bits (268), Expect(2) = 5e-33
Identities = 64/126 (50%), Positives = 80/126 (63%), Gaps = 1/126 (0%)
Frame = +2
Query: 38 MALSKEQKLTISEQQHTASSEQQVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQH 217
MAL+ + KL ++ L +L H +G+VKSMALK A++LGIP AIH H
Sbjct: 1 MALTGDYKLISTDDM----------LQGHAELCIHAYGFVKSMALKCAIELGIPGAIHGH 50
Query: 218 GGSATIPQIVTRITLHPSKTPCLRRLMRVLTVTGVFGTQ-EPHDDGGGCDDELVYTLTPA 394
GG AT+ ++ T I L PS+ P LRRLMRVLTV+GVF Q + DD GC +VY LT A
Sbjct: 51 GGGATLGELATIIALPPSRLPRLRRLMRVLTVSGVFSVQNQQPDDPAGC-AAVVYGLTAA 109
Query: 395 SRLLVG 412
SRLLVG
Sbjct: 110 SRLLVG 115
Score = 55.5 bits (132), Expect(2) = 5e-33
Identities = 26/72 (36%), Positives = 38/72 (52%)
Frame = +3
Query: 435 PLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAGMVA 614
P +M+ P + F + WF + S F + HG D+WE+ A DA SR GM
Sbjct: 125 PPRSLMVDPNLTAPFSGMSAWFMDDEQPRSFFEMHHGEDMWEMAARDAALSRTIGDGMTD 184
Query: 615 DSGFIMNVVVRE 650
DS F++ V++RE
Sbjct: 185 DSRFVVEVLLRE 196
>sptr|Q7XH65|Q7XH65 Putative o-methyltransferase ZRP4.
Length = 366
Score = 95.9 bits (237), Expect(2) = 7e-29
Identities = 52/123 (42%), Positives = 80/123 (65%)
Frame = +2
Query: 56 QKLTISEQQHTASSEQQVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATI 235
Q++ ++Q T++ + + AE++L++H F ++KS AL+AA DL I DAIH++GG+AT+
Sbjct: 3 QRVQEEDEQMTSTDD---LIQAEIELYHHCFAFIKSTALRAATDLCISDAIHRNGGAATL 59
Query: 236 PQIVTRITLHPSKTPCLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGP 415
+ I LHP+K LRRLMRV TV+GVF ++ + E +YTLT SRLL+
Sbjct: 60 SDLALNIGLHPTKLSHLRRLMRVPTVSGVFAVEDH-------NGEAMYTLTRVSRLLLNG 112
Query: 416 TGQ 424
G+
Sbjct: 113 DGE 115
Score = 53.5 bits (127), Expect(2) = 7e-29
Identities = 28/83 (33%), Positives = 48/83 (57%), Gaps = 2/83 (2%)
Frame = +3
Query: 414 QRGKNVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDP-SPFNVTHGRDIWELDAHDAGFSR 590
+R ++ L++V++ P+ V+S + WF E +PF V HG WE+ A+DA
Sbjct: 115 ERTHALSHLVRVLVNPLTVASHFSIHEWFTIEQAAAMTPFEVAHGCTRWEIIANDAKDGS 174
Query: 591 LFDAGMVADSGFIMNVVVRE-CG 656
+F+ MV DS M+++++E CG
Sbjct: 175 VFNTAMVEDSRVAMDIILKESCG 197
>sptr|Q8VWG4|Q8VWG4 Putative to o-methyltransferase ZRP4 (Putative
o-methyltransferase ZRP4).
Length = 366
Score = 95.9 bits (237), Expect(2) = 7e-29
Identities = 52/123 (42%), Positives = 80/123 (65%)
Frame = +2
Query: 56 QKLTISEQQHTASSEQQVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATI 235
Q++ ++Q T++ + + AE++L++H F ++KS AL+AA DL I DAIH++GG+AT+
Sbjct: 3 QRVQEEDEQMTSTDD---LIQAEIELYHHCFAFIKSTALRAATDLCISDAIHRNGGAATL 59
Query: 236 PQIVTRITLHPSKTPCLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGP 415
+ I LHP+K LRRLMRV TV+GVF ++ + E +YTLT SRLL+
Sbjct: 60 SDLALNIGLHPTKLSHLRRLMRVPTVSGVFAVEDH-------NGEAMYTLTRVSRLLLNG 112
Query: 416 TGQ 424
G+
Sbjct: 113 DGE 115
Score = 53.5 bits (127), Expect(2) = 7e-29
Identities = 28/83 (33%), Positives = 48/83 (57%), Gaps = 2/83 (2%)
Frame = +3
Query: 414 QRGKNVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDP-SPFNVTHGRDIWELDAHDAGFSR 590
+R ++ L++V++ P+ V+S + WF E +PF V HG WE+ A+DA
Sbjct: 115 ERTHALSHLVRVLVNPLTVASHFSIHEWFTIEQAAAMTPFEVAHGCTRWEIIANDAKDGS 174
Query: 591 LFDAGMVADSGFIMNVVVRE-CG 656
+F+ MV DS M+++++E CG
Sbjct: 175 VFNTAMVEDSRVAMDIILKESCG 197
>sptr|Q7XH62|Q7XH62 Putative o-methyltransferase ZRP4.
