BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= 3204472.2.1
(757 letters)
Database: /db/trembl-ebi/tmp/swall
1,313,376 sequences; 419,152,180 total letters
Searching..................................................done
Score E
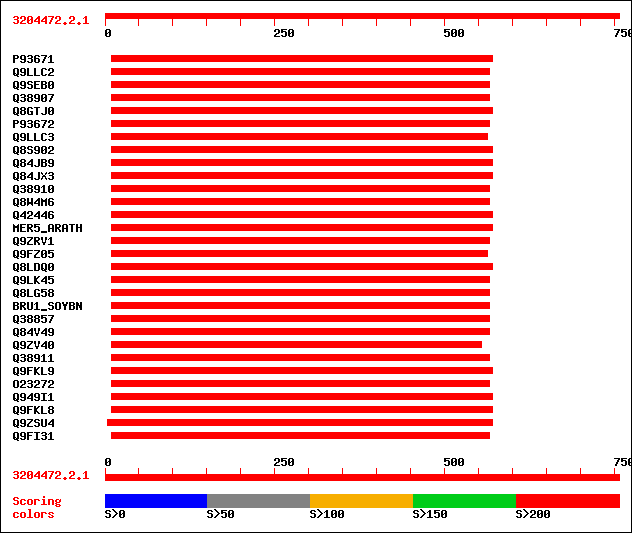
Sequences producing significant alignments: (bits) Value
sptr|P93671|P93671 Xyloglucan endotransglycosylase (XET). 290 1e-77
sptr|Q9LLC2|Q9LLC2 Xyloglucan endotransglycosylase XET2 (EC 2.4.... 266 2e-70
sptr|Q9SEB0|Q9SEB0 ENDOXYLOGLUCAN transferase (AT5g57550/MUA2_12). 260 1e-68
sptr|Q38907|Q38907 Xyloglucan endotransglycosylase-related prote... 260 1e-68
sptr|Q8GTJ0|Q8GTJ0 Xyloglucan endotransglycosylase. 260 2e-68
sptr|P93672|P93672 Xyloglucan endotransglycosylase (XET). 256 2e-67
sptr|Q9LLC3|Q9LLC3 Xyloglucan endotransglycosylase XET1 (EC 2.4.... 256 3e-67
sptr|Q8S902|Q8S902 Syringolide-induced protein 19-1-5. 256 3e-67
sptr|Q84JB9|Q84JB9 Putative xyloglucan endotransglycosylase. 246 2e-64
sptr|Q84JX3|Q84JX3 Putative xyloglucan endotransglycosylase. 246 2e-64
sptr|Q38910|Q38910 XYLOGLUCAN ENDOTRANSGLYCOSYLASE-related prote... 244 1e-63
sptr|Q8W4M6|Q8W4M6 Xyloglucan endo-1, 4-beta-D-glucanase (XTR-6). 243 2e-63
sptr|Q42446|Q42446 XYLOGLUCAN endo-transglycosylase homolog. 241 1e-62
sw|P24806|MER5_ARATH MERI-5 protein precursor (Endo-xyloglucan t... 241 1e-62
sptr|Q9ZRV1|Q9ZRV1 Xyloglucan endotransglycosylase 1. 240 1e-62
sptr|Q9FZ05|Q9FZ05 Xyloglucan endotransglycosylase LeXET2. 240 1e-62
sptr|Q8LDQ0|Q8LDQ0 Xyloglucan endo-1,4-beta-D-glucanase. 239 3e-62
sptr|Q9LK45|Q9LK45 Endoxyloglucan endotransglycosylase. 235 5e-61
sptr|Q8LG58|Q8LG58 Xyloglucan endotransglycosylase, putative. 235 5e-61
sw|P35694|BRU1_SOYBN Brassinosteroid-regulated protein BRU1 prec... 235 5e-61
sptr|Q38857|Q38857 XYLOGLUCAN ENDOTRANSGLYCOSYLASE TCH4 (TCH4 pr... 234 9e-61
sptr|Q84V49|Q84V49 Putative xyloglucan endotransglycosylase. 234 9e-61
sptr|Q9ZV40|Q9ZV40 Putative xyloglucan endo-transglycosylase. 233 2e-60
sptr|Q38911|Q38911 Xyloglucan endotransglycosylase-related prote... 233 2e-60
sptr|Q9FKL9|Q9FKL9 Xyloglucan endotransglycosylase (AT5g57530/MU... 232 5e-60
sptr|O23272|O23272 XYLOGLUCAN ENDOTRANSGLYCOSYLASE-related prote... 230 1e-59
sptr|Q949I1|Q949I1 Xet1 protein. 229 2e-59
sptr|Q9FKL8|Q9FKL8 Xyloglucan endotransglycosylase. 229 3e-59
sptr|Q9ZSU4|Q9ZSU4 XYLOGLUCAN ENDOTRANSGLYCOSYLASE (Putative xyl... 229 4e-59
sptr|Q9FI31|Q9FI31 Xyloglucan endo-1,4-beta-D-glucanase. 227 1e-58
>sptr|P93671|P93671 Xyloglucan endotransglycosylase (XET).
