BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
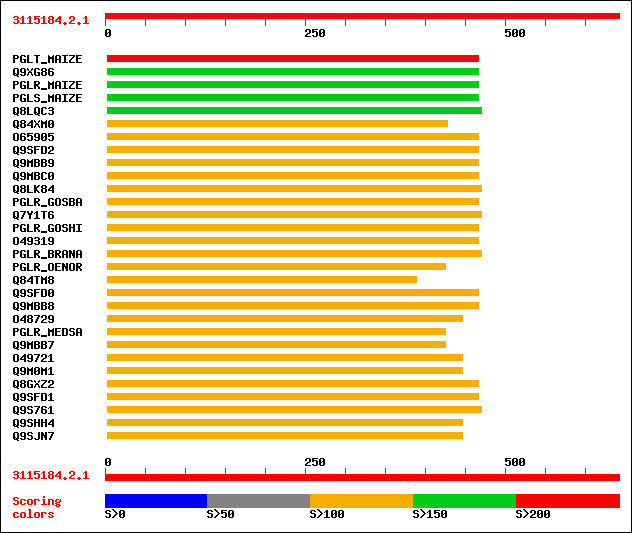
Query= 3115184.2.1
(642 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sw|P35339|PGLT_MAIZE Exopolygalacturonase precursor (EC 3.2.1.67... 201 7e-51
sptr|Q9XG86|Q9XG86 Polygalacturonase (Fragment). 187 8e-47
sw|P26216|PGLR_MAIZE Exopolygalacturonase precursor (EC 3.2.1.67... 186 2e-46
sw|P35338|PGLS_MAIZE Exopolygalacturonase precursor (EC 3.2.1.67... 185 4e-46
sptr|Q8LQC3|Q8LQC3 Putative exopolygalacturorase. 179 2e-44
sptr|Q84XM0|Q84XM0 Putative pollen polygalacturonase. 150 1e-35
sptr|O65905|O65905 Polygalacturonase. 146 3e-34
sptr|Q9SFD2|Q9SFD2 Putative polygalacturonase (Polygalacturonase... 146 3e-34
sptr|Q9MBB9|Q9MBB9 Polygalacturonase. 142 3e-33
sptr|Q9MBC0|Q9MBC0 Polygalacturonase. 142 3e-33
sptr|Q8LK84|Q8LK84 Putative pollen polygalacturonase. 142 3e-33
sw|Q39766|PGLR_GOSBA Polygalacturonase precursor (EC 3.2.1.15) (... 142 5e-33
sptr|Q7Y1T6|Q7Y1T6 Polygalacturonase. 142 5e-33
sw|Q39786|PGLR_GOSHI Polygalacturonase precursor (EC 3.2.1.15) (... 141 8e-33
sptr|O49319|O49319 Putative polygalacturonase. 140 1e-32
sw|P35337|PGLR_BRANA Polygalacturonase precursor (EC 3.2.1.15) (... 140 1e-32
sw|P24548|PGLR_OENOR Exopolygalacturonase (EC 3.2.1.67) (ExoPG) ... 140 2e-32
sptr|Q84TM8|Q84TM8 Polygalactorunase PG11 precursor (EC 3.2.1.15). 139 2e-32
sptr|Q9SFD0|Q9SFD0 Putative polygalacturonase. 139 3e-32
sptr|Q9MBB8|Q9MBB8 Polygalacturonase. 139 4e-32
sptr|O48729|O48729 Putative polygalacturonase. 137 1e-31
sw|Q40312|PGLR_MEDSA Polygalacturonase precursor (EC 3.2.1.15) (... 137 1e-31
sptr|Q9MBB7|Q9MBB7 Polygalacturonase. 137 2e-31
sptr|O49721|O49721 Polygalacturonase-like protein (Fragment). 136 2e-31
sptr|Q9M0M1|Q9M0M1 Polygalacturonase-like protein. 136 2e-31
sptr|Q8GXZ2|Q8GXZ2 Putative polygalacturonase (At3g07830). 135 5e-31
sptr|Q9SFD1|Q9SFD1 Putative polygalacturonase. 135 5e-31
sptr|Q9S761|Q9S761 Putative polygalacturonase. 134 8e-31
sptr|Q9SHH4|Q9SHH4 Similar to polygalacturonase. 134 1e-30
sptr|Q9SJN7|Q9SJN7 Putative polygalacturonase. 133 2e-30
>sw|P35339|PGLT_MAIZE Exopolygalacturonase precursor (EC 3.2.1.67)
(ExoPG) (Pectinase) (Galacturan
1,4-alpha-galacturonidase).
