BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
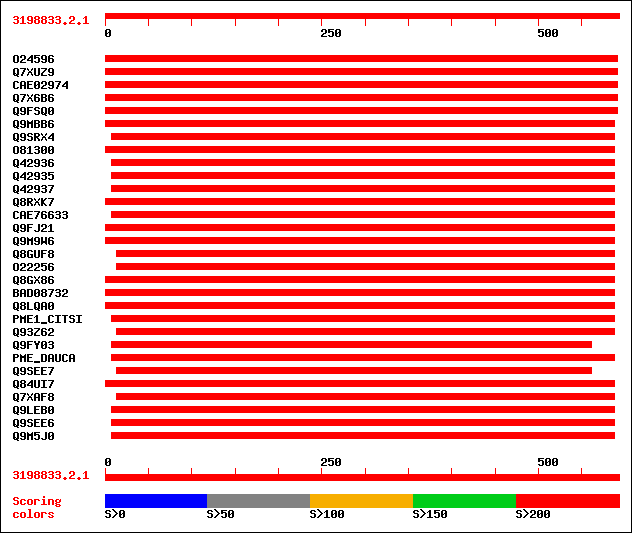
Query= 3198833.2.1
(593 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|O24596|O24596 Pectin methylesterase-like protein. 397 e-110
sptr|Q7XUZ9|Q7XUZ9 OSJNBa0036B21.20 protein. 390 e-108
sptrnew|CAE02974|CAE02974 OSJNBb0079B02.7 protein. 313 8e-85
sptr|Q7X6B6|Q7X6B6 OSJNBb0079B02.8 protein (OSJNBa0063C18.2 prot... 313 8e-85
sptr|Q9FSQ0|Q9FSQ0 Putative pectin methylesterase. 313 8e-85
sptr|Q9MBB6|Q9MBB6 Pectin methylesterase. 278 4e-74
sptr|Q9SRX4|Q9SRX4 F22D16.20 protein. 254 7e-67
sptr|O81300|O81300 T14P8.14 protein (AT4G02330 protein) (Hypothe... 249 1e-65
sptr|Q42936|Q42936 Pectin methylesterase (EC 3.1.1.11) (Pectines... 249 1e-65
sptr|Q42935|Q42935 Pectin methylesterase (EC 3.1.1.11) (Pectines... 249 1e-65
sptr|Q42937|Q42937 Pectin methylesterase (EC 3.1.1.11) (Fragment). 247 9e-65
sptr|Q8RXK7|Q8RXK7 Hypothetical protein. 246 1e-64
sptrnew|CAE76633|CAE76633 Pectin methylesterase (EC 3.1.1.11) (F... 246 1e-64
sptr|Q9FJ21|Q9FJ21 Pectin methylesterase (EC 3.1.1.11) (AT5g4918... 245 3e-64
sptr|Q9M9W6|Q9M9W6 Putative pectinesterase. 241 6e-63
sptr|Q8GUF8|Q8GUF8 At2g47550/T30B22.15. 239 2e-62
sptr|O22256|O22256 Putative pectinesterase. 239 2e-62
sptr|Q8GX86|Q8GX86 Putative pectinesterase (At3g05610). 239 2e-62
sptrnew|BAD08732|BAD08732 Putative pectinesterase. 238 5e-62
sptr|Q8LQA0|Q8LQA0 Putative pectinesterase. 237 9e-62
sw|O04886|PME1_CITSI Pectinesterase 1.1 precursor (EC 3.1.1.11) ... 236 1e-61
sptr|Q93Z62|Q93Z62 At2g47550/T30B22.15. 236 2e-61
sptr|Q9FY03|Q9FY03 Putative pectin methylesterase precursor (EC ... 235 4e-61
sw|P83218|PME_DAUCA Pectinesterase (EC 3.1.1.11) (Pectin methyle... 235 4e-61
sptr|Q9SEE7|Q9SEE7 Pectin methyl esterase (EC 3.1.1.11) (Pectine... 235 4e-61
sptr|Q84UI7|Q84UI7 Pectin methylesterase. 234 6e-61
sptr|Q7XAF8|Q7XAF8 Papillar cell-specific pectin methylesterase-... 233 1e-60
sptr|Q9LEB0|Q9LEB0 Pectin methylesterase (EC 3.1.1.11) (Pectines... 233 1e-60
sptr|Q9SEE6|Q9SEE6 Pectin methyl esterase (EC 3.1.1.11) (Pectine... 233 2e-60
sptr|Q9M5J0|Q9M5J0 Pectin methylesterase isoform alpha (EC 3.1.1... 233 2e-60
>sptr|O24596|O24596 Pectin methylesterase-like protein.
Length = 563
Score = 397 bits (1020), Expect = e-110
Identities = 186/197 (94%), Positives = 191/197 (96%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYV RRQFFRNCV+SGTIDFIF
Sbjct: 344 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVQPRRQFFRNCVVSGTIDFIF 403
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 360
GNSAAVFQNCLI+TRRPMDNQQNSVTAHG TDPNMKSGLVIQNCRLVPDQKLFPDRFKIP
Sbjct: 404 GNSAAVFQNCLIITRRPMDNQQNSVTAHGPTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 463
Query: 361 SYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVT 540
SYLGRPWKEFSRLVIMESTIADF+KPEGYMPWNGDF +KTLYYAEYNNRGPGAGTSKRV
Sbjct: 464 SYLGRPWKEFSRLVIMESTIADFVKPEGYMPWNGDFALKTLYYAEYNNRGPGAGTSKRVN 523
Query: 541 WPGFHVIGRKDAEQFTA 591
WPGFHVIGRK+AE FTA
Sbjct: 524 WPGFHVIGRKEAEPFTA 540
>sptr|Q7XUZ9|Q7XUZ9 OSJNBa0036B21.20 protein.
