BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
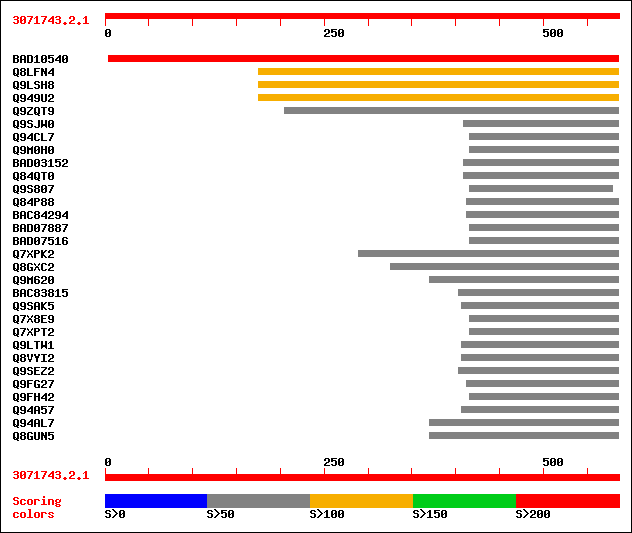
Query= 3071743.2.1
(586 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptrnew|BAD10540|BAD10540 Putative transfactor. 219 2e-56
sptr|Q8LFN4|Q8LFN4 Transfactor-like protein. 142 2e-33
sptr|Q9LSH8|Q9LSH8 Transfactor-like protein. 142 2e-33
sptr|Q949U2|Q949U2 Hypothetical protein. 142 2e-33
sptr|Q9ZQT9|Q9ZQT9 WERBP-1 protein. 99 4e-20
sptr|Q9SJW0|Q9SJW0 Transfactor-like protein. 99 5e-20
sptr|Q94CL7|Q94CL7 Phosphate starvation response regulator 1 (Hy... 96 4e-19
sptr|Q9M0H0|Q9M0H0 Hypothetical protein. 96 4e-19
sptrnew|BAD03152|BAD03152 Putative transfactor. 96 4e-19
sptr|Q84QT0|Q84QT0 Putative transfactor. 96 4e-19
sptr|Q9S807|Q9S807 Phosphate starvation regulator protein. 95 7e-19
sptr|Q84P88|Q84P88 Phosphate starvation response regulator-like ... 93 2e-18
sptrnew|BAC84294|BAC84294 Putative CDPK substrate protein 1. 93 2e-18
sptrnew|BAD07887|BAD07887 Phosphate starvation response regulato... 93 3e-18
sptrnew|BAD07516|BAD07516 Phosphate starvation response regulato... 93 3e-18
sptr|Q7XPK2|Q7XPK2 OSJNBa0087O24.8 protein. 92 4e-18
sptr|Q8GXC2|Q8GXC2 Hypothetical protein. 92 5e-18
sptr|Q9M620|Q9M620 CDPK substrate protein 1. 92 5e-18
sptrnew|BAC83815|BAC83815 Transfactor-like protein. 91 8e-18
sptr|Q9SAK5|Q9SAK5 T8K14.15 protein. 90 2e-17
sptr|Q7X8E9|Q7X8E9 Putative phosphate starvation response regula... 90 2e-17
sptr|Q7XPT2|Q7XPT2 OSJNBa0083N12.9 protein. 89 3e-17
sptr|Q9LTW1|Q9LTW1 Similarity to transfactor. 89 3e-17
sptr|Q8VYI2|Q8VYI2 AT3g12730/MBK21_9. 89 3e-17
sptr|Q9SEZ2|Q9SEZ2 Transfactor, putative, 28697-27224. 89 3e-17
sptr|Q9FG27|Q9FG27 Similarity to transfactor. 88 7e-17
sptr|Q9FH42|Q9FH42 Similarity to transfactor. 88 9e-17
sptr|Q94A57|Q94A57 AT3g24120/MUJ8_3 (Transfactor, putative). 87 2e-16
sptr|Q94AL7|Q94AL7 Hypothetical protein (Fragment). 87 2e-16
sptr|Q8GUN5|Q8GUN5 Hypothetical protein. 87 2e-16
>sptrnew|BAD10540|BAD10540 Putative transfactor.