Length = 292
Score = 290 bits (742), Expect = 1e-77
Identities = 139/188 (73%), Positives = 154/188 (81%), Gaps = 1/188 (0%)
Frame = -2
Query: 570 AAGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
AA S D+E D+TWGDGRGK+L+NG+LLTL LD+ SGSGFQS+HEYLFGKIDMQL+LVPGN
Sbjct: 19 AAASFDKEFDVTWGDGRGKILNNGQLLTLGLDKVSGSGFQSKHEYLFGKIDMQLKLVPGN 78
Query: 390 SAGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTAD 211
SAGTVTAYYLSSQG HDEIDFEFLGNV+GEPYTLHTNVFT+GQGQREQQFRLWFDPT D
Sbjct: 79 SAGTVTAYYLSSQGPTHDEIDFEFLGNVTGEPYTLHTNVFTQGQGQREQQFRLWFDPTND 138
Query: 210 FHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRA-L 34
FHTYSILWNPKH+IFMVDDM IRDF NLE K +AFP+ + A +
Sbjct: 139 FHTYSILWNPKHIIFMVDDMPIRDFKNLEGKGIAFPKNQPMRLYSSLWNADDWATQGARV 198
Query: 33 KTDWSTPP 10
KTDWS P
Sbjct: 199 KTDWSHAP 206
>sptr|Q9LLC2|Q9LLC2 Xyloglucan endotransglycosylase XET2 (EC
2.4.1.207).
Length = 284
Score = 266 bits (681), Expect = 2e-70
Identities = 125/187 (66%), Positives = 148/187 (79%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
AG+ + + ITWGDGRG+++DNG+LL+LSLD+ SGSGF+S HEYLFGKIDMQL+LVPGNS
Sbjct: 19 AGNFNDDFKITWGDGRGRIVDNGQLLSLSLDKASGSGFESTHEYLFGKIDMQLKLVPGNS 78
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVTAYYLSSQG HDEIDFEFLGN+SG+PYTLHTNVFT+G+G REQQF LWFDPT DF
Sbjct: 79 AGTVTAYYLSSQGPTHDEIDFEFLGNLSGDPYTLHTNVFTQGKGNREQQFHLWFDPTKDF 138
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
HTYSILWNP H+IF VD + IRDF N+ES+ VAFP++ + L K
Sbjct: 139 HTYSILWNPSHIIFSVDGIPIRDFKNMESRGVAFPKSQPMKIYSSLWNADDWATRGGLIK 198
Query: 30 TDWSTPP 10
TDW+ P
Sbjct: 199 TDWTKAP 205
>sptr|Q9SEB0|Q9SEB0 ENDOXYLOGLUCAN transferase (AT5g57550/MUA2_12).
Length = 284
Score = 260 bits (665), Expect = 1e-68
Identities = 121/187 (64%), Positives = 145/187 (77%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
AG+ D E DITWGDGRGKVL+NG LLTLSLDR SGSGFQ++ EYLFGKIDMQL+LVPGNS
Sbjct: 27 AGTFDTEFDITWGDGRGKVLNNGELLTLSLDRASGSGFQTKKEYLFGKIDMQLKLVPGNS 86
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVTAYYL S+G DEIDFEFLGN++G+PYT+HTNV+T+G+G REQQF LWFDPTADF
Sbjct: 87 AGTVTAYYLKSKGDTWDEIDFEFLGNLTGDPYTMHTNVYTQGKGDREQQFHLWFDPTADF 146
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
HTYS+LWNP H++FMVDD+ +R+F NL+ + +P+ + L K
Sbjct: 147 HTYSVLWNPHHIVFMVDDIPVREFKNLQHMGIQYPKLQPMRLYSSLWNADQWATRGGLVK 206
Query: 30 TDWSTPP 10
TDWS P
Sbjct: 207 TDWSKAP 213
>sptr|Q38907|Q38907 Xyloglucan endotransglycosylase-related protein
(Fragment).
Length = 277
Score = 260 bits (665), Expect = 1e-68
Identities = 121/187 (64%), Positives = 145/187 (77%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
AG+ D E DITWGDGRGKVL+NG LLTLSLDR SGSGFQ++ EYLFGKIDMQL+LVPGNS
Sbjct: 20 AGTFDTEFDITWGDGRGKVLNNGELLTLSLDRASGSGFQTKKEYLFGKIDMQLKLVPGNS 79
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVTAYYL S+G DEIDFEFLGN++G+PYT+HTNV+T+G+G REQQF LWFDPTADF
Sbjct: 80 AGTVTAYYLKSKGDTWDEIDFEFLGNLTGDPYTMHTNVYTQGKGDREQQFHLWFDPTADF 139
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
HTYS+LWNP H++FMVDD+ +R+F NL+ + +P+ + L K
Sbjct: 140 HTYSVLWNPHHIVFMVDDIPVREFKNLQHMGIQYPKLQPMRLYSSLWNADQWATRGGLVK 199
Query: 30 TDWSTPP 10
TDWS P
Sbjct: 200 TDWSKAP 206
>sptr|Q8GTJ0|Q8GTJ0 Xyloglucan endotransglycosylase.
Length = 282
Score = 260 bits (664), Expect = 2e-68
Identities = 124/188 (65%), Positives = 147/188 (78%), Gaps = 1/188 (0%)
Frame = -2
Query: 570 AAGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
+AG+ +Q+ DITWGDGR K+LDNG+ LTLSLD+TSGSGFQS+++YLFGKIDMQ++LVPGN
Sbjct: 20 SAGNFNQDFDITWGDGRAKILDNGQSLTLSLDKTSGSGFQSKNQYLFGKIDMQIKLVPGN 79
Query: 390 SAGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTAD 211
SAGTVTAYYLSS G HDEIDFEFLGN+SG+PY LHTN+FT+G+G REQQF LWFDPT D
Sbjct: 80 SAGTVTAYYLSSLGSTHDEIDFEFLGNLSGDPYILHTNIFTQGKGNREQQFYLWFDPTKD 139
Query: 210 FHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL- 34
FHTYSILWNP+ +IF VD IR+F NLESK V+FP+ + L
Sbjct: 140 FHTYSILWNPQTIIFSVDGTPIREFKNLESKGVSFPKDQGMWIYSSLWNADDWATRGGLV 199
Query: 33 KTDWSTPP 10
KTDWS P
Sbjct: 200 KTDWSKAP 207
>sptr|P93672|P93672 Xyloglucan endotransglycosylase (XET).