Length = 568
Score = 390 bits (1002), Expect = e-108
Identities = 182/197 (92%), Positives = 190/197 (96%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
MGFHNTAGAERHQAVALR+ GDL AFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF
Sbjct: 349 MGFHNTAGAERHQAVALRINGDLGAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 408
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 360
GNSAAVFQNCLI+TRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP
Sbjct: 409 GNSAAVFQNCLIITRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 468
Query: 361 SYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVT 540
SYLGRPWKE+SRLVIMESTIADFIKPEGYMPWNG+F + TLYYAE+NNRGPGAGTSKRV
Sbjct: 469 SYLGRPWKEYSRLVIMESTIADFIKPEGYMPWNGEFALNTLYYAEFNNRGPGAGTSKRVN 528
Query: 541 WPGFHVIGRKDAEQFTA 591
W GF VIG+K+AEQFTA
Sbjct: 529 WKGFRVIGQKEAEQFTA 545
>sptrnew|CAE02974|CAE02974 OSJNBb0079B02.7 protein.
Length = 971
Score = 313 bits (803), Expect = 8e-85
Identities = 141/197 (71%), Positives = 168/197 (85%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
MGF NTAG E HQAVAL VQGD++ F+NC+F+ +QDTLYVHA RQFFRNC ++GTID+IF
Sbjct: 752 MGFVNTAGPEGHQAVALHVQGDMSVFFNCKFEGYQDTLYVHANRQFFRNCEVTGTIDYIF 811
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 360
GNSAAVFQ+CL+ R+PMDNQ N VTAHGRTDPNM +G+V+Q+CR+VP+Q LFP R +I
Sbjct: 812 GNSAAVFQSCLMTVRKPMDNQANMVTAHGRTDPNMPTGIVLQDCRIVPEQALFPVRLQIA 871
Query: 361 SYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVT 540
SYLGRPWKE++R V+MES I DFIKPEG+ W GD G+KTLYYAEY N GPGAGTSKRVT
Sbjct: 872 SYLGRPWKEYARTVVMESVIGDFIKPEGWSEWMGDVGLKTLYYAEYANTGPGAGTSKRVT 931
Query: 541 WPGFHVIGRKDAEQFTA 591
WPG+ VIG+ +A QFTA
Sbjct: 932 WPGYRVIGQAEATQFTA 948
>sptr|Q7X6B6|Q7X6B6 OSJNBb0079B02.8 protein (OSJNBa0063C18.2 protein).
Length = 971
Score = 313 bits (803), Expect = 8e-85
Identities = 141/197 (71%), Positives = 168/197 (85%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
MGF NTAG E HQAVAL VQGD++ F+NC+F+ +QDTLYVHA RQFFRNC ++GTID+IF
Sbjct: 752 MGFVNTAGPEGHQAVALHVQGDMSVFFNCKFEGYQDTLYVHANRQFFRNCEVTGTIDYIF 811
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 360
GNSAAVFQ+CL+ R+PMDNQ N VTAHGRTDPNM +G+V+Q+CR+VP+Q LFP R +I
Sbjct: 812 GNSAAVFQSCLMTVRKPMDNQANMVTAHGRTDPNMPTGIVLQDCRIVPEQALFPVRLQIA 871
Query: 361 SYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVT 540
SYLGRPWKE++R V+MES I DFIKPEG+ W GD G+KTLYYAEY N GPGAGTSKRVT
Sbjct: 872 SYLGRPWKEYARTVVMESVIGDFIKPEGWSEWMGDVGLKTLYYAEYANTGPGAGTSKRVT 931
Query: 541 WPGFHVIGRKDAEQFTA 591
WPG+ VIG+ +A QFTA
Sbjct: 932 WPGYRVIGQAEATQFTA 948
>sptr|Q9FSQ0|Q9FSQ0 Putative pectin methylesterase.
Length = 717
Score = 313 bits (803), Expect = 8e-85
Identities = 141/197 (71%), Positives = 168/197 (85%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
MGF NTAG E HQAVAL VQGD++ F+NC+F+ +QDTLYVHA RQFFRNC ++GTID+IF
Sbjct: 498 MGFVNTAGPEGHQAVALHVQGDMSVFFNCKFEGYQDTLYVHANRQFFRNCEVTGTIDYIF 557
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 360
GNSAAVFQ+CL+ R+PMDNQ N VTAHGRTDPNM +G+V+Q+CR+VP+Q LFP R +I
Sbjct: 558 GNSAAVFQSCLMTVRKPMDNQANMVTAHGRTDPNMPTGIVLQDCRIVPEQALFPVRLQIA 617
Query: 361 SYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVT 540
SYLGRPWKE++R V+MES I DFIKPEG+ W GD G+KTLYYAEY N GPGAGTSKRVT
Sbjct: 618 SYLGRPWKEYARTVVMESVIGDFIKPEGWSEWMGDVGLKTLYYAEYANTGPGAGTSKRVT 677
Query: 541 WPGFHVIGRKDAEQFTA 591
WPG+ VIG+ +A QFTA
Sbjct: 678 WPGYRVIGQAEATQFTA 694
>sptr|Q9MBB6|Q9MBB6 Pectin methylesterase.