Length = 410
Score = 201 bits (511), Expect = 7e-51
Identities = 94/155 (60%), Positives = 118/155 (76%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGIS+GSLGRYKDEKDV D+ V CTL T NG+RIK+YED+ S L A+K Y+++ M+
Sbjct: 255 GHGISIGSLGRYKDEKDVTDINVKDCTLKKTANGVRIKAYEDAASVLTASKIHYENIKME 314
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
+ YPIIID KYCPN +C +GASKV V DV FKNI GTS+TPEA+ L C +PC GV
Sbjct: 315 DSGYPIIIDMKYCPNKLCTANGASKVTVKDVTFKNITGTSSTPEAVNLLCTAKIPCTGVT 374
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
+ +V+IKY+ ++NKTM+VCKNA G + G KELAC
Sbjct: 375 MDDVNIKYSGTNNKTMAVCKNAKGSAKGCLKELAC 409
>sptr|Q9XG86|Q9XG86 Polygalacturonase (Fragment).
Length = 394
Score = 187 bits (476), Expect = 8e-47
Identities = 90/156 (57%), Positives = 118/156 (75%), Gaps = 1/156 (0%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRYKDEKDV D+ V C L +TNGLRIKSYED+KS L A+K Y++V M+
Sbjct: 237 GHGISVGSLGRYKDEKDVTDITVKNCVLKKSTNGLRIKSYEDAKSPLTASKLTYENVKME 296
Query: 183 NVSYPIIIDQKYCPNNICVKSG-ASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGV 359
+V YPIIIDQKYCPN IC G +++V V DV F+NI GTS+TPEA++L C++ PC GV
Sbjct: 297 DVGYPIIIDQKYCPNKICTSKGDSARVTVKDVTFRNITGTSSTPEAVSLLCSDKQPCNGV 356
Query: 360 QLINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
+ +V I+Y+ ++NKTM+VC NA + G+++ C
Sbjct: 357 TMNDVKIEYSGTNNKTMAVCTNAKVTAKGVSEANTC 392
>sw|P26216|PGLR_MAIZE Exopolygalacturonase precursor (EC 3.2.1.67)
(ExoPG) (Pectinase) (Galacturan
1,4-alpha-galacturonidase).
Length = 410
Score = 186 bits (473), Expect = 2e-46
Identities = 86/155 (55%), Positives = 115/155 (74%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGIS+GSLGRYKDEKDV D+ V CTL T G+RIK+YED+ S L +K Y+++ M+
Sbjct: 255 GHGISIGSLGRYKDEKDVTDINVKDCTLKKTMFGVRIKAYEDAASVLTVSKIHYENIKME 314
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
+ + PI ID KYCPN +C +GASKV V DV FKNI GTS+TPEA++L C +PC GV
Sbjct: 315 DSANPIFIDMKYCPNKLCTANGASKVTVKDVTFKNITGTSSTPEAVSLLCTAKVPCTGVT 374
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
+ +V+++Y+ ++NKTM++C NA G + G KELAC
Sbjct: 375 MDDVNVEYSGTNNKTMAICTNAKGSTKGCLKELAC 409
>sw|P35338|PGLS_MAIZE Exopolygalacturonase precursor (EC 3.2.1.67)
(ExoPG) (Pectinase) (Galacturan
1,4-alpha-galacturonidase).
Length = 410
Score = 185 bits (470), Expect = 4e-46
Identities = 86/155 (55%), Positives = 114/155 (73%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGIS+GSLGRYKDEKDV D+ V CTL T G+RIK+YED+ S L +K Y+++ M+
Sbjct: 255 GHGISIGSLGRYKDEKDVTDINVKDCTLKKTMFGVRIKAYEDAASVLTVSKIHYENIKME 314
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
+ + PI ID KYCPN +C +GASKV V DV FKNI GTS+TPEAI+L C +PC G
Sbjct: 315 DSANPIFIDMKYCPNKLCTANGASKVTVKDVTFKNITGTSSTPEAISLLCTAKVPCTGAT 374
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
+ +V+++Y+ ++NKTM++C NA G + G KELAC
Sbjct: 375 MDDVNVEYSGTNNKTMAICTNAKGSTKGCLKELAC 409
>sptr|Q8LQC3|Q8LQC3 Putative exopolygalacturorase.
