BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
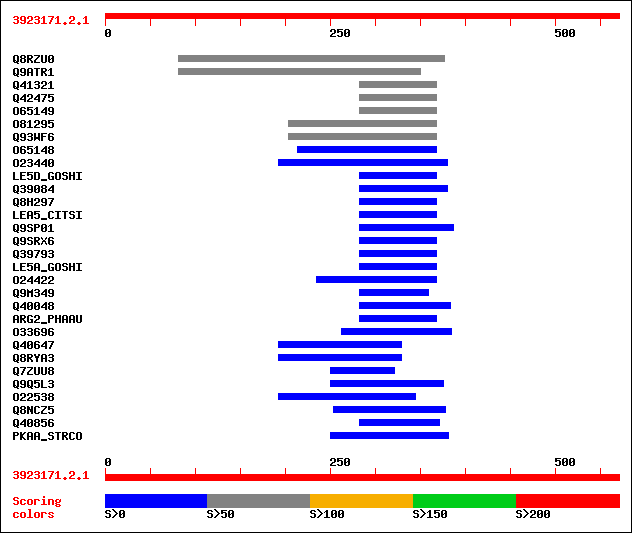
Query= 3923171.2.1
(570 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q8RZU0|Q8RZU0 Zinc-induced protein-like. 85 7e-16
sptr|Q9ATR1|Q9ATR1 Zinc-induced protein. 73 3e-12
sptr|Q41321|Q41321 Protein induced upon tuberization. 54 2e-06
sptr|Q42475|Q42475 Heat shock protein. 54 2e-06
sptr|O65149|O65149 Late embryogenis abundant protein 5. 53 3e-06
sptr|O81295|O81295 T14P8.2 protein (AT4G02380 protein). 51 1e-05
sptr|Q93WF6|Q93WF6 AT4g02380/T14P8_2 (Late embryogenis abundant ... 51 1e-05
sptr|O65148|O65148 Late embryogenesis abundant protein homolog (... 50 2e-05
sptr|O23440|O23440 Drought-induced protein (Drought-induced prot... 48 9e-05
sw|P46522|LE5D_GOSHI Late embryogenesis abundant protein Lea5-D. 47 1e-04
sptr|Q39084|Q39084 Di21 protein (At4g15910). 47 2e-04
sptr|Q8H297|Q8H297 Late-embryogenesis abundant protein-like prot... 47 2e-04
sw|Q39644|LEA5_CITSI Late embryogenesis abundant protein Lea5. 46 3e-04
sptr|Q9SP01|Q9SP01 Indole-3-acetic acid induced protein ARG-2 ho... 46 3e-04
sptr|Q9SRX6|Q9SRX6 F22D16.18 protein (Late embryogenis abundant ... 46 3e-04
sptr|Q39793|Q39793 Late embryogenesis abundant protein. 45 4e-04
sw|P46521|LE5A_GOSHI Late embryogenesis abundant protein Lea5-A. 45 4e-04
sptr|O24422|O24422 Desiccation protective protein LEA5. 45 6e-04
sptr|Q9M349|Q9M349 Hypothetical 14.4 kDa protein. 43 0.002
sptr|Q40048|Q40048 ORF1 protein. 43 0.003
sw|P32292|ARG2_PHAAU Indole-3-acetic acid induced protein ARG2. 43 0.003
sptr|O33696|O33696 15.5 kDa protein precursor. 35 0.46
sptr|Q40647|Q40647 Zn-induced protein. 35 0.78
sptr|Q8RYA3|Q8RYA3 Ramy1. 35 0.78
sptr|Q7ZUU8|Q7ZUU8 Similar to adaptor protein complex AP-1, gamm... 34 1.0
sptr|Q9Q5L3|Q9Q5L3 EBNA2-like protein (EBNA-2). 34 1.3
sptr|O22538|O22538 Zinc inducible protein. 34 1.3
sptr|Q8NCZ5|Q8NCZ5 Hypothetical protein (Fragment). 33 2.3
sptr|Q40856|Q40856 EMB32 protein. 33 3.0
sw|P54739|PKAA_STRCO Serine/threonine protein kinase pkaA (EC 2.... 33 3.0
>sptr|Q8RZU0|Q8RZU0 Zinc-induced protein-like.