Length = 596
Score = 278 bits (711), Expect = 4e-74
Identities = 123/196 (62%), Positives = 159/196 (81%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
MGF NTAG E+HQAVA+RVQ D A F NCRF+ +QDTLY RQF+R+CVI+GT+DFIF
Sbjct: 380 MGFRNTAGPEKHQAVAIRVQADRAIFLNCRFEGYQDTLYAQTHRQFYRSCVITGTVDFIF 439
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 360
G++ +VFQNCLI R+P++NQQN VTA GR D + +G+V+Q+CR+ PD+ L P + KI
Sbjct: 440 GDATSVFQNCLITVRKPLENQQNIVTAQGRIDGHETTGIVLQSCRIEPDKDLVPVKNKIR 499
Query: 361 SYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVT 540
SYLGRPWKEFSR VIM+STI DFI P G++PW GDFG+KTLYYAEY+N+G GA T+ R+
Sbjct: 500 SYLGRPWKEFSRTVIMDSTIGDFIHPGGWLPWQGDFGLKTLYYAEYSNKGGGAQTNARIK 559
Query: 541 WPGFHVIGRKDAEQFT 588
WPG+H+I +++A +FT
Sbjct: 560 WPGYHIIKKEEAMKFT 575
>sptr|Q9SRX4|Q9SRX4 F22D16.20 protein.
Length = 579
Score = 254 bits (648), Expect = 7e-67
Identities = 112/194 (57%), Positives = 141/194 (72%)
Frame = +1
Query: 7 FHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGN 186
F NTAG E+HQAVALR D + FY+C F+A+QDTLY H+ RQF+R C + GT+DFIFGN
Sbjct: 362 FRNTAGPEKHQAVALRSGADFSIFYSCSFEAYQDTLYTHSLRQFYRECDVYGTVDFIFGN 421
Query: 187 SAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSY 366
+A VFQNC + R+PM NQ N++TA GR+DPN +G IQNC + P L + + +Y
Sbjct: 422 AAVVFQNCNLYPRKPMPNQFNAITAQGRSDPNQNTGTSIQNCTIKPADDLVSSNYTVKTY 481
Query: 367 LGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWP 546
LGRPWKE+SR V M+S I F++P G+ WNGDF + TLYYAEYNN GPG+ T+ RVTWP
Sbjct: 482 LGRPWKEYSRTVYMQSYIDGFVEPVGWREWNGDFALSTLYYAEYNNTGPGSNTTNRVTWP 541
Query: 547 GFHVIGRKDAEQFT 588
G+HVI DA FT
Sbjct: 542 GYHVINSTDAANFT 555
>sptr|O81300|O81300 T14P8.14 protein (AT4G02330 protein)
(Hypothetical protein).
Length = 573
Score = 249 bits (637), Expect = 1e-65
Identities = 110/196 (56%), Positives = 141/196 (71%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
M F NTAG E+HQAVA+R DL+ FY+C F+A+QDTLY H+ RQF+R C I GT+DFIF
Sbjct: 354 MTFRNTAGPEKHQAVAMRSSADLSIFYSCSFEAYQDTLYTHSLRQFYRECDIYGTVDFIF 413
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 360
GN+A VFQ+C + R+PM NQ N++TA GRTDPN +G+ I NC + P L + +
Sbjct: 414 GNAAVVFQDCNLYPRQPMQNQFNAITAQGRTDPNQNTGISIHNCTIKPADDLVSSNYTVK 473
Query: 361 SYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVT 540
+YLGRPWKE+SR V M+S I + ++P G+ WNGDF + TLYYAEYNN G G+ T+ RV
Sbjct: 474 TYLGRPWKEYSRTVFMQSYIDEVVEPVGWREWNGDFALSTLYYAEYNNTGSGSSTTDRVV 533
Query: 541 WPGFHVIGRKDAEQFT 588
WPG+HVI DA FT
Sbjct: 534 WPGYHVINSTDANNFT 549
>sptr|Q42936|Q42936 Pectin methylesterase (EC 3.1.1.11)
(Pectinesterase) (Fragment).
Length = 315
Score = 249 bits (637), Expect = 1e-65
Identities = 112/194 (57%), Positives = 140/194 (72%)
Frame = +1
Query: 7 FHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGN 186
F NTAGA +HQAVALRV D + C+ DAFQDTLY H+ RQF+R+C I+GT+DFIFGN
Sbjct: 100 FQNTAGAAKHQAVALRVGADQSVINRCKIDAFQDTLYTHSLRQFYRDCYITGTVDFIFGN 159
Query: 187 SAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSY 366
+A VFQN I R+P Q+N VTA GR DPN +G IQNC ++P L P + + +Y
Sbjct: 160 AAVVFQNSKIAARKPGSGQKNMVTAQGREDPNQNTGTSIQNCDIIPSSDLAPVKGSVKTY 219
Query: 367 LGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWP 546
LGRPWK +SR V M+S I D I PEG+ W+GDF +KTLYY EY N+GPGAGTSKRV WP
Sbjct: 220 LGRPWKAYSRTVFMQSNIGDHIDPEGWSVWDGDFALKTLYYGEYMNKGPGAGTSKRVKWP 279
Query: 547 GFHVIGRKDAEQFT 588
G+H++ +A +FT
Sbjct: 280 GYHILSAAEATKFT 293
>sptr|Q42935|Q42935 Pectin methylesterase (EC 3.1.1.11)
(Pectinesterase) (Fragment).
Length = 315
Score = 249 bits (637), Expect = 1e-65
Identities = 112/194 (57%), Positives = 140/194 (72%)
Frame = +1
Query: 7 FHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGN 186
F NTAGA +HQAVALRV D + C+ DAFQDTLY H+ RQF+R+C I+GT+DFIFGN
Sbjct: 100 FQNTAGAAKHQAVALRVGADQSVINRCKIDAFQDTLYTHSLRQFYRDCYITGTVDFIFGN 159
Query: 187 SAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSY 366
+A VFQN I R+P Q+N VTA GR DPN +G IQNC ++P L P + + +Y
Sbjct: 160 AAVVFQNSKIAARKPGSGQKNMVTAQGREDPNQNTGTSIQNCDIIPSSDLAPVKGSVKTY 219
Query: 367 LGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWP 546
LGRPWK +SR V M+S I D I PEG+ W+GDF +KTLYY EY N+GPGAGTSKRV WP
Sbjct: 220 LGRPWKAYSRTVFMQSNIGDHIDPEGWSVWDGDFALKTLYYGEYMNKGPGAGTSKRVKWP 279
Query: 547 GFHVIGRKDAEQFT 588
G+H++ +A +FT
Sbjct: 280 GYHILSAAEATKFT 293
>sptr|Q42937|Q42937 Pectin methylesterase (EC 3.1.1.11) (Fragment).