Length = 403
Score = 179 bits (455), Expect = 2e-44
Identities = 84/157 (53%), Positives = 115/157 (73%), Gaps = 1/157 (0%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
G GISVG LGR+KDEKDV DV V C L T+NG+RIKSYED S + A++ ++++ MD
Sbjct: 247 GQGISVGCLGRFKDEKDVTDVTVRDCVLRNTSNGVRIKSYEDVLSPITASRLTFENIRMD 306
Query: 183 NVSYPIIIDQKYCPNNIC-VKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGV 359
V+ P+I+DQKYCP C K G+ V + +V F+NI GTSNTPEA++L C++ LPC G+
Sbjct: 307 GVANPVIVDQKYCPEKDCPEKKGSKTVTIKNVTFRNITGTSNTPEAVSLLCSDQLPCSGM 366
Query: 360 QLINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELACV 470
+L++V++KY+ DNKTM+VC NA G S G + LAC+
Sbjct: 367 ELLDVNLKYDGKDNKTMAVCTNAKGISKGSLQALACL 403
>sptr|Q84XM0|Q84XM0 Putative pollen polygalacturonase.
Length = 382
Score = 150 bits (379), Expect = 1e-35
Identities = 72/142 (50%), Positives = 98/142 (69%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGIS+GSLGRY +E+ V V V C L T NG+RIKS+ K +A+ ++D+TM+
Sbjct: 235 GHGISIGSLGRYDNEQPVRGVLVKNCILKNTDNGVRIKSWPAMKGG-EASDIHFEDITME 293
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NV+ P+IIDQ+YCP N C K SKV +++V FKNI GT+NTPEA+ + C++ LPC+ VQ
Sbjct: 294 NVTNPVIIDQEYCPWNQCTKDAPSKVKISNVSFKNIKGTTNTPEAVKIICSSALPCEQVQ 353
Query: 363 LINVDIKYNRSDNKTMSVCKNA 428
L +D+KY + S CKNA
Sbjct: 354 LNGIDLKYTGTQGPAKSECKNA 375
>sptr|O65905|O65905 Polygalacturonase.
Length = 391
Score = 146 bits (368), Expect = 3e-34
Identities = 71/155 (45%), Positives = 97/155 (62%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY E+DV +KV CTL T NGLRIK++ + S A+ ++D+ +
Sbjct: 232 GHGISVGSLGRYGHEQDVSGIKVINCTLQETDNGLRIKTWPSAACSTTASDIHFEDIILK 291
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
+VS PI+IDQ+YCP N C K AS + + ++ FKNI GTS +A+ L C+ PCQ V+
Sbjct: 292 DVSNPILIDQEYCPWNQCNKQKASTIKLVNISFKNIRGTSGNKDAVKLLCSKGYPCQNVE 351
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
+ ++DIKYN +D C N K +G AC
Sbjct: 352 IGDIDIKYNGADGPATFHCSNVSPKILGSQSPKAC 386
>sptr|Q9SFD2|Q9SFD2 Putative polygalacturonase (Polygalacturonase
PGA3).
Length = 391
Score = 146 bits (368), Expect = 3e-34
Identities = 71/155 (45%), Positives = 97/155 (62%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY E+DV +KV CTL T NGLRIK++ + S A+ ++D+ +
Sbjct: 232 GHGISVGSLGRYGHEQDVSGIKVINCTLQETDNGLRIKTWPSAACSTTASDIHFEDIILK 291
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
+VS PI+IDQ+YCP N C K AS + + ++ FKNI GTS +A+ L C+ PCQ V+
Sbjct: 292 DVSNPILIDQEYCPWNQCNKQKASTIKLVNISFKNIRGTSGNKDAVKLLCSKGYPCQNVE 351
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
+ ++DIKYN +D C N K +G AC
Sbjct: 352 IGDIDIKYNGADGPATFHCSNVSPKILGSQSPKAC 386
>sptr|Q9MBB9|Q9MBB9 Polygalacturonase.
