BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
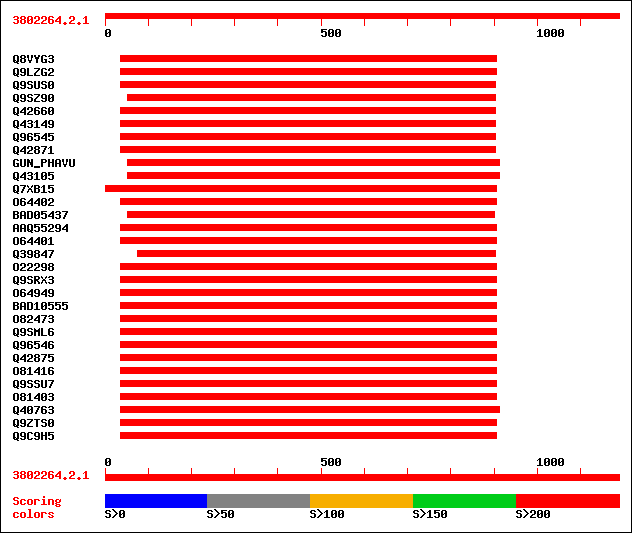
Query= 3802264.2.1
(1189 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q8VYG3|Q8VYG3 Putative cellulase. 336 5e-91
sptr|Q9LZG2|Q9LZG2 Cellulase-like protein. 328 8e-89
sptr|Q9SUS0|Q9SUS0 Putative cellulase. 244 2e-63
sptr|Q9SZ90|Q9SZ90 Cellulase-like protein. 240 4e-62
sptr|Q42660|Q42660 Beta-1,4-endoglycanohydrolase precursor (EC 3... 237 3e-61
sptr|Q43149|Q43149 Cellulase (EC 3.2.1.4) (Fragment). 236 4e-61
sptr|Q96545|Q96545 Cellulase (EC 3.2.1.4). 235 1e-60
sptr|Q42871|Q42871 Endo-1,4-beta-glucanase precursor (EC 3.2.1.4). 232 1e-59
sw|P22503|GUN_PHAVU Endoglucanase precursor (EC 3.2.1.4) (Endo-1... 229 5e-59
sptr|Q43105|Q43105 Cellulase (EC 3.2.1.4). 229 6e-59
sptr|Q7XB15|Q7XB15 Endo-1,4-beta-glucanase (EC 3.2.1.4). 227 2e-58
sptr|O64402|O64402 Endo-beta-1,4-glucanase. 227 3e-58
sptrnew|BAD05437|BAD05437 Putative cellulase. 226 4e-58
sptrnew|AAQ55294|AAQ55294 Endo-1,4-beta-glucanase. 224 3e-57
sptr|O64401|O64401 Endo-beta-1,4-glucanase. 221 2e-56
sptr|Q39847|Q39847 CMCase, cellulase, endo-1,4-beta-D-glucanase ... 220 3e-56
sptr|O22298|O22298 Basic cellulase. 219 9e-56
sptr|Q9SRX3|Q9SRX3 Endo-1,4-beta glucanase. 217 2e-55
sptr|O64949|O64949 Endo-1,4-beta glucanase. 216 4e-55
sptrnew|BAD10555|BAD10555 Putative endoglucanase 1 (Endo-1,4-bet... 216 4e-55
sptr|O82473|O82473 Endo-1,4-beta-glucanase (EC 3.2.1.4). 216 7e-55
sptr|Q9SML6|Q9SML6 Endo-beta-1,4-glucanase precursor (EC 3.2.1.4). 215 9e-55
sptr|Q96546|Q96546 Endo-beta-1,4-glucanase (EC 3.2.1.4). 215 1e-54
sptr|Q42875|Q42875 Endo-1,4-beta-glucanase precursor (EC 3.2.1.4). 215 1e-54
sptr|O81416|O81416 T2H3.5 protein (Putative endo-1, 4-beta gluca... 214 2e-54
sptr|Q9SSU7|Q9SSU7 Endo-1,4-beta-glucanase. 214 2e-54
sptr|O81403|O81403 Endo-1,4-beta-glucanase precursor (EC 3.2.1.4). 213 5e-54
sptr|Q40763|Q40763 Cellulase precursor. 212 8e-54
sptr|Q9ZTS0|Q9ZTS0 Putative cellulase. 212 8e-54
sptr|Q9C9H5|Q9C9H5 Putative beta-glucanase, 74324-76084. 212 1e-53
>sptr|Q8VYG3|Q8VYG3 Putative cellulase.
Length = 486
Score = 336 bits (861), Expect = 5e-91
Identities = 164/291 (56%), Positives = 201/291 (69%), Gaps = 1/291 (0%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGECPFYCSYSGYNDEXXXXXXXXXXXX 213
+VFR D KY+R+LLNKAKLLF AKSH G+YDGECPFYCS SGYNDE
Sbjct: 194 IVFRHVDHKYARRLLNKAKLLFKLAKSHKGTYDGECPFYCSNSGYNDELIWAATWLYKAT 253
Query: 214 RRQVYADFIAHEAISSSVAEFSWDLKFPGAQVLLAEFNMTSAGGAQNFKSQADNFVCAVL 393
R +Y ++ EAIS+ VAEFSWDLK+ GAQ+L+ + G +K QAD+FVC+ L
Sbjct: 254 RNHLYLSYLKFEAISAYVAEFSWDLKYAGAQILITKLIFEGHKGLDLYKQQADSFVCSNL 313
Query: 394 PDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLCGNQPLPPARL 573
P + +HQVF TPGG+IHLRDGANSQYVT+TAFL YAD+L + Q + CG+ L
Sbjct: 314 PGSPYHQVFTTPGGMIHLRDGANSQYVTATAFLFSAYADILQKHNQKISCGSHQFDSTHL 373
Query: 574 HEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVLPPGTDVNCGLSFGDWMA 753
FA++Q+DY+LG NP+ SY+VGFG NPP Q HHRGAS P+ ++C LSF W
Sbjct: 374 MAFAKKQIDYILGHNPQGRSYMVGFGPNPPKQAHHRGASVPMHEANAPLSCPLSFVKWYN 433
Query: 754 PDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAALA 906
+ PN NELTGAI+GGPD+ D F D R S YTEPCTYINS+A+G LA LA
Sbjct: 434 KNVPNANELTGAILGGPDRQDKFQDLRWTSVYTEPCTYINSIAVGVLAKLA 484
>sptr|Q9LZG2|Q9LZG2 Cellulase-like protein.
Length = 483
Score = 328 bits (842), Expect = 8e-89
Identities = 163/291 (56%), Positives = 200/291 (68%), Gaps = 1/291 (0%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGECPFYCSYSGYNDEXXXXXXXXXXXX 213
+VFR D KY+R+LLNKAKLL AKSH G+YDGECPFYCS SGYNDE
Sbjct: 194 IVFRHVDHKYARRLLNKAKLL---AKSHKGTYDGECPFYCSNSGYNDELIWAATWLYKAT 250
Query: 214 RRQVYADFIAHEAISSSVAEFSWDLKFPGAQVLLAEFNMTSAGGAQNFKSQADNFVCAVL 393
R +Y ++ EAIS+ VAEFSWDLK+ GAQ+L+ + G +K QAD+FVC+ L
Sbjct: 251 RNHLYLSYLKFEAISAYVAEFSWDLKYAGAQILITKLIFEGHKGLDLYKQQADSFVCSNL 310
Query: 394 PDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLCGNQPLPPARL 573
P + +HQVF TPGG+IHLRDGANSQYVT+TAFL YAD+L + Q + CG+ L
Sbjct: 311 PGSPYHQVFTTPGGMIHLRDGANSQYVTATAFLFSAYADILQKHNQKISCGSHQFDSTHL 370
Query: 574 HEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVLPPGTDVNCGLSFGDWMA 753
FA++Q+DY+LG NP+ SY+VGFG NPP Q HHRGAS P+ ++C LSF W
Sbjct: 371 MAFAKKQIDYILGHNPQGRSYMVGFGPNPPKQAHHRGASVPMHEANAPLSCPLSFVKWYN 430
Query: 754 PDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAALA 906
+ PN NELTGAI+GGPD+ D F D R S YTEPCTYINS+A+G LA LA
Sbjct: 431 KNVPNANELTGAILGGPDRQDKFQDLRWTSVYTEPCTYINSIAVGVLAKLA 481
>sptr|Q9SUS0|Q9SUS0 Putative cellulase.