Length = 467
Score = 219 bits (558), Expect = 2e-56
Identities = 118/199 (59%), Positives = 132/199 (66%), Gaps = 5/199 (2%)
Frame = +1
Query: 4 GTLPFLPHPPKCEQQV-SAGHXXXXXXXXXGDTGLDVAEQXXXXXXXXXXXXXXXXXXFH 180
G LPFLP PPKCEQQ SAG D FH
Sbjct: 123 GYLPFLPQPPKCEQQQNSAGQSSSSLMLLDADLRNSGHADDEHTDDLKDFLNLSSDCSFH 182
Query: 181 GENNALAFDEQMEFQFLSEQLEIAITDNEKNPHLDDIYGTPPQLSSLPVSSCSNQ----N 348
G+ +A+A++EQMEFQFLSEQL IAI++NE++P LDDIY PPQL SLPVSSCS+Q +
Sbjct: 183 GKCSAMAYNEQMEFQFLSEQLGIAISNNEESPRLDDIYDRPPQLMSLPVSSCSDQEDLQD 242
Query: 349 LGSPVKVQLXXXXXXXXXATANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKV 528
SP KVQL A+ NK RLRWT ELHERFV+AV KLEGPEKATPKGVLKLMKV
Sbjct: 243 ARSPAKVQLSSSRSSSGTASCNKPRLRWTPELHERFVDAVNKLEGPEKATPKGVLKLMKV 302
Query: 529 EGLTIYHVKSHLQKYRLAK 585
EGLTIYH+KSHLQKYRLAK
Sbjct: 303 EGLTIYHIKSHLQKYRLAK 321
>sptr|Q8LFN4|Q8LFN4 Transfactor-like protein.
Length = 443
Score = 142 bits (359), Expect = 2e-33
Identities = 79/139 (56%), Positives = 93/139 (66%), Gaps = 2/139 (1%)
Frame = +1
Query: 175 FHGENNALAFDEQMEFQFLSEQLEIAITDNEKNPHLDDIYGTPPQLSSLPVSSCSNQNLG 354
F N++ +QME QFLS++LE+AITD + P LD+IY TP L+S PV+ S
Sbjct: 155 FGCSNDSYCLSDQMELQFLSDELELAITDRAETPRLDEIYETP--LASNPVTRLSPSQSC 212
Query: 355 SP--VKVQLXXXXXXXXXATANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKV 528
P + V + A KSR+RWT ELHE FV+AV KLEGPEKATPK V KLM V
Sbjct: 213 VPGAMSVDVVSSHPSPGSAANQKSRMRWTPELHESFVKAVIKLEGPEKATPKAVKKLMNV 272
Query: 529 EGLTIYHVKSHLQKYRLAK 585
EGLTIYHVKSHLQKYRLAK
Sbjct: 273 EGLTIYHVKSHLQKYRLAK 291
>sptr|Q9LSH8|Q9LSH8 Transfactor-like protein.
Length = 554
Score = 142 bits (359), Expect = 2e-33
Identities = 79/139 (56%), Positives = 93/139 (66%), Gaps = 2/139 (1%)
Frame = +1
Query: 175 FHGENNALAFDEQMEFQFLSEQLEIAITDNEKNPHLDDIYGTPPQLSSLPVSSCSNQNLG 354
F N++ +QME QFLS++LE+AITD + P LD+IY TP L+S PV+ S
Sbjct: 161 FGCSNDSYCLSDQMELQFLSDELELAITDRAETPRLDEIYETP--LASNPVTRLSPSQSC 218
Query: 355 SP--VKVQLXXXXXXXXXATANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKV 528
P + V + A KSR+RWT ELHE FV+AV KLEGPEKATPK V KLM V
Sbjct: 219 VPGAMSVDVVSSHPSPGSAANQKSRMRWTPELHESFVKAVIKLEGPEKATPKAVKKLMNV 278
Query: 529 EGLTIYHVKSHLQKYRLAK 585
EGLTIYHVKSHLQKYRLAK
Sbjct: 279 EGLTIYHVKSHLQKYRLAK 297
>sptr|Q949U2|Q949U2 Hypothetical protein.