Length = 393
Score = 142 bits (359), Expect = 3e-33
Identities = 69/155 (44%), Positives = 97/155 (62%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY +EK V + V CT++ T NG+RIKS+ D + A+ ++D+ M+
Sbjct: 240 GHGISVGSLGRYPNEKPVSGIFVKNCTISNTANGVRIKSWPDLYGGV-ASNMHFEDIVMN 298
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NV PI++DQ YCP N C SKV ++DV FKNI GTS TP + L C++ +PC+ V+
Sbjct: 299 NVQNPILLDQVYCPWNQCSLKAPSKVKISDVSFKNIRGTSATPVVVKLACSSGIPCEKVE 358
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
L N+++ Y+ S+ S C N K G+ C
Sbjct: 359 LANINLLYSGSEGPAKSQCSNVKPKISGIMSASGC 393
>sptr|Q9MBC0|Q9MBC0 Polygalacturonase.
Length = 393
Score = 142 bits (359), Expect = 3e-33
Identities = 69/155 (44%), Positives = 97/155 (62%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY +EK V + V CT++ T NG+RIKS+ D + A+ ++D+ M+
Sbjct: 240 GHGISVGSLGRYPNEKPVSGIFVKNCTISNTANGVRIKSWPDLYGGV-ASNMHFEDIVMN 298
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NV PI++DQ YCP N C SKV ++DV FKNI GTS TP + L C++ +PC+ V+
Sbjct: 299 NVQNPILLDQVYCPWNQCSLKAPSKVKISDVSFKNIRGTSATPVVVKLACSSGIPCEKVE 358
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
L N+++ Y+ S+ S C N K G+ C
Sbjct: 359 LANINLVYSGSEGPAKSQCSNVKPKISGIMSASGC 393
>sptr|Q8LK84|Q8LK84 Putative pollen polygalacturonase.
Length = 397
Score = 142 bits (359), Expect = 3e-33
Identities = 68/156 (43%), Positives = 98/156 (62%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY E+DV D+KV CTL GT NGLRIK++ + + A ++++ ++
Sbjct: 232 GHGISVGSLGRYGWEQDVNDIKVINCTLEGTDNGLRIKTWPSAACTTTAAGIHFENIILN 291
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
VS PI+IDQ+YCP N C KS S + + D+ F+NI GTS +A+ L C+ PC+ V+
Sbjct: 292 KVSNPILIDQEYCPWNQCNKSKPSTIKLVDITFRNIRGTSGNKDAVKLLCSKGHPCENVE 351
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELACV 470
+ +++I+Y +D C N K +G ACV
Sbjct: 352 IGDINIEYTGADGPPTFECTNVTPKLVGTQNPKACV 387
>sw|Q39766|PGLR_GOSBA Polygalacturonase precursor (EC 3.2.1.15) (PG)
(Pectinase).
Length = 407
Score = 142 bits (357), Expect = 5e-33
Identities = 68/155 (43%), Positives = 105/155 (67%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGIS+GSLG++++E+ VE +K++ CT+ T+NG RIK++ ++ ++D+TM+
Sbjct: 243 GHGISIGSLGKFQNEEPVEGIKISNCTITNTSNGARIKTWPGEHGGA-VSEIHFEDITMN 301
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NVS PI+IDQ+YCP N C K+ SKV ++++ FKNI GTS PEAI C+ + PCQ V+
Sbjct: 302 NVSSPILIDQQYCPWNKCKKNEESKVKLSNISFKNIRGTSALPEAIKFICSGSSPCQNVE 361
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
L ++DI++N ++ T S C N +IG + C
Sbjct: 362 LADIDIQHNGAEPAT-SQCLNVKPITIGKLNPIPC 395
>sptr|Q7Y1T6|Q7Y1T6 Polygalacturonase.
Length = 397
Score = 142 bits (357), Expect = 5e-33
Identities = 68/156 (43%), Positives = 97/156 (62%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY E+DV D+KV CTL GT NGLRIK++ + + A ++++ ++
Sbjct: 232 GHGISVGSLGRYGWEQDVNDIKVINCTLEGTDNGLRIKTWPSAACTTTAAGIHFENIILN 291
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
VS PI+IDQ+YCP N C KS S + + D+ F+NI GTS +A+ L C+ PC+ V+
Sbjct: 292 KVSNPILIDQEYCPWNQCNKSKPSTIKLVDITFRNIRGTSGNKDAVKLLCSKGHPCENVE 351
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELACV 470
+ +++I+Y D C N K +G ACV
Sbjct: 352 IGDINIEYTGPDGPPTFECTNVTPKLVGTQNPKACV 387
>sw|Q39786|PGLR_GOSHI Polygalacturonase precursor (EC 3.2.1.15) (PG)
(Pectinase).