Length = 479
Score = 244 bits (623), Expect = 2e-63
Identities = 130/295 (44%), Positives = 178/295 (60%), Gaps = 6/295 (2%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGECPFYCSYSGYNDEXXXXXXXXXXXX 213
+VF+ D YS LLN AK LF FA + GSY CPFYCSYSGY DE
Sbjct: 183 LVFKSVDSTYSSTLLNHAKTLFEFADKYRGSYQASCPFYCSYSGYQDELLWAAAWLYKAT 242
Query: 214 RRQVYADF-IAHEAISSSVAEFSWDLKFPGAQVLLAEFNMTSAGGAQNFKSQADNFVCAV 390
++Y ++ I+++ S +V EFSWD KF GAQ LL A FKS ++FVCA+
Sbjct: 243 GDKIYINYVISNKDWSQAVNEFSWDNKFVGAQALLVSEFYNGANDLAKFKSDVESFVCAM 302
Query: 391 LPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTG-QTVLCGNQPLPPA 567
+P ++ Q+ TPGG++ +RD +N QYVT+ +L Y+ L + G ++ CG+ +
Sbjct: 303 MPGSSSQQIKPTPGGLLFIRDSSNLQYVTTATTVLFHYSKTLTKAGVGSIQCGSTKFTVS 362
Query: 568 RLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVL---PPGTDVNCGLSF 738
++ FA+ Q+DY+LG NP SY+VGFG PTQPHHRG+S P + P D N G S+
Sbjct: 363 QIRNFAKSQVDYILGNNPMKMSYMVGFGTKYPTQPHHRGSSLPSIQSKPEKIDCNGGYSY 422
Query: 739 GDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAAL 903
+ D PNPN GAIVGGP+ +D + DK+ + S+ EP TYIN+ IGP+AAL
Sbjct: 423 YN---SDTPNPNVHIGAIVGGPNSSDQYSDKKSDYSHAEPTTYINAAFIGPVAAL 474
>sptr|Q9SZ90|Q9SZ90 Cellulase-like protein.
Length = 480
Score = 240 bits (612), Expect = 4e-62
Identities = 127/291 (43%), Positives = 176/291 (60%), Gaps = 7/291 (2%)
Frame = +1
Query: 52 DKKYSRKLLNKAKL--LFTFAKSHLGSYDGECPFYCSYSGYNDEXXXXXXXXXXXXRRQV 225
D YS KLLN AK LF FA + GSY CPFYCS+SGY DE +
Sbjct: 189 DSTYSSKLLNNAKSVSLFEFADKYRGSYQASCPFYCSHSGYQDELLWAAAWLYKATGEKS 248
Query: 226 YADF-IAHEAISSSVAEFSWDLKFPGAQVLLAEFNMTSAGGAQNFKSQADNFVCAVLPDT 402
Y ++ I+++ S ++ EFSWD KF G Q LLA A + FK+ ++FVCA++P +
Sbjct: 249 YLNYVISNKDWSKAINEFSWDNKFAGVQALLASEFYNGANDLEKFKTDVESFVCALMPGS 308
Query: 403 AFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTG-QTVLCGNQPLPPARLHE 579
+ Q+ TPGG++ +RD +N QYVT+ +L Y+ L + G ++ CG+ +++
Sbjct: 309 SSQQIKPTPGGILFIRDSSNLQYVTTATTILFYYSKTLTKAGVGSIQCGSTQFTVSQIRN 368
Query: 580 FARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVL---PPGTDVNCGLSFGDWM 750
FA+ Q+DY+LG NP SY+VGFG PTQPHHRG+S P + P D N G S+ ++
Sbjct: 369 FAKSQVDYILGNNPLKMSYMVGFGTKYPTQPHHRGSSLPSIQSKPEKIDCNGGFSYYNF- 427
Query: 751 APDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAAL 903
D PNPN TGAIVGGP+ +D + DKR + S+ EP TYIN+ IG +AAL
Sbjct: 428 --DTPNPNVHTGAIVGGPNSSDQYSDKRTDYSHAEPTTYINAAFIGSVAAL 476
>sptr|Q42660|Q42660 Beta-1,4-endoglycanohydrolase precursor (EC
3.2.1.4).
Length = 506
Score = 237 bits (604), Expect = 3e-61
Identities = 121/293 (41%), Positives = 175/293 (59%), Gaps = 4/293 (1%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGECPFYCSYSGYNDEXXXXXXXXXXXX 213
+VF++ D YS KLL +++ LF FA + GSY CPFYCSYSGY DE
Sbjct: 185 IVFKNIDSNYSAKLLRRSQSLFAFADKYRGSYQASCPFYCSYSGYQDELLWAAAWLYKAG 244
Query: 214 RRQVYADF-IAHEAISSSVAEFSWDLKFPGAQVLLAEFNMTSAGGAQNFKSQADNFVCAV 390
Y ++ + ++ S +EFSWD KF GAQ+LLA+ + + FK AD+FVCA+
Sbjct: 245 GGNNYLNYALNNQGWSQCPSEFSWDNKFAGAQILLAKEFLNGKSNLEKFKKDADSFVCAL 304
Query: 391 LPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTG-QTVLCGNQPLPPA 567
+P ++ Q+ TPGG++ RD +N QYV+ +L +Y+ +L G + + CG+ +
Sbjct: 305 MPGSSSVQIKTTPGGLLFFRDSSNLQYVSGATMVLFMYSKVLDAAGKEGITCGSVNFSTS 364
Query: 568 RLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP-VLPPGTDVNCGLSFGD 744
++ FA+ Q+DY+LG NP SY+VGFG PTQ HHR +S P + T V C +
Sbjct: 365 KIKAFAKSQVDYILGNNPLQMSYMVGFGNKYPTQLHHRASSLPSIYNHPTRVGCNDGYSS 424
Query: 745 WMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAAL 903
W + + PNPN GAIVGGP+ D FVD R + S++EP TY+N+ +G +AAL
Sbjct: 425 WYSINNPNPNTHVGAIVGGPNSGDQFVDSRSDYSHSEPTTYMNAAFVGSVAAL 477
>sptr|Q43149|Q43149 Cellulase (EC 3.2.1.4) (Fragment).
Length = 508
Score = 236 bits (603), Expect = 4e-61
Identities = 123/293 (41%), Positives = 175/293 (59%), Gaps = 4/293 (1%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGECPFYCSYSGYNDEXXXXXXXXXXXX 213
+VF++ D YS KLL +KLLF FA + GSY CPFYCSYSGY DE
Sbjct: 193 IVFQEVDSNYSAKLLRHSKLLFQFADQYRGSYQASCPFYCSYSGYQDELLWAAAWLYKAS 252
Query: 214 RRQVYADFI-AHEAISSSVAEFSWDLKFPGAQVLLAEFNMTSAGGAQNFKSQADNFVCAV 390
Y +++ ++ S +V+EFSWD KF GAQ LLA+ FKS +++F+CA+
Sbjct: 253 GDATYLNYVLGNQGWSHAVSEFSWDNKFAGAQTLLAKEFFGGKTNLGKFKSDSESFICAL 312
Query: 391 LPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTG-QTVLCGNQPLPPA 567
+P ++ Q+ TPGG+++ RD +N QYVTS + + ++Y+ +L + V CG+ P +
Sbjct: 313 MPGSSSVQIQTTPGGLLYTRDSSNLQYVTSASMVFLIYSKILNAANIKGVQCGSVNFPTS 372
Query: 568 RLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVLP-PGTDVNCGLSFGD 744
++ FA+ Q+DY+LG NP SY+VGFG+ P Q HHRGAS P + V C +
Sbjct: 373 QIKAFAKSQVDYILGNNPMKMSYMVGFGSKYPLQLHHRGASIPSMQIHPARVGCNEGYSA 432
Query: 745 WMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAAL 903
W + K NPN GAIVGGP+ ND F D R + S+ E TY+N+ +G +AAL
Sbjct: 433 WYSSSKSNPNTHVGAIVGGPNSNDQFNDVRSDYSHLETTTYMNAAFVGSVAAL 485
>sptr|Q96545|Q96545 Cellulase (EC 3.2.1.4).