Length = 286
Score = 256 bits (654), Expect = 2e-67
Identities = 123/187 (65%), Positives = 146/187 (78%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
AG+ Q+ +++WGDGRGKV+D GR L L+LD+TSGSGFQS+ EYLFGKIDMQ++LVPGNS
Sbjct: 20 AGNFFQDSEMSWGDGRGKVVDGGRGLDLTLDKTSGSGFQSKSEYLFGKIDMQIKLVPGNS 79
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVT +YLSSQG AHDEIDFEFLGNV+GEPYTLHTNVF +GQGQREQQFRLWFDPT F
Sbjct: 80 AGTVTTFYLSSQGTAHDEIDFEFLGNVTGEPYTLHTNVFAQGQGQREQQFRLWFDPTKAF 139
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPAC-XXXXXXXXXRALK 31
HTYSI+WNP+HVIF VD IRDF N E++ V+FP++ + +K
Sbjct: 140 HTYSIIWNPQHVIFAVDGTAIRDFKNHEARGVSFPKSQPMRLYASLWNADDWATQGGRVK 199
Query: 30 TDWSTPP 10
TDWS P
Sbjct: 200 TDWSKAP 206
>sptr|Q9LLC3|Q9LLC3 Xyloglucan endotransglycosylase XET1 (EC
2.4.1.207).
Length = 284
Score = 256 bits (653), Expect = 3e-67
Identities = 120/186 (64%), Positives = 144/186 (77%), Gaps = 1/186 (0%)
Frame = -2
Query: 564 GSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNSA 385
G+ ++ D+TWGDGR K+L+NG++LTLSLD+ SGSGFQS+++YLFGKIDMQL+LVPGNSA
Sbjct: 27 GNFYRDFDLTWGDGRAKILNNGQMLTLSLDKVSGSGFQSKNQYLFGKIDMQLKLVPGNSA 86
Query: 384 GTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADFH 205
GTVTAYYLSSQG HDEIDFEFLGN+SG+PY LHTN+FT+G+G REQQF LWFDPT DFH
Sbjct: 87 GTVTAYYLSSQGPTHDEIDFEFLGNLSGDPYILHTNIFTQGKGNREQQFYLWFDPTKDFH 146
Query: 204 TYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-KT 28
TYSILWNP+H+IF VD + IRDF NLE + FP+ + L KT
Sbjct: 147 TYSILWNPQHIIFSVDGIPIRDFKNLEKYRIPFPKIQPMRLYSSLWDADNWATRGGLVKT 206
Query: 27 DWSTPP 10
DWS P
Sbjct: 207 DWSKAP 212
>sptr|Q8S902|Q8S902 Syringolide-induced protein 19-1-5.
Length = 285
Score = 256 bits (653), Expect = 3e-67
Identities = 122/188 (64%), Positives = 145/188 (77%), Gaps = 1/188 (0%)
Frame = -2
Query: 570 AAGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
+AG+ Q+ DITWGDGR K+L+NG LLTLSLD+ SGSGFQS++EYLFGKIDMQL+LVPGN
Sbjct: 20 SAGNFHQDFDITWGDGRAKILNNGELLTLSLDKASGSGFQSKNEYLFGKIDMQLKLVPGN 79
Query: 390 SAGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTAD 211
SAGTVTAYYLSS+G DEIDFEFLGN+SG+PY LHTNVF++G+G REQQF LWFDPTAD
Sbjct: 80 SAGTVTAYYLSSKGATWDEIDFEFLGNLSGDPYILHTNVFSQGKGNREQQFYLWFDPTAD 139
Query: 210 FHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL- 34
FHTYSILWNP+ ++F VD IR+F N+ESK V FP+ + L
Sbjct: 140 FHTYSILWNPQRIVFSVDGSPIREFKNMESKGVPFPKNQAMRIYSSLWNADDWATRGGLV 199
Query: 33 KTDWSTPP 10
KTDW+ P
Sbjct: 200 KTDWTQAP 207
>sptr|Q84JB9|Q84JB9 Putative xyloglucan endotransglycosylase.
Length = 291
Score = 246 bits (629), Expect = 2e-64
Identities = 117/188 (62%), Positives = 143/188 (76%), Gaps = 1/188 (0%)
Frame = -2
Query: 570 AAGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
+A + Q DITWGDGR ++L+NG LL+LSLD+ SGSGFQSR+EYL+GKIDMQ++LVPGN
Sbjct: 25 SASNFHQSTDITWGDGRAQILNNGDLLSLSLDKASGSGFQSRNEYLYGKIDMQIKLVPGN 84
Query: 390 SAGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTAD 211
SAGTVTAYYL S+G DEIDFEFLGN+SG+PYT+HTNVF++G+G REQQF LWFDPTAD
Sbjct: 85 SAGTVTAYYLRSEGSTWDEIDFEFLGNLSGDPYTVHTNVFSQGKGNREQQFHLWFDPTAD 144
Query: 210 FHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL- 34
FHTYSILWNP+ ++F VD IR+F N+ES VAFP+ + L
Sbjct: 145 FHTYSILWNPQRIVFYVDGTPIREFKNMESIGVAFPKNQPMRLQSSLWNADDWATRGGLI 204
Query: 33 KTDWSTPP 10
KTDW+ P
Sbjct: 205 KTDWTQAP 212
>sptr|Q84JX3|Q84JX3 Putative xyloglucan endotransglycosylase.