Length = 274
Score = 247 bits (630), Expect = 9e-65
Identities = 111/194 (57%), Positives = 139/194 (71%)
Frame = +1
Query: 7 FHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGN 186
F NTAGA +HQAVALRV + C+ DAFQDTLY H+ RQF+R+C I+GT+DFIFGN
Sbjct: 59 FQNTAGAAKHQAVALRVGAGQSVINRCKIDAFQDTLYTHSLRQFYRDCYITGTVDFIFGN 118
Query: 187 SAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSY 366
+A VFQN I R+P Q+N VTA GR DPN +G IQNC ++P L P + + +Y
Sbjct: 119 AAVVFQNSKIAARKPGSGQKNMVTAQGREDPNQNTGTSIQNCDIIPSSDLAPVKGSVKTY 178
Query: 367 LGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWP 546
LGRPWK +SR V M+S I D I PEG+ W+GDF +KTLYY EY N+GPGAGTSKRV WP
Sbjct: 179 LGRPWKAYSRTVFMQSNIGDHIDPEGWSVWDGDFALKTLYYGEYMNKGPGAGTSKRVKWP 238
Query: 547 GFHVIGRKDAEQFT 588
G+H++ +A +FT
Sbjct: 239 GYHILSAAEATKFT 252
>sptr|Q8RXK7|Q8RXK7 Hypothetical protein.
Length = 573
Score = 246 bits (629), Expect = 1e-64
Identities = 109/196 (55%), Positives = 140/196 (71%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
M F NTAG E+HQAVA+R DL+ FY+C F+A+QDTLY H+ RQF+R C I GT+DFIF
Sbjct: 354 MTFRNTAGPEKHQAVAMRSSADLSIFYSCSFEAYQDTLYTHSLRQFYRECDIYGTVDFIF 413
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 360
GN+A VFQ+C + R+PM NQ N++TA GRTD N +G+ I NC + P L + +
Sbjct: 414 GNAAVVFQDCNLYPRQPMQNQFNAITAQGRTDQNQNTGISIHNCTIKPADDLVSSNYTVK 473
Query: 361 SYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVT 540
+YLGRPWKE+SR V M+S I + ++P G+ WNGDF + TLYYAEYNN G G+ T+ RV
Sbjct: 474 TYLGRPWKEYSRTVFMQSYIDEVVEPVGWREWNGDFALSTLYYAEYNNTGSGSSTTDRVV 533
Query: 541 WPGFHVIGRKDAEQFT 588
WPG+HVI DA FT
Sbjct: 534 WPGYHVINSTDANNFT 549
>sptrnew|CAE76633|CAE76633 Pectin methylesterase (EC 3.1.1.11)
(Fragment).
Length = 426
Score = 246 bits (629), Expect = 1e-64
Identities = 120/195 (61%), Positives = 141/195 (72%), Gaps = 1/195 (0%)
Frame = +1
Query: 7 FHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGN 186
F NTAG +HQAVALRV GDL+AFYNC A+QDTLYVH RQFF NC ISGT+DFIFGN
Sbjct: 210 FQNTAGPSKHQAVALRVGGDLSAFYNCDIIAYQDTLYVHTNRQFFVNCFISGTVDFIFGN 269
Query: 187 SAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSY 366
SA VFQNC I R+P Q+N VTA GR DPN +G+VIQ CR+ + L + P+Y
Sbjct: 270 SAVVFQNCDIHARKPDSGQKNMVTAQGRVDPNQNTGIVIQKCRIGATKDLEGLKGTFPTY 329
Query: 367 LGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWP 546
LGRPWKE+SR VIM+S+I+D I P G+ WNG+F + TL Y EY N GPGAGTSKRV W
Sbjct: 330 LGRPWKEYSRTVIMQSSISDVIDPIGWHEWNGNFALNTLVYREYQNTGPGAGTSKRVNWK 389
Query: 547 GFHVI-GRKDAEQFT 588
GF VI +A+ FT
Sbjct: 390 GFKVITSASEAQTFT 404
>sptr|Q9FJ21|Q9FJ21 Pectin methylesterase (EC 3.1.1.11)
(AT5g49180/K21P3_5) (Pectinesterase).
Length = 571
Score = 245 bits (625), Expect = 3e-64
Identities = 111/196 (56%), Positives = 139/196 (70%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
+GF NTAG E HQAVALRV DLA FYNC+ D +QDTLYVH+ RQFFR+C +SGT+DFIF
Sbjct: 353 IGFENTAGPEGHQAVALRVSADLAVFYNCQIDGYQDTLYVHSHRQFFRDCTVSGTVDFIF 412
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 360
G+ V QNC IV R+PM +Q +TA GR+D +GLV+QNC + + P +
Sbjct: 413 GDGIVVLQNCNIVVRKPMKSQSCMITAQGRSDKRESTGLVLQNCHITGEPAYIPVKSINK 472
Query: 361 SYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVT 540
+YLGRPWKEFSR +IM +TI D I P G++PWNGDF + TLYYAEY N GPG+ ++RV
Sbjct: 473 AYLGRPWKEFSRTIIMGTTIDDVIDPAGWLPWNGDFALNTLYYAEYENNGPGSNQAQRVK 532
Query: 541 WPGFHVIGRKDAEQFT 588
WPG + K A +FT
Sbjct: 533 WPGIKKLSPKQALRFT 548
>sptr|Q9M9W6|Q9M9W6 Putative pectinesterase.