Length = 506
Score = 235 bits (599), Expect = 1e-60
Identities = 120/293 (40%), Positives = 174/293 (59%), Gaps = 4/293 (1%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGECPFYCSYSGYNDEXXXXXXXXXXXX 213
+VF++ D YS KLL +++ LF FA + GSY CPFYCSYSGY DE
Sbjct: 185 IVFKNIDSNYSAKLLRRSQSLFAFADKYRGSYQASCPFYCSYSGYQDELLWAAAWLYKAG 244
Query: 214 RRQVYADF-IAHEAISSSVAEFSWDLKFPGAQVLLAEFNMTSAGGAQNFKSQADNFVCAV 390
Y ++ + ++ S +EFSWD KF GAQ+LLA+ + + FK AD+FVCA+
Sbjct: 245 GGNNYLNYALNNQGWSQCPSEFSWDNKFAGAQILLAKEFLNGKSNLEKFKKDADSFVCAL 304
Query: 391 LPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTG-QTVLCGNQPLPPA 567
+P ++ Q+ TPGG++ RD +N QYV+ +L +Y+ +L G + + CG+ +
Sbjct: 305 MPGSSSVQIKTTPGGLLFFRDSSNLQYVSGATMVLFMYSKVLDAAGKEGITCGSVNFSTS 364
Query: 568 RLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP-VLPPGTDVNCGLSFGD 744
++ FA+ Q+DY+LG NP SY+VGFG PTQ HHR +S P + V C +
Sbjct: 365 KIKAFAKSQVDYILGNNPLQMSYMVGFGNKYPTQLHHRASSLPSIYNHPARVGCNDGYSS 424
Query: 745 WMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAAL 903
W + + PNPN GAIVGGP+ D FVD R + S++EP TY+N+ +G +AAL
Sbjct: 425 WYSINNPNPNTHVGAIVGGPNSGDQFVDSRSDYSHSEPTTYMNAAFVGSVAAL 477
>sptr|Q42871|Q42871 Endo-1,4-beta-glucanase precursor (EC 3.2.1.4).
Length = 501
Score = 232 bits (591), Expect = 1e-59
Identities = 119/293 (40%), Positives = 171/293 (58%), Gaps = 4/293 (1%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGECPFYCSYSGYNDEXXXXXXXXXXXX 213
+VF++ D YS KLL +++ LF FA + GSY CPFYCSYSGY DE
Sbjct: 186 IVFKNIDSNYSTKLLKRSRSLFAFADKYRGSYQASCPFYCSYSGYKDELLWAAAWLYKAG 245
Query: 214 RRQVYADFIA-HEAISSSVAEFSWDLKFPGAQVLLAEFNMTSAGGAQNFKSQADNFVCAV 390
Y ++ + ++ S +EFSWD KF GAQ LLA+ + + FK AD+F+C +
Sbjct: 246 GGNNYLNYASINQGWSQVASEFSWDDKFAGAQTLLAKEYLNGKSNLEKFKKDADSFICGL 305
Query: 391 LPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTG-QTVLCGNQPLPPA 567
+P+++ Q+ TPGG+++ RD +N QYV +L +Y +L G V CG+ +
Sbjct: 306 MPESSSIQIKTTPGGLLYYRDSSNLQYVNGATMVLFMYTKVLEAAGIGGVTCGSVNFSTS 365
Query: 568 RLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP-VLPPGTDVNCGLSFGD 744
++ FA+ Q+DY+LG NP SY+VGFG PT+ HHR +S P + T V C +
Sbjct: 366 KIKAFAKLQVDYILGNNPLKMSYMVGFGNKYPTKLHHRASSLPSIYNHPTRVGCNDGYSS 425
Query: 745 WMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAAL 903
W + PNPN GAIVGGP+ D F+D R + S++EP TY+N+ IG +AAL
Sbjct: 426 WYNSNNPNPNTHVGAIVGGPNSGDQFIDSRSDYSHSEPTTYMNAAFIGSVAAL 478
>sw|P22503|GUN_PHAVU Endoglucanase precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (Abscission cellulase).
Length = 496
Score = 229 bits (585), Expect = 5e-59
Identities = 123/292 (42%), Positives = 168/292 (57%), Gaps = 5/292 (1%)
Frame = +1
Query: 52 DKKYSRKLLNKAKLLFTFAKSHLGSYDGECPFYCSYSGYNDEXXXXXXXXXXXXRRQVYA 231
D KYS LL+ +K LF FA + GSY G CPFYCSYSGY DE Y
Sbjct: 206 DAKYSSTLLSHSKSLFDFADKNRGSYSGSCPFYCSYSGYQDELLWAAAWLYKASGESKYL 265
Query: 232 DFI-AHEAISSSVAEFSWDLKFPGAQVLLAEFNMTSAGGAQNFKSQADNFVCAVLPDTAF 408
+I +++ S +V+EFSWD KF GAQ LL E K+ A++F+CAV+P +
Sbjct: 266 SYIISNQGWSQTVSEFSWDNKFVGAQTLLTEEFYGGKKDLAKIKTDAESFICAVMPGSNS 325
Query: 409 HQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTG-QTVLCGNQPLPPARLHEFA 585
Q+ TPGG++ RD +N QY TS+ +L +++ +L R + CG+ +++ FA
Sbjct: 326 RQIKTTPGGLLFTRDSSNLQYTTSSTMVLFIFSRILNRNHINGINCGSSHFTASQIRGFA 385
Query: 586 RQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP---VLPPGTDVNCGLSFGDWMAP 756
+ Q++Y+LG NP SY+VGFG+ P Q HHRG+S P V P N GLS D+
Sbjct: 386 KTQVEYILGKNPMKMSYMVGFGSKYPKQLHHRGSSIPSIKVHPAKVGCNAGLS--DYYNS 443
Query: 757 DKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAALAVR 912
PNPN GAIVGGPD ND F D R + S+ EP TYIN+ + ++AL +
Sbjct: 444 ANPNPNTHVGAIVGGPDSNDRFNDARSDYSHAEPTTYINAAFVASISALLAK 495
>sptr|Q43105|Q43105 Cellulase (EC 3.2.1.4).
Length = 496
Score = 229 bits (584), Expect = 6e-59
Identities = 123/292 (42%), Positives = 168/292 (57%), Gaps = 5/292 (1%)
Frame = +1
Query: 52 DKKYSRKLLNKAKLLFTFAKSHLGSYDGECPFYCSYSGYNDEXXXXXXXXXXXXRRQVYA 231
D KYS LL+ +K LF FA + GSY G CPFYCSYSGY DE Y
Sbjct: 206 DAKYSSTLLSHSKSLFDFADKNRGSYSGSCPFYCSYSGYQDELLWAAAWLYKASGESKYL 265
Query: 232 DFI-AHEAISSSVAEFSWDLKFPGAQVLLAEFNMTSAGGAQNFKSQADNFVCAVLPDTAF 408
+I +++ S +V+EFSWD KF GAQ LL E K+ A++F+CAV+P +
Sbjct: 266 SYIISNQGWSQTVSEFSWDNKFVGAQTLLTEEFYGGKKDLAKIKTDAESFICAVMPGSNS 325
Query: 409 HQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTG-QTVLCGNQPLPPARLHEFA 585
Q+ TPGG++ RD +N QY TS+ +L +++ +L R + CG+ +++ FA
Sbjct: 326 RQIKTTPGGLLFTRDSSNLQYTTSSTMVLFIFSRILNRNHINGINCGSSHFTASQIRGFA 385
Query: 586 RQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP---VLPPGTDVNCGLSFGDWMAP 756
+ Q++Y+LG NP SY+VGFG+ P Q HHRG+S P V P N GLS D+
Sbjct: 386 KTQVEYILGNNPMKMSYMVGFGSKYPKQLHHRGSSIPSIKVHPAKVGCNAGLS--DYYNS 443
Query: 757 DKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAALAVR 912
PNPN GAIVGGPD ND F D R + S+ EP TYIN+ + ++AL +
Sbjct: 444 ANPNPNTHVGAIVGGPDSNDRFNDARSDYSHAEPTTYINAAFVASISALLAK 495
>sptr|Q7XB15|Q7XB15 Endo-1,4-beta-glucanase (EC 3.2.1.4).