Length = 283
Score = 246 bits (628), Expect = 2e-64
Identities = 119/188 (63%), Positives = 141/188 (75%), Gaps = 1/188 (0%)
Frame = -2
Query: 570 AAGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
++ + +Q+ ITWGDGR K+L+NG LLTLSLD+ SGSGFQS++EYLFGKIDMQL+LV GN
Sbjct: 20 SSANFNQDFQITWGDGRAKILNNGELLTLSLDKASGSGFQSQNEYLFGKIDMQLKLVAGN 79
Query: 390 SAGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTAD 211
SAGTVTAYYLSS+G DEIDFEFLGN+SG+PYTLHTNVF++G+G REQQF LWFDPTAD
Sbjct: 80 SAGTVTAYYLSSKGSTWDEIDFEFLGNLSGDPYTLHTNVFSQGKGNREQQFHLWFDPTAD 139
Query: 210 FHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL- 34
FHTYSILWNP +IF VD IR+F N ES V FP+ + L
Sbjct: 140 FHTYSILWNPNRIIFSVDGTPIREFKNWESNGVPFPKDQPMRIYSSLWNADDWATRGGLV 199
Query: 33 KTDWSTPP 10
KTDW+ P
Sbjct: 200 KTDWTKAP 207
>sptr|Q38910|Q38910 XYLOGLUCAN ENDOTRANSGLYCOSYLASE-related protein
(XYLOGLUCAN endo-1, 4-beta-D-glucanase) (XTR-6).
Length = 286
Score = 244 bits (622), Expect = 1e-63
Identities = 115/187 (61%), Positives = 142/187 (75%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
+ + ++++ITWGDGRG++ +NG LLTLSLD+ SGSGFQS++EYLFGKIDMQ++LV GNS
Sbjct: 23 SANFQRDVEITWGDGRGQITNNGDLLTLSLDKASGSGFQSKNEYLFGKIDMQIKLVAGNS 82
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVTAYYL S G DEIDFEFLGN+SG+PYTLHTNVFT+G+G REQQF+LWFDPT+DF
Sbjct: 83 AGTVTAYYLKSPGSTWDEIDFEFLGNLSGDPYTLHTNVFTQGKGDREQQFKLWFDPTSDF 142
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
HTYSILWNP+ +IF VD IR+F N+ES+ FP+ + L K
Sbjct: 143 HTYSILWNPQRIIFSVDGTPIREFKNMESQGTLFPKNQPMRMYSSLWNAEEWATRGGLVK 202
Query: 30 TDWSTPP 10
TDWS P
Sbjct: 203 TDWSKAP 209
>sptr|Q8W4M6|Q8W4M6 Xyloglucan endo-1, 4-beta-D-glucanase (XTR-6).
Length = 286
Score = 243 bits (620), Expect = 2e-63
Identities = 115/187 (61%), Positives = 141/187 (75%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
+ + ++++ITWGDGRG++ +NG LLTLSLD+ SGSGFQS++EYLFGKIDMQ+ LV GNS
Sbjct: 23 SANFQRDVEITWGDGRGQITNNGDLLTLSLDKASGSGFQSKNEYLFGKIDMQIELVAGNS 82
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVTAYYL S G DEIDFEFLGN+SG+PYTLHTNVFT+G+G REQQF+LWFDPT+DF
Sbjct: 83 AGTVTAYYLKSPGSTWDEIDFEFLGNLSGDPYTLHTNVFTQGKGDREQQFKLWFDPTSDF 142
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
HTYSILWNP+ +IF VD IR+F N+ES+ FP+ + L K
Sbjct: 143 HTYSILWNPQRIIFSVDGTPIREFKNMESQGTLFPKNQPMRMYSSLWNAEEWATRGGLVK 202
Query: 30 TDWSTPP 10
TDWS P
Sbjct: 203 TDWSKAP 209
>sptr|Q42446|Q42446 XYLOGLUCAN endo-transglycosylase homolog.
Length = 280
Score = 241 bits (614), Expect = 1e-62
Identities = 112/188 (59%), Positives = 139/188 (73%), Gaps = 1/188 (0%)
Frame = -2
Query: 570 AAGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
A G+ Q++DITWGDGRGK+LDNG+LLTLS+DR+SGSGFQS+ +YL+G+ DMQL+LVPG+
Sbjct: 20 AGGNFYQDVDITWGDGRGKILDNGQLLTLSMDRSSGSGFQSKAQYLYGRFDMQLKLVPGD 79
Query: 390 SAGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTAD 211
SAGTV +YLSSQG HDEIDFEFLGN SGEPYT+HTNV+++G+G REQQFR+WFDPTA
Sbjct: 80 SAGTVATFYLSSQGSQHDEIDFEFLGNASGEPYTVHTNVYSQGKGGREQQFRMWFDPTAA 139
Query: 210 FHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXC-XSTPACXXXXXXXXXRAL 34
FH YS+LWNP HV+F VD + IR+F V FP + +
Sbjct: 140 FHAYSVLWNPAHVVFYVDGVPIREFRRRGDGTVPFPTSQPMRVYASVWDAEEWATQGGRV 199
Query: 33 KTDWSTPP 10
+TDWS P
Sbjct: 200 RTDWSKAP 207
>sw|P24806|MER5_ARATH MERI-5 protein precursor (Endo-xyloglucan
transferase) (Xyloglucan endo-1,4-beta-D-glucanase).