Length = 449
Score = 142 bits (359), Expect = 2e-33
Identities = 79/139 (56%), Positives = 93/139 (66%), Gaps = 2/139 (1%)
Frame = +1
Query: 175 FHGENNALAFDEQMEFQFLSEQLEIAITDNEKNPHLDDIYGTPPQLSSLPVSSCSNQNLG 354
F N++ +QME QFLS++LE+AITD + P LD+IY TP L+S PV+ S
Sbjct: 161 FGCSNDSYCLSDQMELQFLSDELELAITDRAETPRLDEIYETP--LASNPVTRLSPSQSC 218
Query: 355 SP--VKVQLXXXXXXXXXATANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKV 528
P + V + A KSR+RWT ELHE FV+AV KLEGPEKATPK V KLM V
Sbjct: 219 VPGAMSVDVVSSHPSPGSAANQKSRMRWTPELHESFVKAVIKLEGPEKATPKAVKKLMNV 278
Query: 529 EGLTIYHVKSHLQKYRLAK 585
EGLTIYHVKSHLQKYRLAK
Sbjct: 279 EGLTIYHVKSHLQKYRLAK 297
>sptr|Q9ZQT9|Q9ZQT9 WERBP-1 protein.
Length = 291
Score = 99.0 bits (245), Expect = 4e-20
Identities = 59/129 (45%), Positives = 71/129 (55%), Gaps = 2/129 (1%)
Frame = +1
Query: 205 DEQMEFQFLSEQLEIAITDNEKNPHLDDIYGTPPQLSSLPVSSCS--NQNLGSPVKVQLX 378
D+ + + L+ I D E Y Q S+ PV Q + V+
Sbjct: 4 DDALTSNWNDIMLDTGIADAEPKMQ----YQEQKQPSNFPVHQGQPLQQVPTASVETSAI 59
Query: 379 XXXXXXXXATANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKS 558
++K R+RWT ELHE FVEAV KL G E+ATPKGVLKLMKVEGLTIYHVKS
Sbjct: 60 VPASSTASGASSKQRMRWTPELHEAFVEAVNKLGGSERATPKGVLKLMKVEGLTIYHVKS 119
Query: 559 HLQKYRLAK 585
HLQKYR A+
Sbjct: 120 HLQKYRTAR 128
>sptr|Q9SJW0|Q9SJW0 Transfactor-like protein.
Length = 286
Score = 98.6 bits (244), Expect = 5e-20
Identities = 46/59 (77%), Positives = 54/59 (91%)
Frame = +1
Query: 409 ANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
A+K RLRWT ELHERFV+AV +L GP++ATPKGVL++M V+GLTIYHVKSHLQKYRLAK
Sbjct: 13 ASKQRLRWTHELHERFVDAVAQLGGPDRATPKGVLRVMGVQGLTIYHVKSHLQKYRLAK 71
>sptr|Q94CL7|Q94CL7 Phosphate starvation response regulator 1
(Hypothetical protein).
Length = 409
Score = 95.5 bits (236), Expect = 4e-19
Identities = 45/57 (78%), Positives = 50/57 (87%)
Frame = +1
Query: 415 KSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
K+R+RWT ELHE FVEAV L G E+ATPKGVLK+MKVEGLTIYHVKSHLQKYR A+
Sbjct: 225 KARMRWTPELHEAFVEAVNSLGGSERATPKGVLKIMKVEGLTIYHVKSHLQKYRTAR 281
>sptr|Q9M0H0|Q9M0H0 Hypothetical protein.
Length = 422
Score = 95.5 bits (236), Expect = 4e-19
Identities = 45/57 (78%), Positives = 50/57 (87%)
Frame = +1
Query: 415 KSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
K+R+RWT ELHE FVEAV L G E+ATPKGVLK+MKVEGLTIYHVKSHLQKYR A+
Sbjct: 225 KARMRWTPELHEAFVEAVNSLGGSERATPKGVLKIMKVEGLTIYHVKSHLQKYRTAR 281
>sptrnew|BAD03152|BAD03152 Putative transfactor.
Length = 307
Score = 95.5 bits (236), Expect = 4e-19
Identities = 45/59 (76%), Positives = 52/59 (88%)
Frame = +1
Query: 409 ANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
A + RLRWT ELH+RFVEAV +L GP++ATPKGVL++M V GLTIYHVKSHLQKYRLAK
Sbjct: 43 AARQRLRWTNELHDRFVEAVTQLGGPDRATPKGVLRIMGVPGLTIYHVKSHLQKYRLAK 101
>sptr|Q84QT0|Q84QT0 Putative transfactor.
Length = 307
Score = 95.5 bits (236), Expect = 4e-19
Identities = 45/59 (76%), Positives = 52/59 (88%)
Frame = +1
Query: 409 ANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
A + RLRWT ELH+RFVEAV +L GP++ATPKGVL++M V GLTIYHVKSHLQKYRLAK
Sbjct: 43 AARQRLRWTNELHDRFVEAVTQLGGPDRATPKGVLRIMGVPGLTIYHVKSHLQKYRLAK 101
>sptr|Q9S807|Q9S807 Phosphate starvation regulator protein.