Length = 490
Score = 227 bits (579), Expect = 2e-58
Identities = 127/309 (41%), Positives = 168/309 (54%), Gaps = 7/309 (2%)
Frame = +1
Query: 1 GXXXXXXXXXXXMVFRDDKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSG 165
G + + D KY++ L AK FTFA+SH GSY CP+YCSYSG
Sbjct: 174 GETAAALAAASLVFHKIDHKYAKTLRETAKKAFTFAESHRGSYSDSLSSAVCPYYCSYSG 233
Query: 166 YNDEXXXXXXXXXXXXRRQVYADFIAHEAISSSVAEFSWDLKFPGAQVLLAE-FNMTSAG 342
+ DE Y +F + S+ FSWD KFPGA +LLA+ F +
Sbjct: 234 FQDELLWAAGWLYKATHNASYLNFAQSLNVDSNADTFSWDNKFPGADILLAQDFIINKNQ 293
Query: 343 GAQNFKSQADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLR 522
A ++K QA+ F+C VLP++ T GG+++ +N QYVTST+FLL VYA L
Sbjct: 294 AASSYKQQAEKFLCTVLPNSPTLSAKYTAGGLLYKMTPSNLQYVTSTSFLLGVYAKYLRI 353
Query: 523 TGQTVLCGNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP-V 699
+ +T CGN + P+ L +Q+ Y+LG NP+ SY+V FG N P HHRG+S P V
Sbjct: 354 SRRTFSCGNMVVTPSVLIRLVEKQIGYILGHNPQALSYMVDFGNNFPLHIHHRGSSIPSV 413
Query: 700 LPPGTDVNCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSL 879
+ C F + PNPN LTGA+VGGPD+ND F D R N S +EP TYIN+
Sbjct: 414 HADAQAIGCDEGF-QYYNTSNPNPNILTGAVVGGPDENDSFADDRDNYSQSEPATYINAP 472
Query: 880 AIGPLAALA 906
+G LA LA
Sbjct: 473 LVGTLAFLA 481
>sptr|O64402|O64402 Endo-beta-1,4-glucanase.
Length = 515
Score = 227 bits (578), Expect = 3e-58
Identities = 130/302 (43%), Positives = 170/302 (56%), Gaps = 12/302 (3%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
MVFR D YSR LL A +F FA H G+Y CPFYCSYSGYNDE
Sbjct: 211 MVFRTTDPIYSRHLLQTAMQVFDFADKHRGAYSDSLQSEVCPFYCSYSGYNDELLWGAAW 270
Query: 199 XXXXXRRQVYADFIAHEAIS----SSVAEFSWDLKFPGAQVLLA-EFNMTSAGGAQNFKS 363
R Y +I + +V FSWD K GA+VLLA E + ++ + ++
Sbjct: 271 LQRASRNVSYISYIQSHGQNLGGEDNVNIFSWDDKHAGARVLLAKEVLLRNSKSLEEYRG 330
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
ADNFVC++LP T Q TPGG+++ N QYVTST FLL Y L + Q V C
Sbjct: 331 HADNFVCSLLPGTPNSQAQYTPGGLLYKMSDCNLQYVTSTTFLLFTYPKYLRVSKQVVRC 390
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP-VLPPGTDV 720
GN + P R+ A++Q+DY+LG NP SY+VG+GA P + HHRG+S P + +
Sbjct: 391 GNMVVTPTRIRTLAKRQVDYILGDNPLRMSYMVGYGAKFPERIHHRGSSLPSIYQHPQII 450
Query: 721 NCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAA 900
C F + + PNPN L GA+VGGPD ND F D+R + +++EP TYIN+ +G LA
Sbjct: 451 PCNDGFQS-LYSNAPNPNRLVGAVVGGPDNNDHFSDERNDYAHSEPTTYINAPLVGSLAY 509
Query: 901 LA 906
LA
Sbjct: 510 LA 511
>sptrnew|BAD05437|BAD05437 Putative cellulase.
Length = 516
Score = 226 bits (577), Expect = 4e-58
Identities = 118/286 (41%), Positives = 163/286 (56%), Gaps = 3/286 (1%)
Frame = +1
Query: 52 DKKYSRKLLNKAKLLFTFAKSHLGSYDGECPFYCSYSGYNDEXXXXXXXXXXXXRRQVYA 231
D +S +LL ++ LF FA ++ GS+ CPFYCSYSG+ DE R Y
Sbjct: 207 DTAFSSRLLAASRSLFDFANNYRGSFQSSCPFYCSYSGFQDELLWASAWLFKATRDAKYL 266
Query: 232 DFIAHEAISSS-VAEFSWDLKFPGAQVLLAEFNMTSAGGAQNFKSQADNFVCAVLPDTAF 408
DF+ + SS+ V EFSWD K+ GAQ+L A+ + +K D+FVCA++P++
Sbjct: 267 DFLTNNQGSSNPVNEFSWDNKYAGAQMLAAQEYLGGRTQLARYKDNLDSFVCALMPNSGN 326
Query: 409 HQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTG-QTVLCGNQPLPPARLHEFA 585
Q+ TPGG++ RD N QY T+ +L +Y+ +L +G + V C P ++ FA
Sbjct: 327 VQIRTTPGGLLFTRDSVNLQYTTTATLVLSIYSKVLKSSGSRGVRCSAATFSPNQISSFA 386
Query: 586 RQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVLPP-GTDVNCGLSFGDWMAPDK 762
Q+DY+LG NP SY+VGF P + HHRG+S P + V C F W+
Sbjct: 387 TSQVDYILGKNPLGMSYMVGFSTKFPRRIHHRGSSIPSIKVLSRKVTCKEGFSSWLPTSD 446
Query: 763 PNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAA 900
PNPN GAIVGGPD ND F D RG+SS++EP TYIN+ +G AA
Sbjct: 447 PNPNIHVGAIVGGPDGNDQFSDNRGDSSHSEPATYINAAFVGACAA 492
>sptrnew|AAQ55294|AAQ55294 Endo-1,4-beta-glucanase.
Length = 497
Score = 224 bits (570), Expect = 3e-57
Identities = 125/298 (41%), Positives = 165/298 (55%), Gaps = 8/298 (2%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VFR D KYS+ LLN A+ + FA + G+Y CPFYCSYSGYNDE
Sbjct: 195 LVFRRVDPKYSKLLLNTARNVMQFAIQYRGAYSDSLGSAVCPFYCSYSGYNDELLWGAAW 254
Query: 199 XXXXXRRQVYADFIAHEAISSSVAEFSWDLKFPGAQVLLAEFNMTSAG-GAQNFKSQADN 375
Y +FI S S FSWD KF GA VLL+ + + + +K +A+
Sbjct: 255 LFRATNELYYYNFIKSLGASDSTDIFSWDNKFAGAYVLLSRRALLNNDKNFEPYKQEAEQ 314
Query: 376 FVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLCGNQP 555
F+C +LP++ T GG+I+ G+N QYVTS FLL Y+ + T CGN
Sbjct: 315 FMCRILPNSPSSSTQYTQGGLIYKLPGSNLQYVTSITFLLTTYSKYMAARKLTFDCGNLV 374
Query: 556 LPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVLPP-GTDVNCGL 732
+ P L A+QQ+DY+LG NP SY+VG+G P + HHRG+S P L + C
Sbjct: 375 VTPMALRNLAKQQVDYILGVNPLKMSYMVGYGPYYPKRIHHRGSSLPSLTSHRQSIGCDG 434
Query: 733 SFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAALA 906
F + PNPN L GA+VGGP++NDGF D RG+ S++EP TYIN +GPLA A
Sbjct: 435 GFQPFFYSLNPNPNILVGAVVGGPNQNDGFPDDRGDYSHSEPATYINGAIVGPLAFFA 492
>sptr|O64401|O64401 Endo-beta-1,4-glucanase.