Length = 93
Score = 84.7 bits (208), Expect = 7e-16
Identities = 52/99 (52%), Positives = 56/99 (56%)
Frame = +3
Query: 81 MERVASSYCVSLLAQSXXXXXXXXXXXXXXXXXXXVDEKKAAAALTKRVMGKKEVNTXXX 260
MERVASS C+SLLAQ DEKK AAA+ KR M K
Sbjct: 1 MERVASS-CLSLLAQRRGYSVAAAVAKGAGRRA---DEKKVAAAVAKRTMAK-------- 48
Query: 261 XXXXXXXWVPDPVTGYYRPAGSAKEVDAAELRAKLLTEA 377
WVPDPVTGYYRPAG AKEVDAAELRAKLL+ +
Sbjct: 49 AAEEKTAWVPDPVTGYYRPAGGAKEVDAAELRAKLLSNS 87
>sptr|Q9ATR1|Q9ATR1 Zinc-induced protein.
Length = 117
Score = 72.8 bits (177), Expect = 3e-12
Identities = 45/90 (50%), Positives = 48/90 (53%)
Frame = +3
Query: 81 MERVASSYCVSLLAQSXXXXXXXXXXXXXXXXXXXVDEKKAAAALTKRVMGKKEVNTXXX 260
MERVASS C+SLLAQ DEKK AAA+ KR M K
Sbjct: 1 MERVASS-CLSLLAQRRGYSVAAAVAKGAGRRA---DEKKVAAAVAKRTMAK-------- 48
Query: 261 XXXXXXXWVPDPVTGYYRPAGSAKEVDAAE 350
WVPDPVTGYYRPAG AKEVDAA+
Sbjct: 49 AAEEKTAWVPDPVTGYYRPAGGAKEVDAAD 78
>sptr|Q41321|Q41321 Protein induced upon tuberization.
Length = 98
Score = 53.5 bits (127), Expect = 2e-06
Identities = 23/29 (79%), Positives = 24/29 (82%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
WVPDPVTGYYRP G A E+DAAELR LL
Sbjct: 63 WVPDPVTGYYRPEGQANEIDAAELRKMLL 91
>sptr|Q42475|Q42475 Heat shock protein.
Length = 97
Score = 53.5 bits (127), Expect = 2e-06
Identities = 23/29 (79%), Positives = 24/29 (82%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
WVPDPVTGYYRP G A E+DAAELR LL
Sbjct: 62 WVPDPVTGYYRPEGQANEIDAAELRKMLL 90
>sptr|O65149|O65149 Late embryogenis abundant protein 5.
Length = 97
Score = 52.8 bits (125), Expect = 3e-06
Identities = 23/29 (79%), Positives = 24/29 (82%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
WVPDPVTGYYRP AKE+DAAELR LL
Sbjct: 62 WVPDPVTGYYRPESHAKEIDAAELRQMLL 90
>sptr|O81295|O81295 T14P8.2 protein (AT4G02380 protein).
Length = 206
Score = 50.8 bits (120), Expect = 1e-05
Identities = 27/55 (49%), Positives = 31/55 (56%)
Frame = +3
Query: 204 AAALTKRVMGKKEVNTXXXXXXXXXXWVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
+ A+ VM KK V WVPDP TGYYRP + E+DAAELRA LL
Sbjct: 152 SGAVASAVMKKKGVEESTQKIS----WVPDPKTGYYRPETGSNEIDAAELRAALL 202
>sptr|Q93WF6|Q93WF6 AT4g02380/T14P8_2 (Late embryogenis abundant
protein).