Length = 407
Score = 141 bits (355), Expect = 8e-33
Identities = 68/155 (43%), Positives = 104/155 (67%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGIS+GSLG++++E+ VE +K++ CT+ T+NG RIK++ ++ ++D+TM+
Sbjct: 243 GHGISIGSLGKFQNEEPVEGIKISNCTITNTSNGARIKTWPGEHGGA-VSEIHFEDITMN 301
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NVS PI+IDQ+YCP N C K+ SKV ++++ FKNI GTS PEAI C+ + PCQ V+
Sbjct: 302 NVSSPILIDQQYCPWNKCKKNEESKVKLSNISFKNIRGTSALPEAIKFICSGSSPCQNVE 361
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
L ++DIK+N ++ T S C N + G + C
Sbjct: 362 LADIDIKHNGAEPAT-SQCLNVKPITSGKLNPIPC 395
>sptr|O49319|O49319 Putative polygalacturonase.
Length = 664
Score = 140 bits (354), Expect = 1e-32
Identities = 69/157 (43%), Positives = 98/157 (62%), Gaps = 2/157 (1%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRYK+EK+V+ + V + GTT+GLRIK++ S S + + FLY+++ M
Sbjct: 246 GHGISVGSLGRYKEEKNVQGLTVRNSIINGTTDGLRIKTWAKSVSQISVSNFLYENIQMI 305
Query: 183 NVSYPIIIDQKYCPNNICVKSG--ASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQG 356
NV PI+IDQ+YCP+ C G AS V + DV + NI GTS + EA+ + C+ PCQ
Sbjct: 306 NVGNPIVIDQQYCPHGQCDSPGKYASHVQIKDVKYNNIWGTSTSKEALKMQCSKTFPCQD 365
Query: 357 VQLINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
V+L N+++ Y D ++C+N G G C
Sbjct: 366 VELSNINLHYVGRDGLVTALCENVGGSIRGKIVPANC 402
>sw|P35337|PGLR_BRANA Polygalacturonase precursor (EC 3.2.1.15) (PG)
(Pectinase).
Length = 397
Score = 140 bits (354), Expect = 1e-32
Identities = 67/156 (42%), Positives = 97/156 (62%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY E+DV D+ V CTL GT+NGLRIK++ + + A ++D+ ++
Sbjct: 232 GHGISVGSLGRYGWEQDVTDITVKNCTLEGTSNGLRIKTWPSAACTTTAAGIHFEDIILN 291
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
VS PI+IDQ+YCP N C K+ S + + D+ F+NI GTS +A+ L C+ PC+ V+
Sbjct: 292 KVSNPILIDQEYCPWNQCNKNKPSTIKLVDITFRNIRGTSENKDAVKLLCSKGHPCENVE 351
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELACV 470
+ +++I+Y D C N K +G ACV
Sbjct: 352 IGDINIEYTGPDGPPTFECTNVTPKLVGAQNPKACV 387
>sw|P24548|PGLR_OENOR Exopolygalacturonase (EC 3.2.1.67) (ExoPG)
(Pectinase) (Galacturan 1,4-alpha-galacturonidase)
(Fragment).
Length = 362
Score = 140 bits (352), Expect = 2e-32
Identities = 66/141 (46%), Positives = 95/141 (67%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRYK+E+ V + V CT+ G+ NG+RIK++ S+ +A++ +QD+TM+
Sbjct: 201 GHGISVGSLGRYKNEESVVGIYVKNCTITGSQNGVRIKTWPKSEPG-EASEMHFQDITMN 259
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
+V PI+IDQ YCP N C S V ++ + FKNI GTS T EA+ L C+ + PC GV+
Sbjct: 260 SVGTPILIDQGYCPYNQCTAEVPSSVKLSKISFKNIKGTSTTKEAVKLVCSKSFPCNGVE 319
Query: 363 LINVDIKYNRSDNKTMSVCKN 425
L ++D+ Y+ SVC+N
Sbjct: 320 LADIDLTYSGKGGPATSVCEN 340
>sptr|Q84TM8|Q84TM8 Polygalactorunase PG11 precursor (EC 3.2.1.15).