Length = 366
Score = 94.7 bits (234), Expect(2) = 1e-28
Identities = 51/120 (42%), Positives = 76/120 (63%), Gaps = 4/120 (3%)
Frame = +2
Query: 77 QQHTASSEQQVALD----AELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQI 244
Q+ EQ ++ D A+++L++H F ++KS AL AA+DL I D IH++GG+AT+ +
Sbjct: 3 QRVQEEDEQMMSTDDLIQAQIKLYHHCFAFIKSTALWAAIDLRIADVIHRNGGAATLSDL 62
Query: 245 VTRITLHPSKTPCLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGPTGQ 424
+ LHP+K LRRLMRVLTVTG+F ++ + E +YTLT SRLL+ G+
Sbjct: 63 ALNVGLHPTKLSHLRRLMRVLTVTGIFAVEDR-------NGEAMYTLTRVSRLLLNSDGE 115
Score = 53.9 bits (128), Expect(2) = 1e-28
Identities = 28/78 (35%), Positives = 45/78 (57%), Gaps = 2/78 (2%)
Frame = +3
Query: 429 VNPLMKVMLRPIFVSSFLDLRGWFQHEMPDP-SPFNVTHGRDIWELDAHDAGFSRLFDAG 605
++ + +V+ P+ V S + WF E +PF V HG WE+ A+DA +F+AG
Sbjct: 120 LSQMARVLANPLAVISHFSIHEWFTTEKATTMTPFEVAHGCTRWEMIANDAKDGSVFNAG 179
Query: 606 MVADSGFIMNVVVRE-CG 656
MV DS M+++++E CG
Sbjct: 180 MVEDSRVAMDIILKESCG 197
>sptr|Q8VWJ6|Q8VWJ6 Putative to o-methyltransferase ZRP4 (Putative
o-methyltransferase ZRP4).
Length = 366
Score = 94.7 bits (234), Expect(2) = 1e-28
Identities = 51/120 (42%), Positives = 76/120 (63%), Gaps = 4/120 (3%)
Frame = +2
Query: 77 QQHTASSEQQVALD----AELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQI 244
Q+ EQ ++ D A+++L++H F ++KS AL AA+DL I D IH++GG+AT+ +
Sbjct: 3 QRVQEEDEQMMSTDDLIQAQIKLYHHCFAFIKSTALWAAIDLRIADVIHRNGGAATLSDL 62
Query: 245 VTRITLHPSKTPCLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGPTGQ 424
+ LHP+K LRRLMRVLTVTG+F ++ + E +YTLT SRLL+ G+
Sbjct: 63 ALNVGLHPTKLSHLRRLMRVLTVTGIFAVEDR-------NGEAMYTLTRVSRLLLNSDGE 115
Score = 53.9 bits (128), Expect(2) = 1e-28
Identities = 28/78 (35%), Positives = 45/78 (57%), Gaps = 2/78 (2%)
Frame = +3
Query: 429 VNPLMKVMLRPIFVSSFLDLRGWFQHEMPDP-SPFNVTHGRDIWELDAHDAGFSRLFDAG 605
++ + +V+ P+ V S + WF E +PF V HG WE+ A+DA +F+AG
Sbjct: 120 LSQMARVLANPLAVISHFSIHEWFTTEKATTMTPFEVAHGCTRWEMIANDAKDGSVFNAG 179
Query: 606 MVADSGFIMNVVVRE-CG 656
MV DS M+++++E CG
Sbjct: 180 MVEDSRVAMDIILKESCG 197
>sptr|Q8LI64|Q8LI64 Putative o-methyltransferase.
Length = 360
Score = 94.0 bits (232), Expect(2) = 2e-27
Identities = 53/109 (48%), Positives = 67/109 (61%), Gaps = 1/109 (0%)
Frame = +2
Query: 98 EQQVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKT 277
E + L A +L NH+ GY++SMAL A LG+ DAIH GG AT+ + ++LHPSK
Sbjct: 2 EHEQLLQASTELMNHSLGYIRSMALGCAAKLGVADAIHHAGGRATMDDLRAALSLHPSKL 61
Query: 278 P-CLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGPTG 421
P LRR+MRVL +GVF E DD DD +Y +TP S LLV TG
Sbjct: 62 PYFLRRVMRVLVASGVFAHDEEEDD----DD--IYRVTPVSSLLVTATG 104
Score = 50.8 bits (120), Expect(2) = 2e-27
Identities = 26/80 (32%), Positives = 43/80 (53%), Gaps = 1/80 (1%)
Frame = +3
Query: 420 GKNVNPLMKVMLRP-IFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLF 596
G+++ P + + L P I+V+ + W + +PF +THG +W + + D LF
Sbjct: 108 GRSLLPFVLLQLSPPIYVTPATSMAEWLTSG-EEETPFEMTHGAGLWTVCSRDPELGELF 166
Query: 597 DAGMVADSGFIMNVVVRECG 656
+ M ADS FIM+V +R G
Sbjct: 167 NDAMAADSAFIMDVAIRGAG 186
>sptrnew|CAE03691|CAE03691 OSJNBb0026E15.9 protein.