Length = 269
Score = 241 bits (614), Expect = 1e-62
Identities = 113/188 (60%), Positives = 142/188 (75%), Gaps = 1/188 (0%)
Frame = -2
Query: 570 AAGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
+A + ++++ WG+GRGK+L+NG+LLTLSLD++SGSGFQS+ EYLFGKIDMQ++LVPGN
Sbjct: 20 SAADFNTDVNVAWGNGRGKILNNGQLLTLSLDKSSGSGFQSKTEYLFGKIDMQIKLVPGN 79
Query: 390 SAGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTAD 211
SAGTVT +YL S+G DEIDFEFLGN+SG+PYTLHTNV+T+G+G +EQQF LWFDPTA+
Sbjct: 80 SAGTVTTFYLKSEGSTWDEIDFEFLGNMSGDPYTLHTNVYTQGKGDKEQQFHLWFDPTAN 139
Query: 210 FHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL- 34
FHTYSILWNP+ +I VDD IR+F N ES V FP+ + L
Sbjct: 140 FHTYSILWNPQRIILTVDDTPIREFKNYESLGVLFPKNKPMRMYASLWNADDWATRGGLV 199
Query: 33 KTDWSTPP 10
KTDWS P
Sbjct: 200 KTDWSKAP 207
>sptr|Q9ZRV1|Q9ZRV1 Xyloglucan endotransglycosylase 1.
Length = 292
Score = 240 bits (613), Expect = 1e-62
Identities = 114/187 (60%), Positives = 141/187 (75%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
AG+L Q+ DITWGDGR +L+NG LLTLSLD+ SGSGFQS++++LFGKID Q++LVPGNS
Sbjct: 25 AGNLHQDFDITWGDGRAMILNNGELLTLSLDKASGSGFQSKNQFLFGKIDTQIKLVPGNS 84
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVTAYYLSS+G DEID+EFLGN+SG+PYT+HTN++T+G+G REQQF LWFDPTA+F
Sbjct: 85 AGTVTAYYLSSKGSTWDEIDYEFLGNLSGDPYTIHTNIYTQGKGNREQQFHLWFDPTANF 144
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXC-XSTPACXXXXXXXXXRALK 31
HTYSILWNP+ +IF VD IR+F N ES V P+ + LK
Sbjct: 145 HTYSILWNPQAIIFSVDGTPIREFKNSESIGVPIPKKQPMRLYSSLWNADDWATRGGLLK 204
Query: 30 TDWSTPP 10
TDW+ P
Sbjct: 205 TDWARTP 211
>sptr|Q9FZ05|Q9FZ05 Xyloglucan endotransglycosylase LeXET2.
Length = 275
Score = 240 bits (613), Expect = 1e-62
Identities = 114/187 (60%), Positives = 141/187 (75%), Gaps = 2/187 (1%)
Frame = -2
Query: 564 GSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNSA 385
G+ DQE D+TWG GR K+L+NG+LLTLSLDR+SGSGF+S+ +Y+F KIDM+++LVPGNSA
Sbjct: 25 GTFDQEFDVTWGYGRVKILENGQLLTLSLDRSSGSGFKSKQQYMFAKIDMKIKLVPGNSA 84
Query: 384 GTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADFH 205
GT T YYLSS G AHDEIDFEFLGNVSGEPYTLHTNV+ +G+G REQQF LWFDPT DFH
Sbjct: 85 GTATTYYLSSVGSAHDEIDFEFLGNVSGEPYTLHTNVYAQGKGDREQQFHLWFDPTKDFH 144
Query: 204 TYSILWNPKHVIFMVDDMXIRDFXNLE-SKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
TYSILWNP+++IF+VD IR + NLE + + +P+ + L +
Sbjct: 145 TYSILWNPRNIIFLVDGTPIRQYKNLEATNGIPYPKNQPMWLYSSLWNAEEWATRGGLVR 204
Query: 30 TDWSTPP 10
TDWS P
Sbjct: 205 TDWSKAP 211
>sptr|Q8LDQ0|Q8LDQ0 Xyloglucan endo-1,4-beta-D-glucanase.
Length = 269
Score = 239 bits (610), Expect = 3e-62
Identities = 112/188 (59%), Positives = 141/188 (75%), Gaps = 1/188 (0%)
Frame = -2
Query: 570 AAGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
+A + ++++ WG+GRGK+L+NG+LLTLSLD++SGSGFQS+ EY FGKIDMQ++LVPGN
Sbjct: 20 SAADFNTDVNVAWGNGRGKILNNGQLLTLSLDKSSGSGFQSKTEYFFGKIDMQIKLVPGN 79
Query: 390 SAGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTAD 211
SAGTVT +YL S+G DEIDFEFLGN+SG+PYTLHTNV+T+G+G +EQQF LWFDPTA+
Sbjct: 80 SAGTVTTFYLKSEGSTWDEIDFEFLGNMSGDPYTLHTNVYTQGKGDKEQQFHLWFDPTAN 139
Query: 210 FHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL- 34
FHTYSILWNP+ +I VDD IR+F N ES V FP+ + L
Sbjct: 140 FHTYSILWNPQRIILTVDDTPIREFKNYESLGVLFPKNKPMRMYASLWNADDWATRGGLV 199
Query: 33 KTDWSTPP 10
KTDWS P
Sbjct: 200 KTDWSKAP 207
>sptr|Q9LK45|Q9LK45 Endoxyloglucan endotransglycosylase.
Length = 291
Score = 235 bits (599), Expect = 5e-61
Identities = 111/187 (59%), Positives = 138/187 (73%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
+GS ++E D+TWG+ RGK+ G++L+LSLDR SGSGF+S+ EYLFG+IDMQL+LV GNS
Sbjct: 24 SGSFNEEFDLTWGEHRGKIFSGGKMLSLSLDRVSGSGFKSKKEYLFGRIDMQLKLVAGNS 83
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVTAYYLSS+G HDEIDFEFLGN +G+PY LHTNVF +G+G REQQF LWFDPT +F
Sbjct: 84 AGTVTAYYLSSEGPTHDEIDFEFLGNETGKPYVLHTNVFAQGKGNREQQFYLWFDPTKNF 143
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
HTYS++W P+H+IFMVD++ IR F N E V FP+ + L K
Sbjct: 144 HTYSLVWRPQHIIFMVDNVPIRVFNNAEQLGVPFPKNQPMKIYSSLWNADDWATRGGLVK 203
Query: 30 TDWSTPP 10
TDWS P
Sbjct: 204 TDWSKAP 210
>sptr|Q8LG58|Q8LG58 Xyloglucan endotransglycosylase, putative.