Length = 669
Score = 241 bits (614), Expect = 6e-63
Identities = 105/196 (53%), Positives = 141/196 (71%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
+GF NTAGA +HQAVA+RVQ D + F+NCRFD +QDTLY H+ RQFFR+C ISGTIDF+F
Sbjct: 348 IGFENTAGAIKHQAVAVRVQSDESIFFNCRFDGYQDTLYTHSHRQFFRDCTISGTIDFLF 407
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 360
G++AAVFQNC ++ R+P+ NQ +TAHGR DP +G V Q C + + +
Sbjct: 408 GDAAAVFQNCTLLVRKPLPNQACPITAHGRKDPRESTGFVFQGCTIAGEPDYLAVKETSK 467
Query: 361 SYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVT 540
+YLGRPWKE+SR +IM + I DF++P+G+ PW GDFG+KTL+Y+E N GPG+ + RVT
Sbjct: 468 AYLGRPWKEYSRTIIMNTFIPDFVQPQGWQPWLGDFGLKTLFYSEVQNTGPGSALANRVT 527
Query: 541 WPGFHVIGRKDAEQFT 588
W G + +D +FT
Sbjct: 528 WAGIKTLSEEDILKFT 543
>sptr|Q8GUF8|Q8GUF8 At2g47550/T30B22.15.
Length = 345
Score = 239 bits (610), Expect = 2e-62
Identities = 106/192 (55%), Positives = 135/192 (70%)
Frame = +1
Query: 13 NTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGNSA 192
NTAG + QAVALR GDL+ FY+C F+A+QDTLY H+ RQF+R C + GT+DFIFGN+A
Sbjct: 130 NTAGPTKGQAVALRSGGDLSVFYSCSFEAYQDTLYTHSLRQFYRECDVYGTVDFIFGNAA 189
Query: 193 AVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLG 372
V QNC + R+P Q N VTA GRTDPN +G I C + P L + + +YLG
Sbjct: 190 VVLQNCNLYPRQPRKGQSNEVTAQGRTDPNQNTGTAIHGCTIRPADDLATSNYTVKTYLG 249
Query: 373 RPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWPGF 552
RPWKE+SR V+M++ I F++P G+ W+GDF + TLYYAEYNN GPG+ T+ RVTWPG+
Sbjct: 250 RPWKEYSRTVVMQTYIDGFLEPSGWNAWSGDFALSTLYYAEYNNTGPGSDTTNRVTWPGY 309
Query: 553 HVIGRKDAEQFT 588
HVI DA FT
Sbjct: 310 HVINATDASNFT 321
>sptr|O22256|O22256 Putative pectinesterase.
Length = 560
Score = 239 bits (610), Expect = 2e-62
Identities = 106/192 (55%), Positives = 135/192 (70%)
Frame = +1
Query: 13 NTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGNSA 192
NTAG + QAVALR GDL+ FY+C F+A+QDTLY H+ RQF+R C + GT+DFIFGN+A
Sbjct: 345 NTAGPTKGQAVALRSGGDLSVFYSCSFEAYQDTLYTHSLRQFYRECDVYGTVDFIFGNAA 404
Query: 193 AVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLG 372
V QNC + R+P Q N VTA GRTDPN +G I C + P L + + +YLG
Sbjct: 405 VVLQNCNLYPRQPRKGQSNEVTAQGRTDPNQNTGTAIHGCTIRPADDLATSNYTVKTYLG 464
Query: 373 RPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWPGF 552
RPWKE+SR V+M++ I F++P G+ W+GDF + TLYYAEYNN GPG+ T+ RVTWPG+
Sbjct: 465 RPWKEYSRTVVMQTYIDGFLEPSGWNAWSGDFALSTLYYAEYNNTGPGSDTTNRVTWPGY 524
Query: 553 HVIGRKDAEQFT 588
HVI DA FT
Sbjct: 525 HVINATDASNFT 536
>sptr|Q8GX86|Q8GX86 Putative pectinesterase (At3g05610).
Length = 669
Score = 239 bits (609), Expect = 2e-62
Identities = 104/196 (53%), Positives = 141/196 (71%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
+GF NTAGA +HQAVA+RVQ D + F+NCRFD +Q+TLY H+ RQFFR+C ISGTIDF+F
Sbjct: 348 IGFENTAGAIKHQAVAVRVQSDESIFFNCRFDGYQNTLYTHSHRQFFRDCTISGTIDFLF 407
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 360
G++AAVFQNC ++ R+P+ NQ +TAHGR DP +G V Q C + + +
Sbjct: 408 GDAAAVFQNCTLLVRKPLPNQACPITAHGRKDPRESTGFVFQGCTIAGEPDYLAVKETSK 467
Query: 361 SYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVT 540
+YLGRPWKE+SR +IM + I DF++P+G+ PW GDFG+KTL+Y+E N GPG+ + RVT
Sbjct: 468 AYLGRPWKEYSRTIIMNTFIPDFVQPQGWQPWLGDFGLKTLFYSEVQNTGPGSALANRVT 527
Query: 541 WPGFHVIGRKDAEQFT 588
W G + +D +FT
Sbjct: 528 WAGIKTLSEEDILKFT 543
>sptrnew|BAD08732|BAD08732 Putative pectinesterase.