Length = 311
Score = 95.9 bits (237), Expect(2) = 6e-27
Identities = 53/103 (51%), Positives = 65/103 (63%)
Frame = +2
Query: 104 QVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPC 283
Q L + L +H +GY KSMAL A DLGIP AIH+ GG+ATI IV + P+K P
Sbjct: 11 QDMLQGHIDLHHHLYGYHKSMALLCATDLGIPGAIHRRGGAATISDIVADTMIPPAKLPH 70
Query: 284 LRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVG 412
LRRLMRVL+V+G+F +E VY LTPASR+LVG
Sbjct: 71 LRRLMRVLSVSGIFAVEED-----------VYKLTPASRILVG 102
Score = 47.0 bits (110), Expect(2) = 6e-27
Identities = 25/80 (31%), Positives = 43/80 (53%), Gaps = 3/80 (3%)
Frame = +3
Query: 426 NVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSP---FNVTHGRDIWELDAHDAGFSRLF 596
N +PL+ +++ P +++F L WF+ + + SP F + HG WE+ D +
Sbjct: 108 NFSPLVHLVVSPAMLTTFSSLSPWFR-DGRNASPTALFEMAHGMPPWEMMKRDDTMNSAL 166
Query: 597 DAGMVADSGFIMNVVVRECG 656
+ VADS F+M + +RE G
Sbjct: 167 NDACVADSSFLMEIALRERG 186
>sptr|Q43771|Q43771 Flavonoid 7-O-methyltransferase (EC 2.1.1.6).
Length = 390
Score = 91.7 bits (226), Expect(2) = 3e-26
Identities = 44/107 (41%), Positives = 73/107 (68%)
Frame = +2
Query: 89 ASSEQQVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHP 268
A+S Q+ L AE +L+ H+FGY+KSMAL++ + L IPD +H++GG+A++P++++ + +HP
Sbjct: 29 ANSNGQI-LQAEAELFCHSFGYLKSMALQSVVKLRIPDVLHRYGGAASLPELLSTVPIHP 87
Query: 269 SKTPCLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLV 409
+K P L RLM++L G+F ++ G + +Y L SRLLV
Sbjct: 88 NKLPYLPRLMKMLAAAGIFTAEDVPATVGDGEPTTLYHLNAVSRLLV 134
Score = 48.9 bits (115), Expect(2) = 3e-26
Identities = 25/84 (29%), Positives = 44/84 (52%), Gaps = 2/84 (2%)
Frame = +3
Query: 408 SAQRGKNVNPLMKVMLRPIFVSSFLDLRGWFQHE--MPDPSPFNVTHGRDIWELDAHDAG 581
S G +++P + + P+F+ + L L W Q E +PF + HG ++ + D+
Sbjct: 138 SVNGGASMSPCVLLGTVPLFLGASLKLHEWLQSEEQATTETPFMLAHGGTLYGIGGRDSE 197
Query: 582 FSRLFDAGMVADSGFIMNVVVREC 653
F+ +F+ M A S F+ + VREC
Sbjct: 198 FNTVFNKAMGASSEFVAALAVREC 221
>sptrnew|BAD01311|BAD01311 Putative catechol O-methyltransferase.
Length = 375
Score = 87.4 bits (215), Expect(2) = 4e-26
Identities = 47/100 (47%), Positives = 64/100 (64%), Gaps = 1/100 (1%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A +LWN TF Y+K+MAL+ A+ LGIP+AIH+ GGSA++ ++V I + ++ P L R
Sbjct: 20 LQAHAELWNLTFSYLKAMALECAIKLGIPNAIHRCGGSASLSELVISIPVPETRKPHLPR 79
Query: 293 LMRVLTVTGVFGTQEPHDDGGGCDDEL-VYTLTPASRLLV 409
LMR L GVF P D + + +Y LTP SRLLV
Sbjct: 80 LMRFLAAVGVFSLDNPTIDEEVTEKGMGIYRLTPLSRLLV 119
Score = 52.8 bits (125), Expect(2) = 4e-26
Identities = 24/80 (30%), Positives = 44/80 (55%), Gaps = 3/80 (3%)
Frame = +3
Query: 426 NVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPS---PFNVTHGRDIWELDAHDAGFSRLF 596
+++P + VS+ ++L WF E + + PF HG D+W + + DA + +F
Sbjct: 128 SLSPFVLSQTTKYHVSAAMNLSDWFMTEDKEVAIEMPFRAAHGTDLWGVMSRDANMNEVF 187
Query: 597 DAGMVADSGFIMNVVVRECG 656
+AGM +DS +N ++ +CG
Sbjct: 188 NAGMGSDSRLAINFIISKCG 207
>sptr|Q8GU24|Q8GU24 Orcinol O-methyltransferase.
Length = 367
Score = 82.0 bits (201), Expect(2) = 1e-25
Identities = 40/99 (40%), Positives = 64/99 (64%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A+ +WNH F ++ SM+LK+A+ LGIPD I++HG T+ ++ + + +HP+K+ + R
Sbjct: 24 LHAQAHIWNHIFSFINSMSLKSAIQLGIPDIINKHGYPMTLSELTSALPIHPTKSHSVYR 83
Query: 293 LMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLV 409
LMR+L +G F ++ DE YTLT AS+LL+
Sbjct: 84 LMRILVHSGFFAKKKLSK-----TDEEGYTLTDASQLLL 117
Score = 56.6 bits (135), Expect(2) = 1e-25
Identities = 22/76 (28%), Positives = 42/76 (55%)
Frame = +3
Query: 426 NVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAG 605
++ P + ML P+ + + L WFQ++ DP+PF+ HG W+ H + LF+
Sbjct: 123 SLTPYLTAMLDPVLTNPWNYLSTWFQND--DPTPFDTAHGMTFWDYGNHQPSIAHLFNDA 180
Query: 606 MVADSGFIMNVVVREC 653
M +D+ + +V++ +C
Sbjct: 181 MASDARLVTSVIINDC 196
>sptr|Q8L5K8|Q8L5K8 Orcinol O-methyltransferase.