Length = 97
Score = 50.8 bits (120), Expect = 1e-05
Identities = 27/55 (49%), Positives = 31/55 (56%)
Frame = +3
Query: 204 AAALTKRVMGKKEVNTXXXXXXXXXXWVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
+ A+ VM KK V WVPDP TGYYRP + E+DAAELRA LL
Sbjct: 43 SGAVASAVMKKKGVEESTQKIS----WVPDPKTGYYRPETGSNEIDAAELRAALL 93
>sptr|O65148|O65148 Late embryogenesis abundant protein homolog
(Fragment).
Length = 72
Score = 49.7 bits (117), Expect = 2e-05
Identities = 27/52 (51%), Positives = 29/52 (55%)
Frame = +3
Query: 213 LTKRVMGKKEVNTXXXXXXXXXXWVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
L VM KK V WVPDP TGYYRP + E+DAAELRA LL
Sbjct: 21 LLSAVMKKKGVEESTQKIS----WVPDPKTGYYRPETGSNEIDAAELRAALL 68
>sptr|O23440|O23440 Drought-induced protein (Drought-induced protein
like).
Length = 87
Score = 47.8 bits (112), Expect = 9e-05
Identities = 32/68 (47%), Positives = 35/68 (51%), Gaps = 5/68 (7%)
Frame = +3
Query: 192 EKKAAAALTK----RVM-GKKEVNTXXXXXXXXXXWVPDPVTGYYRPAGSAKEVDAAELR 356
E AA L+K RVM GK E W PDPVTGYYRP+ A E+D AELR
Sbjct: 19 ENVTAAGLSKGGSTRVMVGKMEQR--GLDQEAESAWGPDPVTGYYRPSNRAAEIDPAELR 76
Query: 357 AKLLTEAA 380
LL A
Sbjct: 77 ELLLKNKA 84
>sw|P46522|LE5D_GOSHI Late embryogenesis abundant protein Lea5-D.
Length = 105
Score = 47.4 bits (111), Expect = 1e-04
Identities = 20/29 (68%), Positives = 21/29 (72%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
W PDPVTGYYRP E+DAAELR LL
Sbjct: 70 WAPDPVTGYYRPENCGAEIDAAELREMLL 98
>sptr|Q39084|Q39084 Di21 protein (At4g15910).
Length = 104
Score = 47.0 bits (110), Expect = 2e-04
Identities = 21/33 (63%), Positives = 23/33 (69%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLLTEAA 380
W PDPVTGYYRP+ A E+D AELR LL A
Sbjct: 69 WGPDPVTGYYRPSNRAAEIDPAELRELLLKNKA 101
>sptr|Q8H297|Q8H297 Late-embryogenesis abundant protein-like protein
(Fragment).
Length = 93
Score = 47.0 bits (110), Expect = 2e-04
Identities = 21/29 (72%), Positives = 23/29 (79%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
W+PDPVTGYYRPA A E+DAA LR LL
Sbjct: 61 WLPDPVTGYYRPANRAGEMDAAGLREVLL 89
>sw|Q39644|LEA5_CITSI Late embryogenesis abundant protein Lea5.
Length = 97
Score = 46.2 bits (108), Expect = 3e-04
Identities = 19/29 (65%), Positives = 21/29 (72%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
W PDP+TGYYRP A E+D AELR LL
Sbjct: 62 WAPDPITGYYRPENRAVEIDPAELREMLL 90
>sptr|Q9SP01|Q9SP01 Indole-3-acetic acid induced protein ARG-2
homolog.
Length = 86
Score = 45.8 bits (107), Expect = 3e-04
Identities = 22/35 (62%), Positives = 26/35 (74%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLLTEAAAN 386
WVPDPVTGYY+P + KEVD A+LRA LL + N
Sbjct: 53 WVPDPVTGYYKPE-NIKEVDVADLRATLLRKKFNN 86
>sptr|Q9SRX6|Q9SRX6 F22D16.18 protein (Late embryogenis abundant
protein, putative).
Length = 91
Score = 45.8 bits (107), Expect = 3e-04
Identities = 20/29 (68%), Positives = 23/29 (79%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
WVPDP TGYYRP ++E+D AELRA LL
Sbjct: 59 WVPDPKTGYYRPETVSEEIDPAELRAILL 87
>sptr|Q39793|Q39793 Late embryogenesis abundant protein.