Length = 664
Score = 238 bits (606), Expect = 5e-62
Identities = 113/198 (57%), Positives = 145/198 (73%), Gaps = 2/198 (1%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
+GF NTA A +HQAVAL VQ D + F NCR + QDTLY H++ QF+RNCVISGT+DFIF
Sbjct: 443 LGFRNTARAAKHQAVALLVQSDKSIFLNCRMEGHQDTLYAHSKAQFYRNCVISGTVDFIF 502
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKL-FPDRFKI 357
G++AAVFQNC+IV RRP+DNQQN TA GR D +G V+Q+ R + L R +
Sbjct: 503 GDAAAVFQNCVIVLRRPLDNQQNIATAQGRADRREATGFVLQHYRFAAESALGDASRPAV 562
Query: 358 PSYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRV 537
SYL RPW+E+SR +IM S I F+ GY+PW+GDFG+KTL+YAEY N+G GA T+ RV
Sbjct: 563 RSYLARPWREYSRTLIMNSDIPAFVDKAGYLPWSGDFGLKTLWYAEYGNKGAGAATAGRV 622
Query: 538 TWPGF-HVIGRKDAEQFT 588
+WPG+ VI +K+A +FT
Sbjct: 623 SWPGYKKVISKKEATKFT 640
>sptr|Q8LQA0|Q8LQA0 Putative pectinesterase.
Length = 557
Score = 237 bits (604), Expect = 9e-62
Identities = 110/198 (55%), Positives = 138/198 (69%), Gaps = 2/198 (1%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
M F NTAG +HQAVALR DL+ FY C F+A+QDTLY H+ RQF+R C + GT+D++F
Sbjct: 337 MTFRNTAGPAKHQAVALRCGADLSTFYQCSFEAYQDTLYTHSLRQFYRACDVYGTVDYVF 396
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPD-RFKI 357
GN+A VFQ+C + R PM Q N+VTA GRTDPN +G IQ C +V L + F
Sbjct: 397 GNAAVVFQDCTLYNRLPMQGQSNTVTAQGRTDPNQNTGTTIQGCAIVAAPDLAANTAFAT 456
Query: 358 PSYLGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRV 537
+YLGRPWK +SR VIM+S + I P G+MPW+GD+ + TLYYAEYNN G GA TS+RV
Sbjct: 457 TNYLGRPWKLYSRTVIMQSVVGGLIDPAGWMPWDGDYALSTLYYAEYNNSGAGADTSRRV 516
Query: 538 TWPGFHVI-GRKDAEQFT 588
TWPG+HV+ DA FT
Sbjct: 517 TWPGYHVLNSTADAGNFT 534
>sw|O04886|PME1_CITSI Pectinesterase 1.1 precursor (EC 3.1.1.11)
(Pectin methylesterase) (PE).
Length = 584
Score = 236 bits (603), Expect = 1e-61
Identities = 114/195 (58%), Positives = 140/195 (71%), Gaps = 1/195 (0%)
Frame = +1
Query: 7 FHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGN 186
F NTAG +HQAVALRV DL+AFYNC A+QDTLYVH+ RQFF NC+I+GT+DFIFGN
Sbjct: 368 FQNTAGPSKHQAVALRVGADLSAFYNCDMLAYQDTLYVHSNRQFFVNCLIAGTVDFIFGN 427
Query: 187 SAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSY 366
+AAV QNC I R+P Q+N VTA GRTDPN +G+VIQ R+ L P + P+Y
Sbjct: 428 AAAVLQNCDIHARKPNSGQKNMVTAQGRTDPNQNTGIVIQKSRIGATSDLKPVQGSFPTY 487
Query: 367 LGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWP 546
LGRPWKE+SR VIM+S+I D I P G+ W+G+F + TL+Y E+ N G GAGTS RV W
Sbjct: 488 LGRPWKEYSRTVIMQSSITDLIHPAGWHEWDGNFALNTLFYGEHQNSGAGAGTSGRVKWK 547
Query: 547 GFHVI-GRKDAEQFT 588
GF VI +A+ FT
Sbjct: 548 GFRVITSATEAQAFT 562
>sptr|Q93Z62|Q93Z62 At2g47550/T30B22.15.
Length = 345
Score = 236 bits (601), Expect = 2e-61
Identities = 105/192 (54%), Positives = 134/192 (69%)
Frame = +1
Query: 13 NTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGNSA 192
NTAG + QAVALR GDL+ FY+C F+A+QDTLY H+ RQF+R C + GT+DFIFGN+A
Sbjct: 130 NTAGPTKGQAVALRSGGDLSVFYSCSFEAYQDTLYTHSLRQFYRECDVYGTVDFIFGNAA 189
Query: 193 AVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLG 372
V QNC + R+P Q N VTA GRT PN +G I C + P L + + +YLG
Sbjct: 190 VVLQNCNLYPRQPRKGQSNEVTAQGRTYPNQNTGTAIHGCTIRPADDLATSNYTVKTYLG 249
Query: 373 RPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWPGF 552
RPWKE+SR V+M++ I F++P G+ W+GDF + TLYYAEYNN GPG+ T+ RVTWPG+
Sbjct: 250 RPWKEYSRTVVMQTYIDGFLEPSGWNAWSGDFALSTLYYAEYNNTGPGSDTTNRVTWPGY 309
Query: 553 HVIGRKDAEQFT 588
HVI DA FT
Sbjct: 310 HVINATDASNFT 321
>sptr|Q9FY03|Q9FY03 Putative pectin methylesterase precursor (EC
3.1.1.11).