Length = 367
Score = 82.0 bits (201), Expect(2) = 1e-25
Identities = 40/99 (40%), Positives = 64/99 (64%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A+ +WNH F ++ SM+LK+A+ LGIPD I++HG T+ ++ + + +HP+K+ + R
Sbjct: 24 LHAQAHIWNHIFSFINSMSLKSAIQLGIPDIINKHGYPMTLSELTSALPIHPTKSHSVYR 83
Query: 293 LMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLV 409
LMR+L +G F ++ DE YTLT AS+LL+
Sbjct: 84 LMRILVHSGFFAKKKLSK-----TDEEGYTLTDASQLLL 117
Score = 56.6 bits (135), Expect(2) = 1e-25
Identities = 22/76 (28%), Positives = 42/76 (55%)
Frame = +3
Query: 426 NVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAG 605
++ P + ML P+ + + L WFQ++ DP+PF+ HG W+ H + LF+
Sbjct: 123 SLTPYLTAMLDPVLTNPWNYLSTWFQND--DPTPFDTAHGMTFWDYGNHQPSIAHLFNDA 180
Query: 606 MVADSGFIMNVVVREC 653
M +D+ + +V++ +C
Sbjct: 181 MASDARLVTSVIINDC 196
>sptr|Q8GU21|Q8GU21 Orcinol O-methyltransferase (Fragment).
Length = 348
Score = 80.9 bits (198), Expect(2) = 1e-25
Identities = 42/109 (38%), Positives = 66/109 (60%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A+ +WNH F ++ SM+LK+A+ LGIPD I++HG T+ ++ + + +HP+K+ + R
Sbjct: 14 LHAQAHIWNHIFSFINSMSLKSAIQLGIPDIINKHGYPMTLSELSSALPIHPTKSHSVYR 73
Query: 293 LMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGPTGQEREPF 439
LMR+L +G F ++ DE YTLT AS+LL+ PF
Sbjct: 74 LMRILVHSGFFAKKKLSK-----IDEEGYTLTDASQLLLKDHPLSLTPF 117
Score = 57.8 bits (138), Expect(2) = 1e-25
Identities = 23/76 (30%), Positives = 42/76 (55%)
Frame = +3
Query: 426 NVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAG 605
++ P + ML P+ + L WFQ++ DP+PF+ THG W+ H + LF+
Sbjct: 113 SLTPFLTAMLDPVLTKPWNYLSTWFQND--DPTPFDTTHGMTFWDYGNHQPNIAHLFNDA 170
Query: 606 MVADSGFIMNVVVREC 653
M +D+ + +V++ +C
Sbjct: 171 MASDARLVTSVIIDDC 186
>sptr|Q8GU23|Q8GU23 Orcinol O-methyltransferase.
Length = 366
Score = 79.0 bits (193), Expect(2) = 5e-25
Identities = 42/109 (38%), Positives = 66/109 (60%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A+ +WNH F ++ SM+LK+A+ LGIPD I++H G T+ ++ + + +HP+K+ + R
Sbjct: 24 LHAQAHIWNHIFSFINSMSLKSAIQLGIPDIINKH-GPMTLSELTSALPIHPTKSHSVYR 82
Query: 293 LMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGPTGQEREPF 439
LMR+L +G F ++ DE YTLT AS+LL+ PF
Sbjct: 83 LMRILVHSGFFAKKKLSK-----TDEEGYTLTDASQLLLKDHPLSLTPF 126
Score = 57.4 bits (137), Expect(2) = 5e-25
Identities = 23/76 (30%), Positives = 42/76 (55%)
Frame = +3
Query: 426 NVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAG 605
++ P + ML P+ + + L WFQ+E DP+PF+ HG W+ H + LF+
Sbjct: 122 SLTPFLTAMLDPVLTTPWNYLSTWFQNE--DPTPFDTAHGMTFWDYGNHQPSIAHLFNDA 179
Query: 606 MVADSGFIMNVVVREC 653
M +D+ + +V++ +C
Sbjct: 180 MASDARLVTSVIIDDC 195
>sptr|Q8L5K7|Q8L5K7 Orcinol O-methyltransferase.
Length = 366
Score = 79.0 bits (193), Expect(2) = 5e-25
Identities = 42/109 (38%), Positives = 66/109 (60%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A+ +WNH F ++ SM+LK+A+ LGIPD I++H G T+ ++ + + +HP+K+ + R
Sbjct: 24 LHAQAHIWNHIFSFINSMSLKSAIQLGIPDIINKH-GPMTLSELTSALPIHPTKSHSVYR 82
Query: 293 LMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGPTGQEREPF 439
LMR+L +G F ++ DE YTLT AS+LL+ PF
Sbjct: 83 LMRILVHSGFFAKKKLSK-----TDEEGYTLTDASQLLLKDHPLSLTPF 126
Score = 57.4 bits (137), Expect(2) = 5e-25
Identities = 23/76 (30%), Positives = 42/76 (55%)
Frame = +3
Query: 426 NVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAG 605
++ P + ML P+ + + L WFQ+E DP+PF+ HG W+ H + LF+
Sbjct: 122 SLTPFLTAMLDPVLTTPWNYLSTWFQNE--DPTPFDTAHGMTFWDYGNHQPSIAHLFNDA 179
Query: 606 MVADSGFIMNVVVREC 653
M +D+ + +V++ +C
Sbjct: 180 MASDARLVTSVIIDDC 195
>sptr|Q8GU22|Q8GU22 Orcinol O-methyltransferase (Fragment).