Length = 100
Score = 45.4 bits (106), Expect = 4e-04
Identities = 18/29 (62%), Positives = 21/29 (72%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
W PDPVTGYYRP E+DAA+LR +L
Sbjct: 65 WAPDPVTGYYRPENCGAEIDAADLREMML 93
>sw|P46521|LE5A_GOSHI Late embryogenesis abundant protein Lea5-A.
Length = 105
Score = 45.4 bits (106), Expect = 4e-04
Identities = 18/29 (62%), Positives = 21/29 (72%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
W PDPVTGYYRP E+DAA+LR +L
Sbjct: 70 WAPDPVTGYYRPENCGAEIDAADLREMML 98
>sptr|O24422|O24422 Desiccation protective protein LEA5.
Length = 113
Score = 45.1 bits (105), Expect = 6e-04
Identities = 19/45 (42%), Positives = 25/45 (55%)
Frame = +3
Query: 234 KKEVNTXXXXXXXXXXWVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
+++ +T W PDPVTGYYRP E+D ELR +LL
Sbjct: 55 EEKPDTRDGSKAYSTDWAPDPVTGYYRPINHTPEIDPVELRHRLL 99
>sptr|Q9M349|Q9M349 Hypothetical 14.4 kDa protein.
Length = 124
Score = 43.1 bits (100), Expect = 0.002
Identities = 17/26 (65%), Positives = 21/26 (80%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRA 359
W+PDP TGYYRP A+E+DA ELR+
Sbjct: 86 WMPDPQTGYYRPDNFARELDAVELRS 111
Score = 32.3 bits (72), Expect = 3.9
Identities = 12/20 (60%), Positives = 15/20 (75%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVD 341
W DP TGY+RP +AKE+D
Sbjct: 46 WASDPDTGYFRPETAAKELD 65
>sptr|Q40048|Q40048 ORF1 protein.
Length = 95
Score = 42.7 bits (99), Expect = 0.003
Identities = 19/34 (55%), Positives = 23/34 (67%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLLTEAAA 383
WVPDPVTG+YRPA + D A+LRA L + A
Sbjct: 60 WVPDPVTGHYRPANRSSGADPADLRAAHLGQTYA 93
>sw|P32292|ARG2_PHAAU Indole-3-acetic acid induced protein ARG2.
Length = 99
Score = 42.7 bits (99), Expect = 0.003
Identities = 18/29 (62%), Positives = 23/29 (79%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLL 368
WVPDPVTGYYRP + E+D A++RA +L
Sbjct: 66 WVPDPVTGYYRPE-NTNEIDVADMRATVL 93
>sptr|O33696|O33696 15.5 kDa protein precursor.
Length = 155
Score = 35.4 bits (80), Expect = 0.46
Identities = 20/45 (44%), Positives = 25/45 (55%), Gaps = 4/45 (8%)
Frame = -1
Query: 384 WPQPPSAAWRVAPPRQPPWR-YLLG---DSSR*QGPAPTPSPQLP 262
W P+ + P QPP+ LLG D+ R +GP PTPSPQ P
Sbjct: 69 WAPYPNPHYSENDPHQPPYGGSLLGLSSDNERERGPRPTPSPQSP 113
>sptr|Q40647|Q40647 Zn-induced protein.
Length = 218
Score = 34.7 bits (78), Expect = 0.78
Identities = 18/46 (39%), Positives = 22/46 (47%)
Frame = +3
Query: 192 EKKAAAALTKRVMGKKEVNTXXXXXXXXXXWVPDPVTGYYRPAGSA 329
E K AA +T+ G WVPDPVTG+YRP+ A
Sbjct: 47 EGKGAAGMTQAKDGSSS------SAAREVSWVPDPVTGHYRPSNFA 86
>sptr|Q8RYA3|Q8RYA3 Ramy1.