Length = 752
Score = 94.7 bits (234), Expect = 7e-19
Identities = 44/55 (80%), Positives = 48/55 (87%)
Frame = +1
Query: 415 KSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRL 579
KSRLRWT ELH RFV AV L GP+KATPKG+LKLM V+GLTIYH+KSHLQKYRL
Sbjct: 187 KSRLRWTPELHNRFVNAVNSLGGPDKATPKGILKLMGVDGLTIYHIKSHLQKYRL 241
>sptr|Q84P88|Q84P88 Phosphate starvation response regulator-like
protein.
Length = 361
Score = 93.2 bits (230), Expect = 2e-18
Identities = 44/58 (75%), Positives = 50/58 (86%)
Frame = +1
Query: 412 NKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
+K+R+RWT ELHERFV+AV L G EKATPKGVLKLMK + LTIYHVKSHLQKYR A+
Sbjct: 245 SKTRMRWTPELHERFVDAVNLLGGSEKATPKGVLKLMKADNLTIYHVKSHLQKYRTAR 302
>sptrnew|BAC84294|BAC84294 Putative CDPK substrate protein 1.
Length = 426
Score = 93.2 bits (230), Expect = 2e-18
Identities = 44/58 (75%), Positives = 50/58 (86%)
Frame = +1
Query: 412 NKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
+K+R+RWT ELHERFV+AV L G EKATPKGVLKLMK + LTIYHVKSHLQKYR A+
Sbjct: 245 SKTRMRWTPELHERFVDAVNLLGGSEKATPKGVLKLMKADNLTIYHVKSHLQKYRTAR 302
>sptrnew|BAD07887|BAD07887 Phosphate starvation response
regulator-like.
Length = 284
Score = 92.8 bits (229), Expect = 3e-18
Identities = 43/57 (75%), Positives = 50/57 (87%)
Frame = +1
Query: 415 KSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
K RLRWT +LHERFVEAV KL GP+KATPK VL+LM ++GLT+YH+KSHLQKYRL K
Sbjct: 27 KPRLRWTPDLHERFVEAVTKLGGPDKATPKSVLRLMGMKGLTLYHLKSHLQKYRLGK 83
>sptrnew|BAD07516|BAD07516 Phosphate starvation response
regulator-like.
Length = 284
Score = 92.8 bits (229), Expect = 3e-18
Identities = 43/57 (75%), Positives = 50/57 (87%)
Frame = +1
Query: 415 KSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
K RLRWT +LHERFVEAV KL GP+KATPK VL+LM ++GLT+YH+KSHLQKYRL K
Sbjct: 27 KPRLRWTPDLHERFVEAVTKLGGPDKATPKSVLRLMGMKGLTLYHLKSHLQKYRLGK 83
>sptr|Q7XPK2|Q7XPK2 OSJNBa0087O24.8 protein.
Length = 419
Score = 92.4 bits (228), Expect = 4e-18
Identities = 48/111 (43%), Positives = 69/111 (62%), Gaps = 12/111 (10%)
Frame = +1
Query: 289 IYGTPPQLSSLPVSSCSNQ-NLGSPVKVQLXX-----------XXXXXXXATANKSRLRW 432
I+ PPQ+++ P+ + +L + ++ QL +K+R+RW
Sbjct: 182 IHSFPPQVAAKPILPAMDAPSLQNQMENQLTRNCIGAATPVTPTGNLAGSGAPSKTRIRW 241
Query: 433 TLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
T +LHERFV+ V +L G +KATPKG+LKLM +GLTIYH+KSHLQKYR+AK
Sbjct: 242 TQDLHERFVDCVNQLGGADKATPKGILKLMNSDGLTIYHIKSHLQKYRIAK 292
>sptr|Q8GXC2|Q8GXC2 Hypothetical protein.