Length = 510
Score = 221 bits (562), Expect = 2e-56
Identities = 122/304 (40%), Positives = 170/304 (55%), Gaps = 14/304 (4%)
Frame = +1
Query: 37 MVFR-DDKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VFR D +YS++L++ A +F FA H G+Y CPFYCSYSGY DE
Sbjct: 205 LVFRKSDPQYSKRLIHTAMRVFDFADKHRGAYSDSLRSCVCPFYCSYSGYEDELLWGAAW 264
Query: 199 XXXXXRRQVYADFIAHEAISSSVAE----FSWDLKFPGAQVLLA-EFNMTSAGGAQNFKS 363
R Y ++I + S E F WD K GA+VLL+ EF + + ++K
Sbjct: 265 LHKATRNPTYLNYIQSKGQSLGADESDNIFGWDNKHAGARVLLSKEFLLRNVKSLHDYKG 324
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
ADN++C++LP ++ Q TPGG+++ +N QYVTS++FLL YA L + V C
Sbjct: 325 HADNYICSLLPGVSYSQAKYTPGGLLYTLSDSNLQYVTSSSFLLFTYAKYLSSSKHVVTC 384
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVL---PPGT 714
G+ P RL A++Q+DY+LG NP SY+VG+G++ P + HHRG+S P P
Sbjct: 385 GSMTFSPKRLRTIAKRQVDYILGDNPLKMSYMVGYGSHYPERIHHRGSSLPSKGNHPQAI 444
Query: 715 DVNCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPL 894
N G + + PNPN L GA+VGGPD D F D R + +EP TYIN+ +G L
Sbjct: 445 PCNAGFQS---LYSNAPNPNILVGAVVGGPDSMDRFPDDRNDYEQSEPTTYINAPFVGSL 501
Query: 895 AALA 906
A LA
Sbjct: 502 AYLA 505
>sptr|Q39847|Q39847 CMCase, cellulase, endo-1,4-beta-D-glucanase
(Fragment).
Length = 299
Score = 220 bits (561), Expect = 3e-56
Identities = 114/279 (40%), Positives = 162/279 (58%), Gaps = 3/279 (1%)
Frame = +1
Query: 76 LNKAKLLFTFAKSHLGSYDGECPFYCSYSGYNDEXXXXXXXXXXXXRRQVYADF-IAHEA 252
L+K+K LF FA + GSY G CPFYCSYSGY DE Y + I ++
Sbjct: 1 LSKSKSLFDFADKYRGSYSGSCPFYCSYSGYQDELLWAASWLYKASGESKYLSYSIGNQG 60
Query: 253 ISSSVAEFSWDLKFPGAQVLLAEFNMTSAGGAQNFKSQADNFVCAVLPDTAFHQVFITPG 432
S +V+EFSWD K+ GAQ LL E FKS ++F+C+V+P ++ Q+ TPG
Sbjct: 61 WSQAVSEFSWDNKYVGAQTLLTEEFYGGKKDLAKFKSDVESFICSVMPASSSLQIKTTPG 120
Query: 433 GVIHLRDGANSQYVTSTAFLLVVYADLLLRTG-QTVLCGNQPLPPARLHEFARQQMDYLL 609
G++ RD +N QY TS+ +L +++ +L R + CG+ P+++ FA+ Q+DY+L
Sbjct: 121 GLLFTRDSSNLQYATSSTMVLFIFSKILNRNHIDRIHCGSALFTPSQIRAFAKTQVDYIL 180
Query: 610 GANPRHSSYVVGFGANPPTQPHHRGASTP-VLPPGTDVNCGLSFGDWMAPDKPNPNELTG 786
G+NP SY+VGFG+ P Q HHRG+S P + T V C + PNPN G
Sbjct: 181 GSNPMKMSYMVGFGSKYPKQLHHRGSSIPSINVHPTKVGCNDGLSVYYNSANPNPNTHVG 240
Query: 787 AIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAAL 903
AIVGGPD ND F D R + S++ P TY+N+ + ++AL
Sbjct: 241 AIVGGPDSNDRFSDARSDYSHSGPTTYMNAAFVASVSAL 279
>sptr|O22298|O22298 Basic cellulase.
Length = 488
Score = 219 bits (557), Expect = 9e-56
Identities = 121/298 (40%), Positives = 163/298 (54%), Gaps = 8/298 (2%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VFR D +Y+ LL AK + FA + G+Y CPFYCSYSGY DE
Sbjct: 186 LVFRKGDPRYASLLLRTAKNVLQFAMQYRGAYSDSLGSAVCPFYCSYSGYKDELLWGAAW 245
Query: 199 XXXXXRRQVYADFIAHEAISSSVAEFSWDLKFPGAQVLLAEFNMTSAG-GAQNFKSQADN 375
Y +FI FSWD KF GA VLLA + + + ++ +A++
Sbjct: 246 LFRATNDVTYYNFIKSVGDDGGTDVFSWDNKFAGAHVLLARGALLNRDKNFEPYRQEAED 305
Query: 376 FVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLCGNQP 555
F+C +LP++ F T GG+++ +N QYVTS +FLL YA + T CGN
Sbjct: 306 FICRILPNSPFTTTQYTQGGLMYKMPESNLQYVTSISFLLTTYAKYMRATKHYFTCGNMV 365
Query: 556 LPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVLP-PGTDVNCGL 732
+ P L A++Q+DY+LG NP SY+VGFG N P + HHRG+S P L + C
Sbjct: 366 VNPGLLTNLAKRQVDYILGVNPIKMSYMVGFGPNFPRRIHHRGSSLPSLANHPQSIRCDG 425
Query: 733 SFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAALA 906
F + PNPN L GAIVGGP++NDGF D R + S++EP TYIN+ +GPLA A
Sbjct: 426 GFEPFFHSSNPNPNILVGAIVGGPNQNDGFPDDRSDYSHSEPATYINAAMVGPLAYFA 483
>sptr|Q9SRX3|Q9SRX3 Endo-1,4-beta glucanase.
Length = 501
Score = 217 bits (553), Expect = 2e-55
Identities = 126/302 (41%), Positives = 164/302 (54%), Gaps = 12/302 (3%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VFR D YS LL +A +FTFA + G Y CPFYCSYSGY DE
Sbjct: 201 IVFRKCDPSYSNHLLQRAITVFTFADKYRGPYSAGLAPEVCPFYCSYSGYQDELLWGAAW 260
Query: 199 XXXXXRRQVYADFIAHEAISSSVAEF----SWDLKFPGAQVLLA-EFNMTSAGGAQNFKS 363
Y ++I EF SWD K GA++LL+ EF + + +K
Sbjct: 261 LQKATNNPTYLNYIKANGQILGADEFDNMFSWDNKHVGARILLSKEFLIQKVKSLEEYKE 320
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
AD+F+C+VLP + Q TPGG++ +N QYVTST+FLL+ YA L C
Sbjct: 321 HADSFICSVLPGASSSQY--TPGGLLFKMGESNMQYVTSTSFLLLTYAKYLTSARTVAYC 378
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP-VLPPGTDV 720
G + PARL A++Q+DYLLG NP SY+VG+G P + HHRG+S P V T +
Sbjct: 379 GGSVVTPARLRSIAKKQVDYLLGGNPLKMSYMVGYGLKYPRRIHHRGSSLPSVAVHPTRI 438
Query: 721 NCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAA 900
C F PNPN+L GA+VGGPD+ND F D+R + +EP TYIN+ +G LA
Sbjct: 439 QCHDGF-SLFTSQSPNPNDLVGAVVGGPDQNDQFPDERSDYGRSEPATYINAPLVGALAY 497
Query: 901 LA 906
LA
Sbjct: 498 LA 499
>sptr|O64949|O64949 Endo-1,4-beta glucanase.