Length = 407
Score = 139 bits (351), Expect = 2e-32
Identities = 63/129 (48%), Positives = 89/129 (68%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLG+Y E +VE V V CTL T NG+RIK++ D+ ++ + ++D+ M+
Sbjct: 241 GHGISVGSLGKYTTEDNVEGVIVKNCTLTATQNGVRIKTWPDAPGTITVSDIHFEDIIMN 300
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NV P+IIDQ+YCP N C K SK+ ++ + FKNI GTS TPE + L C++ +PC GV+
Sbjct: 301 NVMNPVIIDQEYCPWNQCSKKNPSKIKLSKISFKNIQGTSGTPEGVVLICSSGVPCDGVE 360
Query: 363 LINVDIKYN 389
L NV + +N
Sbjct: 361 LNNVGLTFN 369
>sptr|Q9SFD0|Q9SFD0 Putative polygalacturonase.
Length = 401
Score = 139 bits (350), Expect = 3e-32
Identities = 66/155 (42%), Positives = 96/155 (61%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY E+DV + V CTL GT NGLRIK++ + + A+ ++++ ++
Sbjct: 233 GHGISVGSLGRYVHEQDVTGITVVNCTLQGTDNGLRIKTWPSAACATTASGIHFENIILN 292
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NVS PI+IDQ+YCP N C K S + + D+ FKNI GTS +A+ L C+ PC V+
Sbjct: 293 NVSNPILIDQEYCPWNQCNKQKPSTIKLVDISFKNIRGTSGNKDAVKLLCSKAHPCANVE 352
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
+ N++++Y +D +C N K +G AC
Sbjct: 353 IGNINLEYKGADGPPTFMCSNVSPKLVGTQNPKAC 387
>sptr|Q9MBB8|Q9MBB8 Polygalacturonase.
Length = 393
Score = 139 bits (349), Expect = 4e-32
Identities = 66/155 (42%), Positives = 95/155 (61%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY +EK V + V CT++ T NG+RIKS+ D A+ ++D+ M+
Sbjct: 240 GHGISVGSLGRYPNEKPVSGIFVKNCTISNTANGVRIKSWPDLYGGA-ASNMHFEDIVMN 298
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NV P+++DQ+YCP N C SKV ++DV FKNI GTS TP + L C+ +PC+ V+
Sbjct: 299 NVQNPVVVDQEYCPWNQCSLKSPSKVKISDVSFKNIRGTSATPVVVKLACSGGIPCEKVE 358
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
L ++++ Y+ S+ S C N G+ C
Sbjct: 359 LADINLVYSGSEGPAKSQCSNVKPTISGIMSASGC 393
>sptr|O48729|O48729 Putative polygalacturonase.
Length = 402
Score = 137 bits (345), Expect = 1e-31
Identities = 67/149 (44%), Positives = 99/149 (66%), Gaps = 1/149 (0%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLG+ KDEKDV+D+ V GT++G+RIK++E S S + + F+Y+++ M
Sbjct: 245 GHGISVGSLGKNKDEKDVKDLFVRDVIFNGTSDGIRIKTWESSASKILVSNFVYENIQMI 304
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
+V PI IDQKYCP+ C S V + D+ KNI+GTS A+ L C+ + PC+ V+
Sbjct: 305 DVGKPINIDQKYCPHPPCEHERKSHVQIQDLKLKNIYGTSKNKVAVNLQCSKSFPCKNVE 364
Query: 363 LINVDIKYN-RSDNKTMSVCKNAIGKSIG 446
LI+++IK N D +++VC+N G + G
Sbjct: 365 LIDINIKQNGLEDGSSITVCENVDGFARG 393
>sw|Q40312|PGLR_MEDSA Polygalacturonase precursor (EC 3.2.1.15) (PG)
(Pectinase).
Length = 421
Score = 137 bits (345), Expect = 1e-31
Identities = 63/141 (44%), Positives = 95/141 (67%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHG+SVGSLG++ E++VE + V CTL T NG+RIK++ D+ ++ + ++D+TM
Sbjct: 241 GHGLSVGSLGKFTTEENVEGITVKNCTLTATDNGVRIKTWPDAPGTITVSDIHFEDITMT 300
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NV P+IIDQ+Y P N C K SK+ ++ + FKN+ GTS T E + L C++ +PC GV+
Sbjct: 301 NVKNPVIIDQEYYPWNQCSKKNPSKIKLSKISFKNVKGTSGTAEGVVLICSSAVPCDGVE 360
Query: 363 LINVDIKYNRSDNKTMSVCKN 425
L NVD+K+N + T + C N
Sbjct: 361 LNNVDLKFNGA--PTTAKCTN 379
>sptr|Q9MBB7|Q9MBB7 Polygalacturonase.