Length = 291
Score = 235 bits (599), Expect = 5e-61
Identities = 111/187 (59%), Positives = 138/187 (73%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
+GS ++E D+TWG+ RGK+ G++L+LSLDR SGSGF+S+ EYLFG+IDMQL+LV GNS
Sbjct: 24 SGSFNEEFDLTWGEHRGKIFSGGKMLSLSLDRVSGSGFKSKKEYLFGRIDMQLKLVAGNS 83
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVTAYYLSS+G HDEIDFEFLGN +G+PY LHTNVF +G+G REQQF LWFDPT +F
Sbjct: 84 AGTVTAYYLSSEGPTHDEIDFEFLGNETGKPYVLHTNVFAQGKGNREQQFYLWFDPTKNF 143
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
HTYS++W P+H+IFMVD++ IR F N E V FP+ + L K
Sbjct: 144 HTYSLVWRPQHIIFMVDNVPIRVFNNAEQLGVPFPKNQPMKIYSSLWNADDWATRGGLVK 203
Query: 30 TDWSTPP 10
TDWS P
Sbjct: 204 TDWSKAP 210
>sw|P35694|BRU1_SOYBN Brassinosteroid-regulated protein BRU1
precursor.
Length = 283
Score = 235 bits (599), Expect = 5e-61
Identities = 113/187 (60%), Positives = 137/187 (73%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
AGS Q+ D+TWG R K+ + G+LL+LSLD+ SGSGF+S+ EYLFG+IDMQL+LV GNS
Sbjct: 29 AGSFYQDFDLTWGGDRAKIFNGGQLLSLSLDKVSGSGFKSKKEYLFGRIDMQLKLVAGNS 88
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVTAYYLSSQG HDEIDFEFLGN+SG+PY LHTN+FT+G+G REQQF LWFDPT +F
Sbjct: 89 AGTVTAYYLSSQGPTHDEIDFEFLGNLSGDPYILHTNIFTQGKGNREQQFYLWFDPTRNF 148
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
HTYSI+W P+H+IF+VD+ IR F N E V FP+ + L K
Sbjct: 149 HTYSIIWKPQHIIFLVDNTPIRVFKNAEPLGVPFPKNQPMRIYSSLWNADDWATRGGLVK 208
Query: 30 TDWSTPP 10
TDWS P
Sbjct: 209 TDWSKAP 215
>sptr|Q38857|Q38857 XYLOGLUCAN ENDOTRANSGLYCOSYLASE TCH4 (TCH4
protein) (AT5g57560/MUA2_13).
Length = 284
Score = 234 bits (597), Expect = 9e-61
Identities = 110/187 (58%), Positives = 139/187 (74%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
+ + ++++ITWGDGRG++ +NG LLTLSLD++SGSGFQS++EYLFGK+ MQ++LVPGNS
Sbjct: 20 SANFQRDVEITWGDGRGQIKNNGELLTLSLDKSSGSGFQSKNEYLFGKVSMQMKLVPGNS 79
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVT YL S G DEIDFEFLGN SGEPYTLHTNV+T+G+G +EQQF+LWFDPTA+F
Sbjct: 80 AGTVTTLYLKSPGTTWDEIDFEFLGNSSGEPYTLHTNVYTQGKGDKEQQFKLWFDPTANF 139
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
HTY+ILWNP+ +IF VD IR+F N+ES FP+ + L K
Sbjct: 140 HTYTILWNPQRIIFTVDGTPIREFKNMESLGTLFPKNKPMRMYSSLWNADDWATRGGLVK 199
Query: 30 TDWSTPP 10
TDWS P
Sbjct: 200 TDWSKAP 206
>sptr|Q84V49|Q84V49 Putative xyloglucan endotransglycosylase.
Length = 297
Score = 234 bits (597), Expect = 9e-61
Identities = 107/187 (57%), Positives = 137/187 (73%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
AG+ +E+D+TWGDGRGK+++NG L+TLSLD+ SGSGFQS+ +YL G+ DM+++LVPGNS
Sbjct: 34 AGNFYKEVDVTWGDGRGKIIENGNLITLSLDKASGSGFQSKKQYLHGRFDMKIKLVPGNS 93
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVTAYYL S+G + DEIDFEFLGNVSG+PY +HTN++ GQG REQQF LWFDPTADF
Sbjct: 94 AGTVTAYYLRSEGSSWDEIDFEFLGNVSGQPYVVHTNIYVGGQGNREQQFYLWFDPTADF 153
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
HTYS LWNP ++F VD IR+F N+E+ +P+ + L K
Sbjct: 154 HTYSFLWNPARILFYVDGRPIREFKNMEANGAPYPKKQPMRLYASLWNGEDWATRGGLIK 213
Query: 30 TDWSTPP 10
TDW+ P
Sbjct: 214 TDWTKAP 220
>sptr|Q9ZV40|Q9ZV40 Putative xyloglucan endo-transglycosylase.