Length = 501
Score = 216 bits (551), Expect = 4e-55
Identities = 125/302 (41%), Positives = 164/302 (54%), Gaps = 12/302 (3%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VFR D YS LL +A +FTFA + G Y CPFYCSYSGY DE
Sbjct: 201 IVFRKCDPSYSNHLLQRAITVFTFADKYRGPYSAGLAPEVCPFYCSYSGYQDELLWGAAW 260
Query: 199 XXXXXRRQVYADFIAHEAISSSVAEF----SWDLKFPGAQVLLA-EFNMTSAGGAQNFKS 363
Y ++I EF SWD K GA++LL+ EF + + +K
Sbjct: 261 LQKATNNPTYLNYIKANGQILGADEFDNMFSWDNKHVGARILLSKEFLIQKVKSLEEYKE 320
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
AD+F+C+V+P + Q TPGG++ +N QYVTST+FLL+ YA L C
Sbjct: 321 HADSFICSVIPGASSSQY--TPGGLLFKMGESNMQYVTSTSFLLLTYAKYLTSARTVAYC 378
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP-VLPPGTDV 720
G + PARL A++Q+DYLLG NP SY+VG+G P + HHRG+S P V T +
Sbjct: 379 GGSVVTPARLRSIAKKQVDYLLGGNPLKMSYMVGYGLKYPRRIHHRGSSLPSVAVHPTRI 438
Query: 721 NCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAA 900
C F PNPN+L GA+VGGPD+ND F D+R + +EP TYIN+ +G LA
Sbjct: 439 QCHDGF-SLFTSQSPNPNDLVGAVVGGPDQNDQFPDERSDYGRSEPATYINAPLVGALAY 497
Query: 901 LA 906
LA
Sbjct: 498 LA 499
>sptrnew|BAD10555|BAD10555 Putative endoglucanase 1
(Endo-1,4-beta-glucanase) (Abscission cellulase 1).
Length = 523
Score = 216 bits (551), Expect = 4e-55
Identities = 124/309 (40%), Positives = 172/309 (55%), Gaps = 19/309 (6%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
MVFR D YS +LL+ A +F FA H GSY CPFYCSYSGY+DE
Sbjct: 213 MVFRRADPAYSARLLHAATQVFDFADRHRGSYSDSLASSVCPFYCSYSGYHDELLWGASW 272
Query: 199 XXXXXRRQVYADFIAHEAISSSVAE----FSWDLKFPGAQVLLAE-FNMTSAGGAQNFKS 363
R + ++ + + FSWD K G +VLLA+ F G + +K+
Sbjct: 273 LHRASRNASFMSYVEANGMQLGAGDDDYSFSWDDKRVGTKVLLAKGFLRNRLHGLELYKA 332
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
+D+++C+++P TA Q TPGG+++ +N QYVT+ FL++ YA L +G T C
Sbjct: 333 HSDSYICSLVPGTASFQSRYTPGGLLYREGSSNMQYVTTATFLMLAYAKYLRSSGATASC 392
Query: 544 GNQ------PLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVLP 705
G+ + A L A++Q+DY+LG NP SY+VGFG P + HHRGAS P +
Sbjct: 393 GDGGGGARGEVSAAELVAVAKRQVDYILGKNPAGMSYMVGFGCRYPRRAHHRGASMPSVR 452
Query: 706 --PGTDVNCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSL 879
PG ++C FG ++ +PNPN L GA+VGGPD D F D RGN + +EP TYIN+
Sbjct: 453 AHPGR-ISCDAGFG-YLHSGEPNPNVLVGAVVGGPDSRDAFADDRGNFAQSEPATYINAP 510
Query: 880 AIGPLAALA 906
+G LA A
Sbjct: 511 LVGALAYFA 519
>sptr|O82473|O82473 Endo-1,4-beta-glucanase (EC 3.2.1.4).
Length = 497
Score = 216 bits (549), Expect = 7e-55
Identities = 124/303 (40%), Positives = 169/303 (55%), Gaps = 13/303 (4%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VF+D D YS LL A+ +F FA + GSY CPF CSYSGYNDE
Sbjct: 194 IVFKDSDPSYSSSLLRTAQKVFAFADKYRGSYSDSLSSVVCPFSCSYSGYNDELLWGASW 253
Query: 199 XXXXXRRQVYADFIAHEAISSSVAE----FSWDLKFPGAQVLLA-EFNMTSAGGAQNFKS 363
+ Y +I + + FSWD K PG +++L+ +F S+ Q +K
Sbjct: 254 LHRASQDSSYLAYIQSNGQTMGANDDDYSFSWDDKRPGTKIILSKDFLEKSSQEFQAYKV 313
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
+DN++C+++P + Q TPGG+++ G+N QYVTS++FLL+ YA L G V C
Sbjct: 314 HSDNYICSLIPGSPSFQAQYTPGGLLYKGSGSNLQYVTSSSFLLLTYAKYLRSNGGDVSC 373
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVL--PPGTD 717
G+ P RL E A++Q+DY+LG NP SY+VGFG P + HHRG+S P + PG
Sbjct: 374 GSSRFPAERLVELAKKQVDYILGDNPAKISYMVGFGDKYPLRVHHRGSSLPSVHAHPG-H 432
Query: 718 VNCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLA 897
+ C F + PNPN L GAIVGGPD D F D R N +EP TYIN+ +G LA
Sbjct: 433 IGCNDGFQS-LNSGSPNPNVLIGAIVGGPDSRDNFEDDRNNYQQSEPATYINAPLVGALA 491
Query: 898 ALA 906
L+
Sbjct: 492 FLS 494
>sptr|Q9SML6|Q9SML6 Endo-beta-1,4-glucanase precursor (EC 3.2.1.4).
Length = 497
Score = 215 bits (548), Expect = 9e-55
Identities = 124/303 (40%), Positives = 168/303 (55%), Gaps = 13/303 (4%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VF+D D YS LL+ A+ +F FA + GSY CPFYCSYSGYNDE
Sbjct: 191 IVFKDSDPSYSSTLLHTAQKVFAFADKYRGSYSDSLSSVVCPFYCSYSGYNDELLWGASW 250
Query: 199 XXXXXRRQVYADFIAHEAISSSVAE----FSWDLKFPGAQVLLA-EFNMTSAGGAQNFKS 363
+ Y +I + + FSWD K PG +++L+ +F S Q +K
Sbjct: 251 LHRASQDTSYLSYIQSNGQTMGANDDDYSFSWDDKRPGTKIVLSKDFLEKSTQEFQAYKV 310
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
+DN++C+++P + Q TPGG++ +N QYVTS++FLL+ YA L G V C
Sbjct: 311 HSDNYICSLIPGSPSFQAQYTPGGLLFKGSESNLQYVTSSSFLLLTYAKYLRSNGGVVSC 370
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVL--PPGTD 717
G+ P +L E AR+Q+DY+LG NP SY+VGFG P + HHRG+S P + PG
Sbjct: 371 GSSRFPANKLVELARKQVDYILGDNPAKISYMVGFGQKYPLRVHHRGSSLPSVRTHPG-H 429
Query: 718 VNCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLA 897
+ C F + PNPN L GAIVGGPD D F D R N +EP TYIN+ +G LA
Sbjct: 430 IGCNDGFQS-LYSGSPNPNVLVGAIVGGPDSRDNFEDDRNNYQQSEPATYINAPLVGALA 488
Query: 898 ALA 906
L+
Sbjct: 489 FLS 491
>sptr|Q96546|Q96546 Endo-beta-1,4-glucanase (EC 3.2.1.4).