Length = 348
Score = 79.7 bits (195), Expect(2) = 9e-25
Identities = 39/99 (39%), Positives = 64/99 (64%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A+ +WNH F ++ SM+LK+A+ LGIPD I+++G T+ ++ + + +HP+K+ + R
Sbjct: 14 LHAQAHIWNHIFSFINSMSLKSAIQLGIPDIINKYGYPMTLSELTSALPIHPTKSHSVYR 73
Query: 293 LMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLV 409
LMR+L +G F ++ DE YTLT AS+LL+
Sbjct: 74 LMRILVHSGFFAKKKLSK-----TDEEGYTLTDASQLLL 107
Score = 55.8 bits (133), Expect(2) = 9e-25
Identities = 22/76 (28%), Positives = 42/76 (55%)
Frame = +3
Query: 426 NVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAG 605
++ P + ML P+ + + L WFQ++ DP+PF+ HG W+ H + LF+
Sbjct: 113 SLTPYLTAMLDPVLTNPWNYLSTWFQND--DPTPFDTAHGMTFWDYGNHQPSIAHLFNDA 170
Query: 606 MVADSGFIMNVVVREC 653
M +D+ + +V++ +C
Sbjct: 171 MASDARLVTSVIIDDC 186
>sptr|Q9ST37|Q9ST37 O-methyltransferase.
Length = 383
Score = 82.8 bits (203), Expect(2) = 1e-23
Identities = 43/113 (38%), Positives = 65/113 (57%), Gaps = 4/113 (3%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A+ QLWNH F ++ SM+LK A+ LGI D IH HG ++ +++ + +HPSK + R
Sbjct: 15 LGAQAQLWNHIFQFINSMSLKCAVQLGIADVIHNHGQPISLSELIAGLNVHPSKAHFVSR 74
Query: 293 LMRVLTVTGVFGT-QEPHDDGGGCDDE---LVYTLTPASRLLVGPTGQEREPF 439
LM +L + F H D ++E ++Y+LTP+SRLL+ PF
Sbjct: 75 LMLILVHSNFFAQHHHVHHDRADVEEEEAVVLYSLTPSSRLLLKDGPFSTTPF 127
Score = 48.9 bits (115), Expect(2) = 1e-23
Identities = 27/82 (32%), Positives = 38/82 (46%), Gaps = 5/82 (6%)
Frame = +3
Query: 426 NVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDP-----SPFNVTHGRDIWELDAHDAGFSR 590
+ P + L P+ + F + W + D +PF + +G WEL A + F
Sbjct: 123 STTPFLLATLDPVVTTPFHLMGAWLKINGGDDPGATCTPFEMENGMPFWELGAQEPRFGN 182
Query: 591 LFDAGMVADSGFIMNVVVRECG 656
LF+ M ADS I VVV ECG
Sbjct: 183 LFNEAMEADSKLIGRVVVEECG 204
>sptrnew|BAD05699|BAD05699 Putative flavonoid 7-O-methyltransferase.
Length = 371
Score = 87.0 bits (214), Expect(2) = 3e-23
Identities = 50/112 (44%), Positives = 70/112 (62%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A+ +L H + +++SMALK A+DLGIP AIH++GGSA++P ++ + + +K P L R
Sbjct: 26 LQAQAELCCHAWRHMESMALKCAIDLGIPSAIHRNGGSASLPDLLATLPIAENKRPFLHR 85
Query: 293 LMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGPTGQEREPFNES 448
LMR LTV+G+F + DDG VY LT SRLLV PF E+
Sbjct: 86 LMRFLTVSGIFTSA---DDG-------VYQLTRVSRLLVDSIPLNLLPFAET 127
Score = 43.5 bits (101), Expect(2) = 3e-23
Identities = 22/58 (37%), Positives = 32/58 (55%)
Frame = +3
Query: 474 SFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAGMVADSGFIMNVVVR 647
SFL L WF+ + D +PF + HG D+W + D + F A M ADS F+ + +R
Sbjct: 145 SFLCLGEWFR-DGGDTTPFAMAHGIDVWGAMSLDRALAAGFSASMAADSKFLAEIAIR 201
>sptrnew|BAC99560|BAC99560 Putative flavonoid 7-O-methyltransferase.
Length = 371
Score = 87.0 bits (214), Expect(2) = 3e-23
Identities = 50/112 (44%), Positives = 70/112 (62%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A+ +L H + +++SMALK A+DLGIP AIH++GGSA++P ++ + + +K P L R
Sbjct: 26 LQAQAELCCHAWRHMESMALKCAIDLGIPSAIHRNGGSASLPDLLATLPIAENKRPFLHR 85
Query: 293 LMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGPTGQEREPFNES 448
LMR LTV+G+F + DDG VY LT SRLLV PF E+
Sbjct: 86 LMRFLTVSGIFTSA---DDG-------VYQLTRVSRLLVDSIPLNLLPFAET 127
Score = 43.5 bits (101), Expect(2) = 3e-23
Identities = 22/58 (37%), Positives = 32/58 (55%)
Frame = +3
Query: 474 SFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAGMVADSGFIMNVVVR 647
SFL L WF+ + D +PF + HG D+W + D + F A M ADS F+ + +R
Sbjct: 145 SFLCLGEWFR-DGGDTTPFAMAHGIDVWGAMSLDRALAAGFSASMAADSKFLAEIAIR 201
>sptr|Q93WU2|Q93WU2 Eugenol O-methyltransferase.