Length = 305
Score = 233 bits (595), Expect = 2e-60
Identities = 110/183 (60%), Positives = 136/183 (74%), Gaps = 1/183 (0%)
Frame = -2
Query: 555 DQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNSAGTV 376
+Q++DITWGDGRG +L+NG LL L LD++SGSGFQS+ EYL+GK+DMQ++LVPGNSAGTV
Sbjct: 29 NQDIDITWGDGRGNILNNGTLLNLGLDQSSGSGFQSKAEYLYGKVDMQIKLVPGNSAGTV 88
Query: 375 TAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADFHTYS 196
T +YL SQG DEIDFEFLGNVSG+PY +HTNV+T+G+G REQQF LWFDPTA FH YS
Sbjct: 89 TTFYLKSQGLTWDEIDFEFLGNVSGDPYIVHTNVYTQGKGDREQQFYLWFDPTAAFHNYS 148
Query: 195 ILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-KTDWS 19
ILWNP H++F +D IR+F NLE VA+P+ + L KT+WS
Sbjct: 149 ILWNPSHIVFYIDGKPIREFKNLEVLGVAYPKNQPMRMYGSLWNADDWATRGGLVKTNWS 208
Query: 18 TPP 10
P
Sbjct: 209 QGP 211
>sptr|Q38911|Q38911 Xyloglucan endotransglycosylase-related protein
XTR-7 (Putative xyloglucan endotransglycosylase-related
protein).
Length = 289
Score = 233 bits (594), Expect = 2e-60
Identities = 111/187 (59%), Positives = 136/187 (72%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
A + E D+TWGD RGK+ + G +L+LSLD+ SGSGF+S+ EYLFG+IDMQL+LV GNS
Sbjct: 25 ASNFFDEFDLTWGDHRGKIFNGGNMLSLSLDQVSGSGFKSKKEYLFGRIDMQLKLVAGNS 84
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVTAYYLSSQG HDEIDFEFLGN +G+PY LHTNVF +G+G REQQF LWFDPT +F
Sbjct: 85 AGTVTAYYLSSQGATHDEIDFEFLGNETGKPYVLHTNVFAQGKGDREQQFYLWFDPTKNF 144
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
HTYSI+W P+H+IF+VD++ IR F N E V FP++ + L K
Sbjct: 145 HTYSIVWRPQHIIFLVDNLPIRVFNNAEKLGVPFPKSQPMRIYSSLWNADDWATRGGLVK 204
Query: 30 TDWSTPP 10
TDWS P
Sbjct: 205 TDWSKAP 211
>sptr|Q9FKL9|Q9FKL9 Xyloglucan endotransglycosylase
(AT5g57530/MUA2_10).
Length = 285
Score = 232 bits (591), Expect = 5e-60
Identities = 110/188 (58%), Positives = 138/188 (73%), Gaps = 1/188 (0%)
Frame = -2
Query: 570 AAGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
A GS DITWG GR + ++G+LLT +LD+TSGSGFQS+ EYLFGKIDM+++LVPGN
Sbjct: 23 ATGSFYDSFDITWGAGRANIFESGQLLTCTLDKTSGSGFQSKKEYLFGKIDMKIKLVPGN 82
Query: 390 SAGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTAD 211
SAGTVTAYYLSS+G DEIDFEFLGNV+G+PY +HTNVFT G+G RE QF LWFDPTAD
Sbjct: 83 SAGTVTAYYLSSKGETWDEIDFEFLGNVTGQPYVIHTNVFTGGKGNREMQFYLWFDPTAD 142
Query: 210 FHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXC-XSTPACXXXXXXXXXRAL 34
FHTY++LWNP ++IF+VD + IR F N E+ VA+P++ + +
Sbjct: 143 FHTYTVLWNPLNIIFLVDGIPIRVFKNNEANGVAYPKSQPMKIYSSLWEADDWATQGGKV 202
Query: 33 KTDWSTPP 10
KTDW+ P
Sbjct: 203 KTDWTNAP 210
>sptr|O23272|O23272 XYLOGLUCAN ENDOTRANSGLYCOSYLASE-related protein
XTR-7.
Length = 289
Score = 230 bits (587), Expect = 1e-59
Identities = 110/187 (58%), Positives = 135/187 (72%), Gaps = 1/187 (0%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGNS 388
A + E D+TWGD RGK+ + G +L+LSLD+ SGSGF+S+ EYL G+IDMQL+LV GNS
Sbjct: 25 ASNFFDEFDLTWGDHRGKIFNGGNMLSLSLDQVSGSGFKSKKEYLVGRIDMQLKLVAGNS 84
Query: 387 AGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTADF 208
AGTVTAYYLSSQG HDEIDFEFLGN +G+PY LHTNVF +G+G REQQF LWFDPT +F
Sbjct: 85 AGTVTAYYLSSQGATHDEIDFEFLGNETGKPYVLHTNVFAQGKGDREQQFYLWFDPTKNF 144
Query: 207 HTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL-K 31
HTYSI+W P+H+IF+VD++ IR F N E V FP++ + L K
Sbjct: 145 HTYSIVWRPQHIIFLVDNLPIRVFNNAEKLGVPFPKSQPMRIYSSLWNADDWATRGGLVK 204
Query: 30 TDWSTPP 10
TDWS P
Sbjct: 205 TDWSKAP 211
>sptr|Q949I1|Q949I1 Xet1 protein.
Length = 291
Score = 229 bits (585), Expect = 2e-59
Identities = 115/189 (60%), Positives = 135/189 (71%), Gaps = 2/189 (1%)
Frame = -2
Query: 570 AAGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
AA S D+E DITWGDGRGK+L+NG+LLTL LD+TSGSGFQS+ EYLFGKIDMQL+LV G
Sbjct: 20 AAASFDKEFDITWGDGRGKILNNGQLLTLGLDKTSGSGFQSKREYLFGKIDMQLKLVAGQ 79
Query: 390 SAGTVTA-YYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTA 214
+ S G HD+IDFEFLGNV+GEPYTLHTNVF +GQG+REQQFRLWFDPT
Sbjct: 80 LPPAPSPPTTCRSMGATHDKIDFEFLGNVTGEPYTLHTNVFAKGQGKREQQFRLWFDPTK 139
Query: 213 DFHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPAC-XXXXXXXXXRA 37
FHTYSI+WNP+HVIF VD IRDF N E++ V+FP+ +
Sbjct: 140 AFHTYSIVWNPQHVIFAVDGTPIRDFKNHEARGVSFPKNQPMRLYASLWNADDWATQGGR 199
Query: 36 LKTDWSTPP 10
+KTDWS P
Sbjct: 200 VKTDWSHAP 208
>sptr|Q9FKL8|Q9FKL8 Xyloglucan endotransglycosylase.