Length = 393
Score = 137 bits (344), Expect = 2e-31
Identities = 64/141 (45%), Positives = 92/141 (65%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY +EK V + V CT++ T NG+RIKS+ D A+ ++D+ M+
Sbjct: 240 GHGISVGSLGRYPNEKPVSGIFVKNCTISNTANGVRIKSWPDLYGG-DASNMHFEDIVMN 298
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NV P+ +DQ+YCP N C SKV ++DV FKNI GTS TP + L C++ +PC+ V+
Sbjct: 299 NVQNPVAVDQEYCPWNQCSLKAPSKVKISDVSFKNIRGTSATPVVVKLACSSGIPCEKVE 358
Query: 363 LINVDIKYNRSDNKTMSVCKN 425
L ++++ Y+ S+ S C N
Sbjct: 359 LADINLVYSGSEGPAKSQCSN 379
>sptr|O49721|O49721 Polygalacturonase-like protein (Fragment).
Length = 362
Score = 136 bits (343), Expect = 2e-31
Identities = 68/159 (42%), Positives = 99/159 (62%), Gaps = 11/159 (6%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY +EKDV + V C ++GTTNG+RIK++ +S AT ++++ M+
Sbjct: 196 GHGISVGSLGRYPNEKDVNGLVVKDCKISGTTNGIRIKTWANSPGLSAATNMTFENIIMN 255
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NV+ PIIIDQ YCP + C+ + SKV ++++ FKNI GTS++ A+ L+C+ +PC+ V
Sbjct: 256 NVTNPIIIDQSYCPFSSCISNVPSKVELSEIYFKNIRGTSSSLVAVQLHCSRGMPCKKVY 315
Query: 363 LINVDI-----------KYNRSDNKTMSVCKNAIGKSIG 446
L NV + NR + S C+N IG
Sbjct: 316 LENVHLDLSSSDGGRKQSSNRGNEAVSSSCRNVRANYIG 354
>sptr|Q9M0M1|Q9M0M1 Polygalacturonase-like protein.
Length = 414
Score = 136 bits (343), Expect = 2e-31
Identities = 68/159 (42%), Positives = 99/159 (62%), Gaps = 11/159 (6%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY +EKDV + V C ++GTTNG+RIK++ +S AT ++++ M+
Sbjct: 248 GHGISVGSLGRYPNEKDVNGLVVKDCKISGTTNGIRIKTWANSPGLSAATNMTFENIIMN 307
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NV+ PIIIDQ YCP + C+ + SKV ++++ FKNI GTS++ A+ L+C+ +PC+ V
Sbjct: 308 NVTNPIIIDQSYCPFSSCISNVPSKVELSEIYFKNIRGTSSSLVAVQLHCSRGMPCKKVY 367
Query: 363 LINVDI-----------KYNRSDNKTMSVCKNAIGKSIG 446
L NV + NR + S C+N IG
Sbjct: 368 LENVHLDLSSSDGGRKQSSNRGNEAVSSSCRNVRANYIG 406
>sptr|Q8GXZ2|Q8GXZ2 Putative polygalacturonase (At3g07830).
Length = 225
Score = 135 bits (340), Expect = 5e-31
Identities = 64/155 (41%), Positives = 96/155 (61%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY E+DV ++V CTL T NGLRIK++ + S A+ ++++ +
Sbjct: 61 GHGISVGSLGRYGHEQDVSGIRVINCTLQETDNGLRIKTWPSAACSTTASNIHFENIILR 120
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NVS PI+IDQ+YCP N C K +S + + ++ F+ I GTS +A+ L C+ PC+ VQ
Sbjct: 121 NVSNPILIDQEYCPWNQCNKQKSSSIKLANISFRRIRGTSGNKDAVKLLCSKGYPCENVQ 180
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
+ +++I+Y +D +C N K +G AC
Sbjct: 181 VGDINIQYTGADGPATFMCSNVRPKLVGTQFPKAC 215
>sptr|Q9SFD1|Q9SFD1 Putative polygalacturonase.