Length = 357
Score = 82.0 bits (201), Expect(2) = 2e-22
Identities = 46/118 (38%), Positives = 69/118 (58%)
Frame = +2
Query: 56 QKLTISEQQHTASSEQQVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATI 235
QK+ IS S+EQ L A++ +WNH + + SM+LK A+ LGIPD +H+HG T+
Sbjct: 4 QKVDIS-----LSTEQ--LLQAQVHVWNHMYAFANSMSLKCAIQLGIPDILHKHGRPMTL 56
Query: 236 PQIVTRITLHPSKTPCLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLV 409
Q++ I ++ KT C +RLMR L + F ++ + E+ Y LTPAS LL+
Sbjct: 57 SQLLQSIPINKEKTQCFQRLMRALVNSNFF-----IEENNSNNQEVCYWLTPASCLLL 109
Score = 45.4 bits (106), Expect(2) = 2e-22
Identities = 24/74 (32%), Positives = 37/74 (50%)
Frame = +3
Query: 429 VNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAGM 608
V PL++V+L P F + + + WF HE + F +G WE A++ R FD M
Sbjct: 116 VTPLVQVVLDPTFTNPWHHMSEWFTHE-KHATQFEAANGCTFWEKLANEPSKGRFFDEAM 174
Query: 609 VADSGFIMNVVVRE 650
DS I +V ++
Sbjct: 175 SCDSRLIAHVFTKD 188
>sptr|O24249|O24249 Methyltransferase.
Length = 354
Score = 76.3 bits (186), Expect(2) = 8e-22
Identities = 39/109 (35%), Positives = 61/109 (55%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A+ +WNH F ++ S++LK A+ L IPD I +HG T+ ++V+ + + P+K + R
Sbjct: 11 LQAQAHIWNHIFSFINSLSLKCAVQLDIPDVIQKHGQPMTLSELVSALPISPTKAHFIPR 70
Query: 293 LMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGPTGQEREPF 439
LMR+L +G F + GC ++ Y LT AS LL+ PF
Sbjct: 71 LMRILVHSGFFAKESL----SGCGEQ-GYILTDASALLLKDNPMSARPF 114
Score = 49.3 bits (116), Expect(2) = 8e-22
Identities = 22/76 (28%), Positives = 40/76 (52%)
Frame = +3
Query: 426 NVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAG 605
+ P + ML PI + L WFQ++ +P+PF+V +G W+ D + F+
Sbjct: 110 SARPFLLAMLSPILTDPYQYLTTWFQND--NPTPFHVVNGMTCWDYVNQDPTLAHFFNDA 167
Query: 606 MVADSGFIMNVVVREC 653
M +D+ I ++V+ +C
Sbjct: 168 MASDAQLISSLVIDDC 183
>sptr|Q93WU3|Q93WU3 Chavicol O-methyltransferase.
Length = 356
Score = 75.5 bits (184), Expect(2) = 3e-20
Identities = 40/116 (34%), Positives = 65/116 (56%)
Frame = +2
Query: 62 LTISEQQHTASSEQQVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQ 241
+ + + S+EQ L A+ +WNH + + SM+LK A+ LGIPD +H+H T+ Q
Sbjct: 1 MALQNMDISLSTEQ--LLQAQAHVWNHMYAFANSMSLKCAIQLGIPDILHKHDHPMTLSQ 58
Query: 242 IVTRITLHPSKTPCLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLV 409
++ I ++ K+ +RLMR L + F + + + E+ Y LTPASRLL+
Sbjct: 59 LLKAIPINKEKSQSFQRLMRALVNSNFFIEENSN------NQEVCYWLTPASRLLL 108
Score = 44.7 bits (104), Expect(2) = 3e-20
Identities = 23/74 (31%), Positives = 38/74 (51%)
Frame = +3
Query: 429 VNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAGM 608
V PL++V+L P F + + + WF+HE + F +G WE A+ R FD M
Sbjct: 115 VAPLVQVVLDPTFTNPWHYMSEWFKHEN-HATQFEAANGCTFWEKLANKPSMGRFFDEAM 173
Query: 609 VADSGFIMNVVVRE 650
DS + +V+ ++
Sbjct: 174 SCDSRLVAHVLTKD 187
>sptr|O24305|O24305 6a-hydroxymaackiain methyltransferase.
Length = 360
Score = 92.0 bits (227), Expect(2) = 4e-20
Identities = 46/108 (42%), Positives = 67/108 (62%)
Frame = +2
Query: 86 TASSEQQVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLH 265
T SE+ A++ L+ H + +V SMALK+A++LGI DAIH HG T+P++ + + LH
Sbjct: 5 TNGSEESELYHAQIHLYKHVYNFVSSMALKSAMELGIADAIHNHGKPMTLPELSSSLKLH 64
Query: 266 PSKTPCLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLV 409
PSK L R +R+LT G F + G ++E Y LTP+S+LLV
Sbjct: 65 PSKVNILYRFLRLLTHNGFFAKTTVKSNEG--EEETAYVLTPSSKLLV 110
Score = 27.7 bits (60), Expect(2) = 4e-20
Identities = 17/64 (26%), Positives = 29/64 (45%)
Frame = +3
Query: 429 VNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAGM 608
++ L+K L P + + + WF HE + + F G + W+ D+ +F M
Sbjct: 117 LSSLVKGALHPSSLDMWGVSKKWF-HEDKEQTLFECATGENYWDFLNKDSDSLSMFQDAM 175
Query: 609 VADS 620
ADS
Sbjct: 176 AADS 179
>sptr|Q8RV92|Q8RV92 Putative o-methyltransferase.