Length = 284
Score = 229 bits (584), Expect = 3e-59
Identities = 109/188 (57%), Positives = 139/188 (73%), Gaps = 1/188 (0%)
Frame = -2
Query: 570 AAGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
+AGS DITWG+GR ++++G+LLT +LD+ SGSGFQS+ EYLFGKIDM+++LV GN
Sbjct: 22 SAGSFYDNFDITWGNGRANIVESGQLLTCTLDKISGSGFQSKKEYLFGKIDMKMKLVAGN 81
Query: 390 SAGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTAD 211
SAGTVTAYYLSS+G DEIDFEFLGNV+G+PY LHTNVFT G+G RE QF LWFDPTAD
Sbjct: 82 SAGTVTAYYLSSKGETWDEIDFEFLGNVTGQPYVLHTNVFTGGKGNREMQFYLWFDPTAD 141
Query: 210 FHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXC-XSTPACXXXXXXXXXRAL 34
FHTY++LWNP ++IF+VD + IR F N E+ VA+P++ + +
Sbjct: 142 FHTYTVLWNPLNIIFLVDGIPIRVFKNNEANGVAYPKSQPMKIYSSLWEADDWATQGGKV 201
Query: 33 KTDWSTPP 10
KTDW+ P
Sbjct: 202 KTDWTNAP 209
>sptr|Q9ZSU4|Q9ZSU4 XYLOGLUCAN ENDOTRANSGLYCOSYLASE (Putative
xyloglucan endo-1, 4-beta-D-glucanase).
Length = 287
Score = 229 bits (583), Expect = 4e-59
Identities = 110/190 (57%), Positives = 137/190 (72%), Gaps = 1/190 (0%)
Frame = -2
Query: 570 AAGSLDQELDITWGDGRGKVLDNGRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
+AG+ + DITWG+GR + +NG+LLT +LD+ SGSGFQS+ EYLFGKIDM+L+LV GN
Sbjct: 26 SAGNFYESFDITWGNGRANIFENGQLLTCTLDKVSGSGFQSKKEYLFGKIDMKLKLVAGN 85
Query: 390 SAGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTAD 211
SAGTVTAYYLSS+G A DEIDFEFLGN +G PYT+HTNVFT G+G RE QFRLWFDPTAD
Sbjct: 86 SAGTVTAYYLSSKGTAWDEIDFEFLGNRTGHPYTIHTNVFTGGKGDREMQFRLWFDPTAD 145
Query: 210 FHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXC-XSTPACXXXXXXXXXRAL 34
FHTY++ WNP ++IF+VD + IR F N E VA+P+ + +
Sbjct: 146 FHTYTVHWNPVNIIFLVDGIPIRVFKNNEKNGVAYPKNQPMRIYSSLWEADDWATEGGRV 205
Query: 33 KTDWSTPPLR 4
K DWS P +
Sbjct: 206 KIDWSNAPFK 215
>sptr|Q9FI31|Q9FI31 Xyloglucan endo-1,4-beta-D-glucanase.
Length = 282
Score = 227 bits (579), Expect = 1e-58
Identities = 109/188 (57%), Positives = 135/188 (71%), Gaps = 2/188 (1%)
Frame = -2
Query: 567 AGSLDQELDITWGDGRGKVLDN-GRLLTLSLDRTSGSGFQSRHEYLFGKIDMQLRLVPGN 391
AGS +++ I WGDGRGK+LDN G LL+LSLD+ SGSGFQS E+L+GK+++Q++LVPGN
Sbjct: 26 AGSFHKDVQIHWGDGRGKILDNVGNLLSLSLDKFSGSGFQSHQEFLYGKVEVQMKLVPGN 85
Query: 390 SAGTVTAYYLSSQGGAHDEIDFEFLGNVSGEPYTLHTNVFTRGQGQREQQFRLWFDPTAD 211
SAGTVT +YL S G DEIDFEFLGN+SG PYTLHTNV+T+G G +EQQF LWFDPT D
Sbjct: 86 SAGTVTTFYLKSPGTTWDEIDFEFLGNISGHPYTLHTNVYTKGTGDKEQQFHLWFDPTVD 145
Query: 210 FHTYSILWNPKHVIFMVDDMXIRDFXNLESKVVAFPRASXCXSTPACXXXXXXXXXRAL- 34
FHTY I+WNP+ VIF +D + IR+F N E+ V FP+ + L
Sbjct: 146 FHTYCIIWNPQRVIFTIDGIPIREFKNSEALGVPFPKHQPMRLYASLWEAEHWATRGGLE 205
Query: 33 KTDWSTPP 10
KTDWS P
Sbjct: 206 KTDWSKAP 213
Database: /db/trembl-ebi/tmp/swall
Posted date: Jan 2, 2004 7:22 PM
Number of letters in database: 419,152,180
Number of sequences in database: 1,313,376
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 382,356,460
Number of Sequences: 1313376
Number of extensions: 6355897
Number of successful extensions: 21065
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 20106
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 20861
length of database: 419,152,180
effective HSP length: 120
effective length of database: 261,547,060
effective search space used: 34262664860
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