Length = 218
Score = 34.7 bits (78), Expect = 0.78
Identities = 18/46 (39%), Positives = 22/46 (47%)
Frame = +3
Query: 192 EKKAAAALTKRVMGKKEVNTXXXXXXXXXXWVPDPVTGYYRPAGSA 329
E K AA +T+ G WVPDPVTG+YRP+ A
Sbjct: 48 EGKGAAGMTQAKDGSSS------SAAREVSWVPDPVTGHYRPSNFA 87
>sptr|Q7ZUU8|Q7ZUU8 Similar to adaptor protein complex AP-1, gamma 1
subunit.
Length = 819
Score = 34.3 bits (77), Expect = 1.0
Identities = 15/24 (62%), Positives = 17/24 (70%)
Frame = -1
Query: 321 LLGDSSR*QGPAPTPSPQLPQRQP 250
LLGD S GPAPTP+P +P QP
Sbjct: 658 LLGDLSLTGGPAPTPAPSVPMSQP 681
>sptr|Q9Q5L3|Q9Q5L3 EBNA2-like protein (EBNA-2).
Length = 605
Score = 33.9 bits (76), Expect = 1.3
Identities = 21/49 (42%), Positives = 23/49 (46%), Gaps = 7/49 (14%)
Frame = -1
Query: 375 PPSAAWRVAPPR--QPPWRYLLGDSSR*QGPAPTPSP-----QLPQRQP 250
PPSA WR +PPR QP W Y + Q P P P Q P R P
Sbjct: 237 PPSAPWRPSPPRPPQPHWAYRPHYTQHTQPPWPPYRPPPTKHQAPPRVP 285
>sptr|O22538|O22538 Zinc inducible protein.
Length = 221
Score = 33.9 bits (76), Expect = 1.3
Identities = 19/51 (37%), Positives = 23/51 (45%)
Frame = +3
Query: 192 EKKAAAALTKRVMGKKEVNTXXXXXXXXXXWVPDPVTGYYRPAGSAKEVDA 344
E K AA +T+ G WVPDPV G+YRP+ A DA
Sbjct: 47 EGKGAAGITQAKDG------WSSSAAREVSWVPDPVRGHYRPSNFAGGADA 91
>sptr|Q8NCZ5|Q8NCZ5 Hypothetical protein (Fragment).
Length = 150
Score = 33.1 bits (74), Expect = 2.3
Identities = 18/44 (40%), Positives = 23/44 (52%), Gaps = 2/44 (4%)
Frame = -1
Query: 378 QPPSAAWRVAPPRQP-PWRYLLGDSSR*Q-GPAPTPSPQLPQRQ 253
+PP WR+ PPR+P P R + G P P PSP P R+
Sbjct: 8 RPPHLTWRLLPPRRPAPQREINGPRPPLHPSPPPRPSPGSPGRE 51
>sptr|Q40856|Q40856 EMB32 protein.
Length = 95
Score = 32.7 bits (73), Expect = 3.0
Identities = 15/30 (50%), Positives = 18/30 (60%)
Frame = +3
Query: 282 WVPDPVTGYYRPAGSAKEVDAAELRAKLLT 371
W+ DP TG + P E D AELR KLL+
Sbjct: 63 WMRDPATGNWIPEDHFGETDTAELRQKLLS 92
>sw|P54739|PKAA_STRCO Serine/threonine protein kinase pkaA (EC
2.7.1.37).
Length = 543
Score = 32.7 bits (73), Expect = 3.0
Identities = 20/55 (36%), Positives = 25/55 (45%), Gaps = 11/55 (20%)
Frame = -1
Query: 381 PQPPSAAWRVAPPRQP---------PWRYLLGDSSR*Q--GPAPTPSPQLPQRQP 250
PQ P A + PP+QP P RY + Q P P P PQ P+R+P
Sbjct: 413 PQQPRQAPQGPPPQQPGYGYPQQQQPQRYATPQPQQPQRYAPPPAPEPQQPRREP 467
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 326,954,936
Number of Sequences: 1272877
Number of extensions: 5765049
Number of successful extensions: 18123
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 16619
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 17874
length of database: 406,886,216
effective HSP length: 116
effective length of database: 259,232,484
effective search space used: 18923971332
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