Length = 378
Score = 98.6 bits (244), Expect = 6e-20
Identities = 53/112 (47%), Positives = 70/112 (62%), Gaps = 3/112 (2%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A+ +LWNH F Y KSM+L+ A++LGIPDA+H+ GG+ T+P++V + L S+ P LRR
Sbjct: 15 LRAQAELWNHIFAYTKSMSLRCAVELGIPDAVHRRGGAVTVPELVAELALPRSREPFLRR 74
Query: 293 LMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLV---GPTGQEREPF 439
LMR+L G+F D G +D Y LT SRLLV G GQ PF
Sbjct: 75 LMRLLAHGGIF------DAAAGAED--AYGLTAVSRLLVSAPGGAGQGLSPF 118
Score = 53.5 bits (127), Expect = 2e-06
Identities = 29/92 (31%), Positives = 43/92 (46%), Gaps = 10/92 (10%)
Frame = +3
Query: 399 VSLSAQRGKNVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPS---------PFNVTHG-R 548
VS G+ ++P + ML PI VS + L WF+ D PF HG R
Sbjct: 105 VSAPGGAGQGLSPFARAMLHPIIVSPSISLASWFRAAAADDDDEGADAPRVPFAAVHGGR 164
Query: 549 DIWELDAHDAGFSRLFDAGMVADSGFIMNVVV 644
++W + D GF F+ M D F+M+V++
Sbjct: 165 ELWAVAKDDPGFGAAFNDAMACDGRFVMDVLL 196
>sptr|O65859|O65859 O-methyltransferase.
Length = 356
Score = 82.0 bits (201), Expect(2) = 1e-19
Identities = 41/109 (37%), Positives = 63/109 (57%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRR 292
L A+ +WN F ++ M+LK A+ LGIPD I +HG ++ +++ + +HP K+ C+ R
Sbjct: 13 LQAQAHIWNCIFSFINPMSLKCAVQLGIPDIIKKHGNPMSLSDLISALPIHPKKSNCVYR 72
Query: 293 LMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGPTGQEREPF 439
LMR+L +G F Q+ + D+E Y LT ASRLL+ PF
Sbjct: 73 LMRILVHSGFFCRQKLSE----LDEEEGYVLTDASRLLLKDDPLSARPF 117
Score = 36.2 bits (82), Expect(2) = 1e-19
Identities = 19/75 (25%), Positives = 31/75 (41%)
Frame = +3
Query: 426 NVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAG 605
+ P + L P + WFQ++ DP+ HG W+ + S +F+
Sbjct: 113 SARPFLLGALDPFMTKPWHYFSTWFQND--DPTACVTAHGTTFWDFGCLEPSLSHIFNDA 170
Query: 606 MVADSGFIMNVVVRE 650
M +D+ I VV E
Sbjct: 171 MASDARLISKVVSNE 185
>sptr|Q84KK4|Q84KK4 S-adenosyl-L-methionine:
2,7,4'-trihydroxyisoflavanone 4'-O-methyltransferase.
Length = 365
Score = 89.0 bits (219), Expect(2) = 2e-19
Identities = 45/105 (42%), Positives = 63/105 (60%)
Frame = +2
Query: 95 SEQQVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSK 274
SE A++ L+ H + +V SMALK+A++LGI D IH HG T+P++ T + L PSK
Sbjct: 9 SEDTELSQAQIHLYKHVYNFVSSMALKSAMELGIADVIHSHGKPITLPELATALNLRPSK 68
Query: 275 TPCLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLV 409
L R +R+LT G F + G G ++E Y LTP S+LLV
Sbjct: 69 IGVLHRFLRLLTHNGFF-AKTTVSRGEGAEEETAYGLTPPSKLLV 112
Score = 28.5 bits (62), Expect(2) = 2e-19
Identities = 17/75 (22%), Positives = 32/75 (42%), Gaps = 2/75 (2%)
Frame = +3
Query: 435 PLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWEL--DAHDAGFSRLFDAGM 608
P++K L P + + + WF + + + F G WE ++ +F M
Sbjct: 121 PIVKGALHPSSLDMWRSSKKWFLEDNEELTLFESATGESFWEFLNKETESDTLSMFQEAM 180
Query: 609 VADSGFIMNVVVREC 653
ADS + + ++EC
Sbjct: 181 AADS-HMFKLALKEC 194
>sptr|Q84KK6|Q84KK6 S-adenosyl-L-methionine:
2,7,4'-trihydroxyisoflavanone 4'-O-methyltransferase.