Length = 579
Score = 235 bits (599), Expect = 4e-61
Identities = 110/185 (59%), Positives = 134/185 (72%)
Frame = +1
Query: 7 FHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGN 186
F NTAG +HQAVALRV DL+AFY C A+QDTLYVH+ RQFF NC ++GT+DFIFGN
Sbjct: 363 FENTAGPSKHQAVALRVGSDLSAFYECDMLAYQDTLYVHSNRQFFINCFVAGTVDFIFGN 422
Query: 187 SAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSY 366
+AAVFQ+C RRP Q+N VTA GRTDPN +G+VIQ R+ L P + P+Y
Sbjct: 423 AAAVFQDCDYHARRPDSGQKNMVTAQGRTDPNQNTGIVIQKSRIGATSDLLPVQSSFPTY 482
Query: 367 LGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWP 546
LGRPWKE+SR VIM+S+I D I+P G+ W+G F + TL+YAEY N G GAGTS RV W
Sbjct: 483 LGRPWKEYSRTVIMQSSITDVIQPAGWHEWSGSFALSTLFYAEYQNSGAGAGTSSRVKWE 542
Query: 547 GFHVI 561
G+ VI
Sbjct: 543 GYKVI 547
>sw|P83218|PME_DAUCA Pectinesterase (EC 3.1.1.11) (Pectin
methylesterase) (PE).
Length = 319
Score = 235 bits (599), Expect = 4e-61
Identities = 113/195 (57%), Positives = 139/195 (71%), Gaps = 1/195 (0%)
Frame = +1
Query: 7 FHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGN 186
F NTAGA +HQAVALRV DL+AFY C A+QD+LYVH+ RQFF NC I+GT+DFIFGN
Sbjct: 103 FQNTAGAAKHQAVALRVGSDLSAFYRCDILAYQDSLYVHSNRQFFINCFIAGTVDFIFGN 162
Query: 187 SAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSY 366
+A V Q+C I RRP Q+N VTA GRTDPN +G+VIQ R+ L P + P+Y
Sbjct: 163 AAVVLQDCDIHARRPGSGQKNMVTAQGRTDPNQNTGIVIQKSRIGATSDLQPVQSSFPTY 222
Query: 367 LGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWP 546
LGRPWKE+SR V+M+S+I + I P G+ PW+G+F + TLYY EY N G GA TS RVTW
Sbjct: 223 LGRPWKEYSRTVVMQSSITNVINPAGWFPWDGNFALDTLYYGEYQNTGAGAATSGRVTWK 282
Query: 547 GFHVI-GRKDAEQFT 588
GF VI +A+ FT
Sbjct: 283 GFKVITSSTEAQGFT 297
>sptr|Q9SEE7|Q9SEE7 Pectin methyl esterase (EC 3.1.1.11)
(Pectinesterase) (Pectin methylesterase) (Fragment).
Length = 530
Score = 235 bits (599), Expect = 4e-61
Identities = 107/183 (58%), Positives = 134/183 (73%)
Frame = +1
Query: 13 NTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGNSA 192
NTAG E+HQAVALRV GD++ C DA+QDTLY H++RQF+R+ +SGTIDFIFGN+A
Sbjct: 314 NTAGPEKHQAVALRVGGDMSVINRCPIDAYQDTLYAHSQRQFYRDSYVSGTIDFIFGNAA 373
Query: 193 AVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLG 372
VFQ C +V R+P NQ+N VTA GRTDPN +G IQ C ++ L P + +YLG
Sbjct: 374 VVFQKCQLVARKPSKNQKNMVTAQGRTDPNQATGTSIQFCDIIASPDLEPVVKEFKTYLG 433
Query: 373 RPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWPGF 552
RPWKE+SR V+M+S + I P G+ W+G+F +KTLYY EY N GPGAGTSKRV WPG+
Sbjct: 434 RPWKEYSRTVVMQSYLGGLIDPAGWAEWSGEFALKTLYYGEYMNNGPGAGTSKRVKWPGY 493
Query: 553 HVI 561
HVI
Sbjct: 494 HVI 496
>sptr|Q84UI7|Q84UI7 Pectin methylesterase.
Length = 553
Score = 234 bits (597), Expect = 6e-61
Identities = 111/198 (56%), Positives = 138/198 (69%), Gaps = 2/198 (1%)
Frame = +1
Query: 1 MGFHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIF 180
+GF NTAG E+HQAVALRV D + CR DAFQDTLY H+ RQF+R+C I+GTIDFIF
Sbjct: 334 IGFKNTAGPEKHQAVALRVGSDQSVINRCRIDAFQDTLYAHSNRQFYRDCFITGTIDFIF 393
Query: 181 GNSAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIP 360
GN+AAVFQ +V R+PM NQ+N VTA GR DPN + IQ C ++P L P I
Sbjct: 394 GNAAAVFQKSKLVARKPMSNQKNMVTAQGRLDPNQNTATSIQQCDIIPSTDLKPVLGSIK 453
Query: 361 SYLGRPWKEFSRLVIMESTIADFIKPEGYMPWN--GDFGIKTLYYAEYNNRGPGAGTSKR 534
+YLGRPWK +SR V+M+S I + I P G+ W+ +KTLYY EY N GPGAGT+KR
Sbjct: 454 TYLGRPWKPYSRTVVMQSPIGNHIDPTGWAEWDDASKAFLKTLYYGEYLNSGPGAGTAKR 513
Query: 535 VTWPGFHVIGRKDAEQFT 588
V WPG+HV+ +A +FT
Sbjct: 514 VNWPGYHVLNTAEATKFT 531
>sptr|Q7XAF8|Q7XAF8 Papillar cell-specific pectin
methylesterase-like protein.