Length = 497
Score = 215 bits (547), Expect = 1e-54
Identities = 124/303 (40%), Positives = 167/303 (55%), Gaps = 13/303 (4%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VF+D D YS LL A+ +F FA + GSY CPFYCSYSGYNDE
Sbjct: 191 IVFKDSDPSYSSTLLRTAQKVFAFADKYRGSYSDSLSSVVCPFYCSYSGYNDELLWGASW 250
Query: 199 XXXXXRRQVYADFIAHEAISSSVAE----FSWDLKFPGAQVLLA-EFNMTSAGGAQNFKS 363
+ Y +I + + FSWD K PG +++L+ +F S Q +K
Sbjct: 251 LHRASQDTSYLSYIQSNGQTMGANDDDYSFSWDDKRPGTKIVLSKDFLEKSTQEFQAYKV 310
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
+DN++C+++P + Q TPGG++ +N QYVTS++FLL+ YA L G V C
Sbjct: 311 HSDNYICSLIPGSPSFQAQYTPGGLLFKGSESNLQYVTSSSFLLLTYAKYLRSNGGVVSC 370
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVL--PPGTD 717
G+ P +L E AR+Q+DY+LG NP SY+VGFG P + HHRG+S P + PG
Sbjct: 371 GSSRFPANKLVELARKQVDYILGDNPAKISYMVGFGQKYPLRVHHRGSSLPSVRTHPG-H 429
Query: 718 VNCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLA 897
+ C F + PNPN L GAIVGGPD D F D R N +EP TYIN+ +G LA
Sbjct: 430 IGCNDGFQS-LYSGSPNPNVLVGAIVGGPDSRDNFEDDRNNYQQSEPATYINAPLVGALA 488
Query: 898 ALA 906
L+
Sbjct: 489 FLS 491
>sptr|Q42875|Q42875 Endo-1,4-beta-glucanase precursor (EC 3.2.1.4).
Length = 510
Score = 215 bits (547), Expect = 1e-54
Identities = 122/302 (40%), Positives = 167/302 (55%), Gaps = 12/302 (3%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VFR + YS+ L+ +A +F FA + GSY CP+YCS SGY DE
Sbjct: 204 LVFRKCNPSYSKILIKRAIRVFAFADKYRGSYSNGLRKVVCPYYCSVSGYEDELLWGAAW 263
Query: 199 XXXXXRRQVYADFIAHEAISSSVAE----FSWDLKFPGAQVLLAE-FNMTSAGGAQNFKS 363
+ Y ++I + AE F WD K GA++LL++ F + ++KS
Sbjct: 264 LHRATKNPTYLNYIQRNGQTLGAAETDNTFGWDNKHVGARILLSKSFLVQKLQTLHDYKS 323
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
ADN++C+++P T Q TPGG++ D +N QYVTST+FLLV YA L V C
Sbjct: 324 HADNYICSLIPGTPASQAQYTPGGLLFKMDDSNMQYVTSTSFLLVTYAKYLTSARMVVKC 383
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP-VLPPGTDV 720
G + P RL A++Q+DYLLG NP SY+VG+GA P + HHRG+S P V +
Sbjct: 384 GGVVITPKRLRNVAKKQVDYLLGDNPLKMSYMVGYGARYPQRIHHRGSSLPSVANHPAKI 443
Query: 721 NCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAA 900
C F M PNPN L GA+VGGPD++D F D+R + +EP TYIN+ +G L
Sbjct: 444 QCRDGF-SVMNSQSPNPNVLVGAVVGGPDEHDRFPDERSDYEQSEPATYINAPLVGTLTY 502
Query: 901 LA 906
LA
Sbjct: 503 LA 504
>sptr|O81416|O81416 T2H3.5 protein (Putative endo-1, 4-beta glucanase)
(AT4g02290/T2H3_5).
Length = 516
Score = 214 bits (545), Expect = 2e-54
Identities = 124/302 (41%), Positives = 161/302 (53%), Gaps = 12/302 (3%)
Frame = +1
Query: 37 MVFR-DDKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VFR D YS+ LL +A +F FA + G+Y CPFYCSYSGY DE
Sbjct: 210 IVFRKSDPSYSKVLLKRAISVFAFADKYRGTYSAGLKPDVCPFYCSYSGYQDELLWGAAW 269
Query: 199 XXXXXRRQVYADFIAHEAISSSVAE----FSWDLKFPGAQVLLAE-FNMTSAGGAQNFKS 363
+ Y ++I AE F WD K GA++LL + F + + +K
Sbjct: 270 LQKATKNIKYLNYIKINGQILGAAEYDNTFGWDNKHAGARILLTKAFLVQNVKTLHEYKG 329
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
ADNF+C+V+P F TPGG++ AN QYVTST+FLL+ YA L V C
Sbjct: 330 HADNFICSVIPGAPFSSTQYTPGGLLFKMADANMQYVTSTSFLLLTYAKYLTSAKTVVHC 389
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP-VLPPGTDV 720
G P RL A++Q+DYLLG NP SY+VG+G P + HHRG+S P V +
Sbjct: 390 GGSVYTPGRLRSIAKRQVDYLLGDNPLRMSYMVGYGPKFPRRIHHRGSSLPCVASHPAKI 449
Query: 721 NCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAA 900
C F M PNPN L GA+VGGPD++D F D+R + +EP TYINS +G LA
Sbjct: 450 QCHQGFA-IMNSQSPNPNFLVGAVVGGPDQHDRFPDERSDYEQSEPATYINSPLVGALAY 508
Query: 901 LA 906
A
Sbjct: 509 FA 510
>sptr|Q9SSU7|Q9SSU7 Endo-1,4-beta-glucanase.
Length = 506
Score = 214 bits (545), Expect = 2e-54
Identities = 122/303 (40%), Positives = 166/303 (54%), Gaps = 13/303 (4%)
Frame = +1
Query: 37 MVFR-DDKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VFR D Y++ L+ +A +F FA H SY CPFYCSYSGY D
Sbjct: 200 LVFRKSDPTYAKILVRRAIRVFQFADKHRRSYSNALKPFVCPFYCSYSGYQDGLLWGAAW 259
Query: 199 XXXXXRRQVYADFIAHEAISSSVAEFS----WDLKFPGAQVLLA-EFNMTSAGGAQNFKS 363
+ +Y +I AEF WD K GA++LL+ EF + + ++K
Sbjct: 260 LHKATKNPMYLKYIQTNGQILGAAEFDNTFGWDNKHVGARILLSKEFLVQNVKSLHDYKG 319
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
+DNFVC+++P TPGG++ +N QYVTST FLLV YA L ++ V C
Sbjct: 320 HSDNFVCSLIPGAGSSSAQYTPGGLLFKMSDSNMQYVTSTTFLLVTYAKYLTKSHSVVNC 379
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP--VLPPGTD 717
G + P RL A++Q+DYLLG NP SY+VG+G P + HHRG+S P + PG
Sbjct: 380 GGTTVTPKRLRTLAKRQVDYLLGDNPLKMSYMVGYGPRYPQRIHHRGSSLPSMAVHPG-K 438
Query: 718 VNCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLA 897
+ C FG M PNPN L GA+VGGPD++D F D+R + +EP TY+N+ +G LA
Sbjct: 439 IQCSAGFG-VMNSKSPNPNILMGAVVGGPDQHDRFPDQRSDYEQSEPATYVNAPLVGTLA 497
Query: 898 ALA 906
LA
Sbjct: 498 YLA 500
>sptr|O81403|O81403 Endo-1,4-beta-glucanase precursor (EC 3.2.1.4).