Length = 397
Score = 92.0 bits (227), Expect = 5e-18
Identities = 52/87 (59%), Positives = 57/87 (65%)
Frame = +1
Query: 325 VSSCSNQNLGSPVKVQLXXXXXXXXXATANKSRLRWTLELHERFVEAVKKLEGPEKATPK 504
VSS SN N S A A K R+RWT ELHE FV+AV +L G +ATPK
Sbjct: 213 VSSNSNNNSNS------------NNAAAAAKGRMRWTPELHEVFVDAVNQLGGSNEATPK 260
Query: 505 GVLKLMKVEGLTIYHVKSHLQKYRLAK 585
GVLK MKVEGLTI+HVKSHLQKYR AK
Sbjct: 261 GVLKHMKVEGLTIFHVKSHLQKYRTAK 287
>sptr|Q9M620|Q9M620 CDPK substrate protein 1.
Length = 470
Score = 92.0 bits (227), Expect = 5e-18
Identities = 44/72 (61%), Positives = 52/72 (72%)
Frame = +1
Query: 370 QLXXXXXXXXXATANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYH 549
Q A +N+ R+RWT ELHE FV+AV +L G E+ATPKGVL+ M VEGLTIYH
Sbjct: 241 QTSAAANQTSAAHSNRPRMRWTPELHEAFVDAVNQLGGSERATPKGVLRHMNVEGLTIYH 300
Query: 550 VKSHLQKYRLAK 585
VKSHLQKYR A+
Sbjct: 301 VKSHLQKYRTAR 312
>sptrnew|BAC83815|BAC83815 Transfactor-like protein.
Length = 339
Score = 91.3 bits (225), Expect = 8e-18
Identities = 41/61 (67%), Positives = 52/61 (85%)
Frame = +1
Query: 403 ATANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLA 582
+T K RL+WT ELHERFVEAV +L GP+KATPK +++LM + GLT+YH+KSHLQKYRL+
Sbjct: 42 STDAKPRLKWTSELHERFVEAVNQLGGPDKATPKTIMRLMGIPGLTLYHLKSHLQKYRLS 101
Query: 583 K 585
K
Sbjct: 102 K 102
>sptr|Q9SAK5|Q9SAK5 T8K14.15 protein.
Length = 367
Score = 89.7 bits (221), Expect = 2e-17
Identities = 40/60 (66%), Positives = 52/60 (86%)
Frame = +1
Query: 406 TANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
T K RLRWT+ELHERFV+AV +L GP+KATPK ++++M V+GLT+YH+KSHLQK+RL K
Sbjct: 31 TDPKPRLRWTVELHERFVDAVAQLGGPDKATPKTIMRVMGVKGLTLYHLKSHLQKFRLGK 90
>sptr|Q7X8E9|Q7X8E9 Putative phosphate starvation response regulator
(Putative calcium-dependent protein kinase substrate
protein).
Length = 282
Score = 89.7 bits (221), Expect = 2e-17
Identities = 40/57 (70%), Positives = 50/57 (87%)
Frame = +1
Query: 415 KSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
K RLRWT +LH+RFV+AV KL GP+KATPK VL+LM ++GLT+YH+KSHLQKYRL +
Sbjct: 26 KPRLRWTADLHDRFVDAVTKLGGPDKATPKSVLRLMGLKGLTLYHLKSHLQKYRLGQ 82
>sptr|Q7XPT2|Q7XPT2 OSJNBa0083N12.9 protein.
Length = 249
Score = 89.4 bits (220), Expect = 3e-17
Identities = 40/57 (70%), Positives = 50/57 (87%)
Frame = +1
Query: 415 KSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
K RLRWT +LHERFV+AV +L GP+KATPK VL+LM ++GLT+YH+KSHLQKYRL +
Sbjct: 21 KPRLRWTPDLHERFVDAVTRLGGPDKATPKSVLRLMGMKGLTLYHLKSHLQKYRLGR 77
>sptr|Q9LTW1|Q9LTW1 Similarity to transfactor.
Length = 228
Score = 89.4 bits (220), Expect = 3e-17
Identities = 41/60 (68%), Positives = 50/60 (83%)
Frame = +1
Query: 406 TANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
T K RLRWT ELHERFV+AV L GPEKATPK ++++M V+GLT+YH+KSHLQK+RL K
Sbjct: 20 TDPKPRLRWTTELHERFVDAVTHLGGPEKATPKTIMRVMGVKGLTLYHLKSHLQKFRLGK 79
>sptr|Q8VYI2|Q8VYI2 AT3g12730/MBK21_9.