Length = 562
Score = 233 bits (595), Expect = 1e-60
Identities = 103/192 (53%), Positives = 133/192 (69%)
Frame = +1
Query: 13 NTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGNSA 192
NTAG + QAVALR GD + FY+C F+A+QDTLY H+ RQF+R C + GT+DFIFGN+A
Sbjct: 347 NTAGPTKGQAVALRSGGDFSVFYSCSFEAYQDTLYTHSLRQFYRECDVYGTVDFIFGNAA 406
Query: 193 AVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSYLG 372
V Q C + R+P Q N VTA GRTDPN +G V+ C + P L + + +YLG
Sbjct: 407 VVLQKCNLYPRQPRQGQANEVTAQGRTDPNQNTGTVLHGCTIRPADDLASSNYTVKTYLG 466
Query: 373 RPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWPGF 552
RPWKE+SR V+M++ I F+ P G+ W+G+F + TLYYAEYNN GPG+ T+ RVTWPG+
Sbjct: 467 RPWKEYSRTVVMQTYIDGFLDPTGWNAWSGNFALSTLYYAEYNNTGPGSSTTNRVTWPGY 526
Query: 553 HVIGRKDAEQFT 588
HVI DA FT
Sbjct: 527 HVINATDASNFT 538
>sptr|Q9LEB0|Q9LEB0 Pectin methylesterase (EC 3.1.1.11)
(Pectinesterase).
Length = 579
Score = 233 bits (594), Expect = 1e-60
Identities = 112/195 (57%), Positives = 139/195 (71%), Gaps = 1/195 (0%)
Frame = +1
Query: 7 FHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGN 186
F NTAGA +HQAVALRV DL+AFY C A+QD+LYVH+ RQ+F C+I+GT+DFIFGN
Sbjct: 363 FQNTAGAAKHQAVALRVGSDLSAFYRCDILAYQDSLYVHSNRQYFVQCLIAGTVDFIFGN 422
Query: 187 SAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSY 366
+AAV QNC I RRP Q+N VTA GR+DPN +G+VIQ CR+ L P + P+Y
Sbjct: 423 AAAVLQNCDIHARRPGSGQKNMVTAQGRSDPNQNTGIVIQKCRIGATSDLRPVQKSFPTY 482
Query: 367 LGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWP 546
LGRPWKE+SR VIM+S+I D I G+ WNG+F + TL+Y EY N G GAGTS RV W
Sbjct: 483 LGRPWKEYSRTVIMQSSITDVINSAGWHEWNGNFALNTLFYGEYQNTGAGAGTSGRVKWK 542
Query: 547 GFHVI-GRKDAEQFT 588
GF VI +A+ +T
Sbjct: 543 GFKVITSATEAQAYT 557
>sptr|Q9SEE6|Q9SEE6 Pectin methyl esterase (EC 3.1.1.11)
(Pectinesterase) (Pectin methylesterase).
Length = 576
Score = 233 bits (593), Expect = 2e-60
Identities = 112/195 (57%), Positives = 138/195 (70%), Gaps = 1/195 (0%)
Frame = +1
Query: 7 FHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGN 186
F NTAGA +HQAVALRV DL+AFY C A+QDTLYVH+ RQFF C+++GT+DFIFGN
Sbjct: 360 FQNTAGASKHQAVALRVGSDLSAFYKCDILAYQDTLYVHSNRQFFVQCLVAGTVDFIFGN 419
Query: 187 SAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSY 366
AAV Q+C I RRP Q+N VTA GRTDPN +G+VIQ CR+ L P + P+Y
Sbjct: 420 GAAVLQDCDIHARRPGSGQKNMVTAQGRTDPNQNTGIVIQKCRIGATSDLRPVQKSFPTY 479
Query: 367 LGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWP 546
LGRPWKE+SR VIM+S+I D I+P G+ WNG+F + TL+Y EY N G GA TS RV W
Sbjct: 480 LGRPWKEYSRTVIMQSSITDVIQPAGWHEWNGNFALNTLFYGEYANTGAGAATSGRVKWK 539
Query: 547 GFHVI-GRKDAEQFT 588
G VI +A+ +T
Sbjct: 540 GHKVITSSTEAQAYT 554
>sptr|Q9M5J0|Q9M5J0 Pectin methylesterase isoform alpha (EC
3.1.1.11) (Fragment).
Length = 277
Score = 233 bits (593), Expect = 2e-60
Identities = 108/195 (55%), Positives = 136/195 (69%), Gaps = 1/195 (0%)
Frame = +1
Query: 7 FHNTAGAERHQAVALRVQGDLAAFYNCRFDAFQDTLYVHARRQFFRNCVISGTIDFIFGN 186
F N+AG +HQAVALR D +AFY C F A+QDTLYVH+ RQF+R C + GT+DFIFGN
Sbjct: 60 FENSAGPSKHQAVALRNGADFSAFYQCSFVAYQDTLYVHSLRQFYRECDVYGTVDFIFGN 119
Query: 187 SAAVFQNCLIVTRRPMDNQQNSVTAHGRTDPNMKSGLVIQNCRLVPDQKLFPDRFKIPSY 366
+AAV QNC + R+P NQ+N TA GR DPN +G+ I NC++ L P + + +Y
Sbjct: 120 AAAVLQNCNLYARKPNKNQRNLFTAQGREDPNQSTGISIINCKVAAAADLIPVKSEFRNY 179
Query: 367 LGRPWKEFSRLVIMESTIADFIKPEGYMPWNGDFGIKTLYYAEYNNRGPGAGTSKRVTWP 546
LGRPWK +SR V + S + D I+P G++ WNG F + TLYY EYNNRGPGA TS RVTWP
Sbjct: 180 LGRPWKMYSRTVFLNSLMEDLIEPAGWLEWNGTFALDTLYYGEYNNRGPGANTSGRVTWP 239
Query: 547 GFHVI-GRKDAEQFT 588
G+ VI +A QFT
Sbjct: 240 GYRVITNSTEASQFT 254
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 391,603,367
Number of Sequences: 1395590
Number of extensions: 8940303
Number of successful extensions: 46314
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 41227
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 45356
length of database: 442,889,342
effective HSP length: 117
effective length of database: 279,605,312
effective search space used: 22368424960
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