Length = 367
Score = 90.9 bits (224), Expect = 1e-17
Identities = 48/111 (43%), Positives = 68/111 (61%), Gaps = 3/111 (2%)
Frame = +2
Query: 86 TASSEQQVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLH 265
T SE+ A++ L+ H + +V SMALK+A++LGI D IH HG T+P++ + + LH
Sbjct: 5 TNGSEEIELYHAQIHLYKHVYNFVSSMALKSAMELGIADVIHNHGKPITLPELASALKLH 64
Query: 266 PSKTPCLRRLMRVLTVTGVFG-TQEPHDDG--GGCDDELVYTLTPASRLLV 409
PSK L R +R+LT G F T P +G G ++E Y LTP S+LLV
Sbjct: 65 PSKVGILYRFLRLLTHNGFFAKTTVPSQNGKDGEEEEETAYALTPPSKLLV 115
>sptr|Q8GSN1|Q8GSN1 Flavonoid O-methyltransferase.
Length = 348
Score = 70.9 bits (172), Expect(2) = 9e-17
Identities = 35/98 (35%), Positives = 53/98 (54%)
Frame = +2
Query: 116 DAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKTPCLRRL 295
+A+ + F + +LK A+ LGIPDAIH HG + + + ++PSK P + RL
Sbjct: 10 NAQAHFFTQVFSFTSMSSLKCAVQLGIPDAIHSHGKPMALSDLTNSLPINPSKAPYIYRL 69
Query: 296 MRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLV 409
MR+L G F +E + VY+LTP +RLL+
Sbjct: 70 MRILVAAGYFSEEEKN----------VYSLTPFTRLLL 97
Score = 37.7 bits (86), Expect(2) = 9e-17
Identities = 18/52 (34%), Positives = 29/52 (55%)
Frame = +3
Query: 495 WFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAGMVADSGFIMNVVVRE 650
WFQ+E D + F HG++ W+ A D + + FD M ADS + +++ E
Sbjct: 126 WFQNE--DLTAFETAHGKNFWDFGAEDK-YGKNFDGVMAADSILVSKMLIPE 174
>sptr|Q8LI58|Q8LI58 OJ1634_H04.28 protein.
Length = 341
Score = 87.0 bits (214), Expect = 2e-16
Identities = 49/114 (42%), Positives = 67/114 (58%)
Frame = +2
Query: 98 EQQVALDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRITLHPSKT 277
E + + A +L +H+ GYV+SMAL A LG+ DAIH+ GG AT+ + ++LHP+K
Sbjct: 2 EHEQLVQASTELMHHSLGYVRSMALGCAAKLGVADAIHRAGGRATLHDLHAALSLHPTKL 61
Query: 278 PCLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLVGPTGQEREPF 439
P LRR+MRVL +GVF + +D Y LTP S LLV G+ PF
Sbjct: 62 PFLRRVMRVLVASGVFAQVKEEEDH--------YRLTPVSSLLV-TAGRTLLPF 106
Score = 36.2 bits (82), Expect = 0.39
Identities = 20/82 (24%), Positives = 35/82 (42%)
Frame = +3
Query: 402 SLSAQRGKNVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAG 581
SL G+ + P + + P+ V+ + W + + + F + HG +W
Sbjct: 94 SLLVTAGRTLLPFVLLQHSPLCVTPATSMAEWLKTG-EEETAFEMAHGAGLWGACRRAPE 152
Query: 582 FSRLFDAGMVADSGFIMNVVVR 647
F+ M ADS FIM+ +R
Sbjct: 153 LGDFFNDAMAADSAFIMDAAIR 174
>sptrnew|AAQ01668|AAQ01668 (R,S)-reticuline 7-O-methyltransferase.
Length = 355
Score = 68.6 bits (166), Expect(2) = 1e-14
Identities = 42/103 (40%), Positives = 62/103 (60%), Gaps = 4/103 (3%)
Frame = +2
Query: 113 LDAELQLWNHTFGYVKSMALKAALDLGIPDAIHQHGGSATIPQIVTRI-TLHPSKTP--- 280
L + ++W H F +V SMALK A++LGIPD I+ HG TI +IV + T PS +P
Sbjct: 8 LKGQAEIWEHMFAFVDSMALKCAVELGIPDIINSHGRPVTISEIVDSLKTNTPSSSPNID 67
Query: 281 CLRRLMRVLTVTGVFGTQEPHDDGGGCDDELVYTLTPASRLLV 409
L R+MR+L +F T E H + ++L+Y LT +S+ L+
Sbjct: 68 YLTRIMRLLVHKRLF-TSELHQE----SNQLLYNLTRSSKWLL 105
Score = 33.1 bits (74), Expect(2) = 1e-14
Identities = 21/75 (28%), Positives = 35/75 (46%)
Frame = +3
Query: 426 NVNPLMKVMLRPIFVSSFLDLRGWFQHEMPDPSPFNVTHGRDIWELDAHDAGFSRLFDAG 605
N++PL+ PI + + L Q + SPF HG +IW+L D F+ +
Sbjct: 111 NLSPLVLWETNPILLKPWQYLGKCAQEKS---SPFERAHGCEIWDLALADPKFNNFLNGA 167
Query: 606 MVADSGFIMNVVVRE 650
M + I+N ++ E
Sbjct: 168 MQCSTTTIINEMLLE 182
Database: swall
Posted date: Feb 26, 2004 3:00 PM
Number of letters in database: 439,479,560
Number of sequences in database: 1,381,838
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 470,883,751
Number of Sequences: 1381838
Number of extensions: 9733380
Number of successful extensions: 35835
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 34383
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 35767
length of database: 439,479,560
effective HSP length: 118
effective length of database: 276,422,676
effective search space used: 27642267600
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