Length = 235
Score = 89.4 bits (220), Expect = 3e-17
Identities = 41/60 (68%), Positives = 50/60 (83%)
Frame = +1
Query: 406 TANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
T K RLRWT ELHERFV+AV L GPEKATPK ++++M V+GLT+YH+KSHLQK+RL K
Sbjct: 20 TDPKPRLRWTTELHERFVDAVTHLGGPEKATPKTIMRVMGVKGLTLYHLKSHLQKFRLGK 79
>sptr|Q9SEZ2|Q9SEZ2 Transfactor, putative, 28697-27224.
Length = 332
Score = 89.4 bits (220), Expect = 3e-17
Identities = 38/61 (62%), Positives = 51/61 (83%)
Frame = +1
Query: 403 ATANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLA 582
+T K RL+WT +LH +F+EAV +L GP KATPKG++K+M++ GLT+YH+KSHLQKYRL
Sbjct: 26 STDAKPRLKWTCDLHHKFIEAVNQLGGPNKATPKGLMKVMEIPGLTLYHLKSHLQKYRLG 85
Query: 583 K 585
K
Sbjct: 86 K 86
>sptr|Q9FG27|Q9FG27 Similarity to transfactor.
Length = 374
Score = 88.2 bits (217), Expect = 7e-17
Identities = 39/58 (67%), Positives = 49/58 (84%)
Frame = +1
Query: 412 NKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
NK+R+RWT +LHE+FVE V +L G +KATPK +LK M +GLTI+HVKSHLQKYR+AK
Sbjct: 190 NKTRIRWTQDLHEKFVECVNRLGGADKATPKAILKRMDSDGLTIFHVKSHLQKYRIAK 247
>sptr|Q9FH42|Q9FH42 Similarity to transfactor.
Length = 308
Score = 87.8 bits (216), Expect = 9e-17
Identities = 40/57 (70%), Positives = 49/57 (85%)
Frame = +1
Query: 415 KSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
K RLRWT +LH+RFV+AV KL G +KATPK VLKLM ++GLT+YH+KSHLQKYRL +
Sbjct: 23 KPRLRWTADLHDRFVDAVAKLGGADKATPKSVLKLMGLKGLTLYHLKSHLQKYRLGQ 79
>sptr|Q94A57|Q94A57 AT3g24120/MUJ8_3 (Transfactor, putative).
Length = 295
Score = 87.0 bits (214), Expect = 2e-16
Identities = 39/60 (65%), Positives = 50/60 (83%)
Frame = +1
Query: 406 TANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRLAK 585
T K RLRWT ELHERFV+AV +L GP+KATPK +++ M V+GLT+YH+KSHLQK+RL +
Sbjct: 38 TDPKPRLRWTTELHERFVDAVTQLGGPDKATPKTIMRTMGVKGLTLYHLKSHLQKFRLGR 97
>sptr|Q94AL7|Q94AL7 Hypothetical protein (Fragment).
Length = 385
Score = 87.0 bits (214), Expect = 2e-16
Identities = 45/72 (62%), Positives = 51/72 (70%)
Frame = +1
Query: 370 QLXXXXXXXXXATANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYH 549
QL AT+ K R+RWT ELHE FVEAV +L G E+ATPK VLKL+ GLTIYH
Sbjct: 189 QLSGRNSSSSVATS-KQRMRWTPELHEAFVEAVNQLGGSERATPKAVLKLLNNPGLTIYH 247
Query: 550 VKSHLQKYRLAK 585
VKSHLQKYR A+
Sbjct: 248 VKSHLQKYRTAR 259
>sptr|Q8GUN5|Q8GUN5 Hypothetical protein.
Length = 413
Score = 87.0 bits (214), Expect = 2e-16
Identities = 45/72 (62%), Positives = 51/72 (70%)
Frame = +1
Query: 370 QLXXXXXXXXXATANKSRLRWTLELHERFVEAVKKLEGPEKATPKGVLKLMKVEGLTIYH 549
QL AT+ K R+RWT ELHE FVEAV +L G E+ATPK VLKL+ GLTIYH
Sbjct: 217 QLSGRNSSSSVATS-KQRMRWTPELHEAFVEAVNQLGGSERATPKAVLKLLNNPGLTIYH 275
Query: 550 VKSHLQKYRLAK 585
VKSHLQKYR A+
Sbjct: 276 VKSHLQKYRTAR 287
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.314 0.133 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 416,986,211
Number of Sequences: 1395590
Number of extensions: 7097920
Number of successful extensions: 15052
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 14118
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 14948
length of database: 442,889,342
effective HSP length: 117
effective length of database: 279,605,312
effective search space used: 21529609024
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)