Length = 397
Score = 135 bits (340), Expect = 5e-31
Identities = 64/155 (41%), Positives = 96/155 (61%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLGRY E+DV ++V CTL T NGLRIK++ + S A+ ++++ +
Sbjct: 233 GHGISVGSLGRYGHEQDVSGIRVINCTLQETDNGLRIKTWPSAACSTTASNIHFENIILR 292
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NVS PI+IDQ+YCP N C K +S + + ++ F+ I GTS +A+ L C+ PC+ VQ
Sbjct: 293 NVSNPILIDQEYCPWNQCNKQKSSSIKLANISFRRIRGTSGNKDAVKLLCSKGYPCENVQ 352
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIGMAKELAC 467
+ +++I+Y +D +C N K +G AC
Sbjct: 353 VGDINIQYTGADGPATFMCSNVRPKLVGTQFPKAC 387
>sptr|Q9S761|Q9S761 Putative polygalacturonase.
Length = 404
Score = 134 bits (338), Expect = 8e-31
Identities = 67/159 (42%), Positives = 103/159 (64%), Gaps = 3/159 (1%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLG+ KDEKDV+++ V GT++G+RIK++E S S + + F+Y+++ M
Sbjct: 245 GHGISVGSLGKNKDEKDVKNLTVRDVIFNGTSDGIRIKTWESSASKILVSNFVYENIQMI 304
Query: 183 NVSYPIIIDQKYCPNNIC--VKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQG 356
+V PI IDQKYCP+ C + G S V + D+ KNI+GTS A+ L C+ + PC+
Sbjct: 305 DVGKPINIDQKYCPHPPCEHEQKGESHVQIQDLKLKNIYGTSKNKVAMNLQCSKSFPCKN 364
Query: 357 VQLINVDIKYNRSDN-KTMSVCKNAIGKSIGMAKELACV 470
++LI+++IK N +N +++VC+N G G C+
Sbjct: 365 IELIDINIKSNGLENSSSIAVCENVDGSMSGKMVPQHCL 403
>sptr|Q9SHH4|Q9SHH4 Similar to polygalacturonase.
Length = 369
Score = 134 bits (337), Expect = 1e-30
Identities = 62/148 (41%), Positives = 92/148 (62%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGIS+GSLG+ KDEK+V + V GTTNG+RIK++E S S+++ +Y+++ M
Sbjct: 213 GHGISIGSLGKNKDEKNVNGLMVRNSVFTGTTNGIRIKTWESSASTIRIINLVYENLQMI 272
Query: 183 NVSYPIIIDQKYCPNNICVKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQGVQ 362
NV PI IDQKYCP C G S + + +V KNI GTS A+ C+ PC+ VQ
Sbjct: 273 NVENPIGIDQKYCPYPPCSNMGDSHIQIRNVTLKNIWGTSKNKVAVKFQCSKTFPCKDVQ 332
Query: 363 LINVDIKYNRSDNKTMSVCKNAIGKSIG 446
LI++++ ++ D ++C+N G + G
Sbjct: 333 LIDINLTHHGVDGPASALCENVDGSATG 360
>sptr|Q9SJN7|Q9SJN7 Putative polygalacturonase.
Length = 404
Score = 133 bits (335), Expect = 2e-30
Identities = 68/151 (45%), Positives = 99/151 (65%), Gaps = 3/151 (1%)
Frame = +3
Query: 3 GHGISVGSLGRYKDEKDVEDVKVTGCTLAGTTNGLRIKSYEDSKSSLKATKFLYQDVTMD 182
GHGISVGSLG+ KDEKDV+D+ V GT++G+RIK++E S S + + F+Y+++ M
Sbjct: 245 GHGISVGSLGKNKDEKDVKDLIVRDVIFNGTSDGIRIKNWESSASKILVSNFVYENIQMI 304
Query: 183 NVSYPIIIDQKYCPNNIC--VKSGASKVAVNDVVFKNIHGTSNTPEAITLNCANNLPCQG 356
+V PI IDQKYCP+ C + G S V + ++ KNI+GTS A+ L C+ PC+
Sbjct: 305 DVGKPINIDQKYCPHPPCEHERKGESHVQIQNLKLKNIYGTSKNKVAVNLQCSKIFPCKN 364
Query: 357 VQLINVDIKYNR-SDNKTMSVCKNAIGKSIG 446
V+LI+++IK N D + SVC+N G + G
Sbjct: 365 VELIDINIKQNGVKDGSSTSVCENVDGFARG 395
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 330,752,943
Number of Sequences: 1395590
Number of extensions: 5259865
Number of successful extensions: 23352
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 22519
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 23063
length of database: 442,889,342
effective HSP length: 118
effective length of database: 278,209,722
effective search space used: 26429923590
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