Length = 496
Score = 213 bits (542), Expect = 5e-54
Identities = 120/302 (39%), Positives = 169/302 (55%), Gaps = 12/302 (3%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VFR D YSR LLN+A +F FA +H G Y CPFYC +G+ DE
Sbjct: 193 IVFRSHDPAYSRLLLNRAVRVFEFADTHRGGYSSSLKNAVCPFYCDVNGFQDELLWGAAW 252
Query: 199 XXXXXRRQVYADFIAHEAI----SSSVAEFSWDLKFPGAQVLLA-EFNMTSAGGAQNFKS 363
RR+ Y ++I + ++ EF WD K G +L++ E M A ++FK
Sbjct: 253 LHKASRRRQYREYIVRNEVVLRAGDTINEFGWDNKHAGINILISKEVLMGKADYFESFKQ 312
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
AD F+C+VLP A QV +PGG+I G+N Q+VTS +FLL+ Y++ L + V C
Sbjct: 313 NADGFICSVLPGLAHTQVQYSPGGLIFKPGGSNMQHVTSLSFLLLTYSNYLSHANKNVPC 372
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP-VLPPGTDV 720
G PA L + A++Q+DY+LG NP SY+VG+G P + HHRG+S P V +
Sbjct: 373 GMTSASPAFLKQLAKRQVDYILGDNPLRMSYMVGYGPRYPQRIHHRGSSLPSVQAHPARI 432
Query: 721 NCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAA 900
C +M+P+ PNPN+L GA+VGGP+ +D F D R +EP TYIN+ +G L+
Sbjct: 433 GCKAGSRYFMSPN-PNPNKLVGAVVGGPNSSDAFPDSRPYFQESEPTTYINAPLVGLLSY 491
Query: 901 LA 906
A
Sbjct: 492 FA 493
>sptr|Q40763|Q40763 Cellulase precursor.
Length = 494
Score = 212 bits (540), Expect = 8e-54
Identities = 121/306 (39%), Positives = 164/306 (53%), Gaps = 14/306 (4%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VF++ D YS KLL+ A +F FA + GSY CPFYCSYSGY DE
Sbjct: 191 IVFKESDPSYSTKLLHTAMKVFDFADRYRGSYSNSLNSVVCPFYCSYSGYQDELLWGASW 250
Query: 199 XXXXXRRQVYADFIAHEAISSSVAE----FSWDLKFPGAQVLLA-EFNMTSAGGAQNFKS 363
+ Y +I + + FSWD K PG ++LL+ EF + Q +KS
Sbjct: 251 IHRASQNGSYLTYIQSNGHTMGSDDDDYSFSWDDKRPGTKILLSKEFLEKTTEEFQLYKS 310
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
+DN++C+++P T+ Q TPGG+ + +N QYVTST FLL+ YA L G C
Sbjct: 311 HSDNYICSLIPGTSSFQAQYTPGGLFYKASESNLQYVTSTTFLLLTYAKYLGSNGGVARC 370
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPVLPPGTDVN 723
G + L A++Q+DY+LG NP SY+VGFG P HHRG+S P + D
Sbjct: 371 GGSTVTAESLIAQAKKQVDYILGDNPARMSYMVGFGNRYPQHVHHRGSSVPSI---HDTR 427
Query: 724 CGLSFGD---WMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPL 894
G+S D ++ PNPN L GAI+GGPD D F D R N +EP TYIN+ +G L
Sbjct: 428 IGISCNDGFQFLYSSSPNPNVLVGAIIGGPDNRDNFADSRNNYQQSEPATYINAPFVGAL 487
Query: 895 AALAVR 912
A + +
Sbjct: 488 AFFSAK 493
>sptr|Q9ZTS0|Q9ZTS0 Putative cellulase.
Length = 496
Score = 212 bits (540), Expect = 8e-54
Identities = 119/302 (39%), Positives = 170/302 (56%), Gaps = 12/302 (3%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
+VFR D YSR LLN+A +F FA +H G+Y CPFYC +G+ DE
Sbjct: 193 IVFRSRDPAYSRLLLNRAVKVFEFADTHRGAYSSSLKNAVCPFYCDVNGFQDELLWGAAW 252
Query: 199 XXXXXRRQVYADFIAHEAI----SSSVAEFSWDLKFPGAQVLLA-EFNMTSAGGAQNFKS 363
RR+ Y ++I + ++ EF WD K G +L++ E M A ++FK
Sbjct: 253 LHKASRRRQYREYIVRNEVILRAGDTINEFGWDNKHAGINILISKEVLMGKADYFESFKQ 312
Query: 364 QADNFVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLC 543
AD F+C+VLP A QV +PGG+I G+N Q+VTS +FLL+ Y++ L + V C
Sbjct: 313 NADGFICSVLPGLAHTQVQYSPGGLIFKPGGSNMQHVTSLSFLLLTYSNYLSHANKNVPC 372
Query: 544 GNQPLPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTP-VLPPGTDV 720
G PA L + A++Q+DY+LG NP SY+VG+G P + HHRG+S P V +
Sbjct: 373 GMTSASPAFLKQLAKRQVDYILGDNPLRMSYMVGYGPRYPQRIHHRGSSLPSVQAHPARI 432
Query: 721 NCGLSFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAA 900
C +++P+ PNPN+L GA+VGGP+ +D F D R +EP TYIN+ +G L+
Sbjct: 433 GCKAGSHYFLSPN-PNPNKLVGAVVGGPNSSDAFPDSRPYFQESEPTTYINAPLVGLLSY 491
Query: 901 LA 906
A
Sbjct: 492 FA 493
>sptr|Q9C9H5|Q9C9H5 Putative beta-glucanase, 74324-76084.
Length = 484
Score = 212 bits (539), Expect = 1e-53
Identities = 122/298 (40%), Positives = 162/298 (54%), Gaps = 8/298 (2%)
Frame = +1
Query: 37 MVFRD-DKKYSRKLLNKAKLLFTFAKSHLGSYDGE-----CPFYCSYSGYNDEXXXXXXX 198
MVFR D KYSR LL AK + FA + G+Y CPFYCSYSGY DE
Sbjct: 183 MVFRKVDSKYSRLLLATAKDVMQFAIQYQGAYSDSLSSSVCPFYCSYSGYKDELMWGASW 242
Query: 199 XXXXXRRQVYADFIAHEAISSSVAEFSWDLKFPGAQVLLAEFNMTSA-GGAQNFKSQADN 375
YA+FI FSWD K+ GA VLL+ + + + +K A+N
Sbjct: 243 LLRATNNPYYANFIKSLGGGDQPDIFSWDNKYAGAYVLLSRRALLNKDSNFEQYKQAAEN 302
Query: 376 FVCAVLPDTAFHQVFITPGGVIHLRDGANSQYVTSTAFLLVVYADLLLRTGQTVLCGNQP 555
F+C +LPD+ T GG+++ +N QYVTS FLL YA + T T CG+
Sbjct: 303 FICKILPDSPSSSTQYTQGGLMYKLPQSNLQYVTSITFLLTTYAKYMKATKHTFNCGSSV 362
Query: 556 LPPARLHEFARQQMDYLLGANPRHSSYVVGFGANPPTQPHHRGASTPV-LPPGTDVNCGL 732
+ P L +++Q+DY+LG NP SY+VGF +N P + HHR +S P + C
Sbjct: 363 IVPNALISLSKRQVDYILGDNPIKMSYMVGFSSNFPKRIHHRASSLPSHALRSQSLGCNG 422
Query: 733 SFGDWMAPDKPNPNELTGAIVGGPDKNDGFVDKRGNSSYTEPCTYINSLAIGPLAALA 906
F + PNPN LTGAIVGGP++NDG+ D+R + S+ EP TYIN+ +GPLA A
Sbjct: 423 GFQSFYT-QNPNPNILTGAIVGGPNQNDGYPDQRDDYSHAEPATYINAAFVGPLAYFA 479
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 676,563,205
Number of Sequences: 1395590
Number of extensions: 15176030
Number of successful extensions: 51280
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 47105
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 50675
length of database: 442,889,342
effective HSP length: 125
effective length of database: 268,440,592
effective search space used: 72478959840
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

