BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
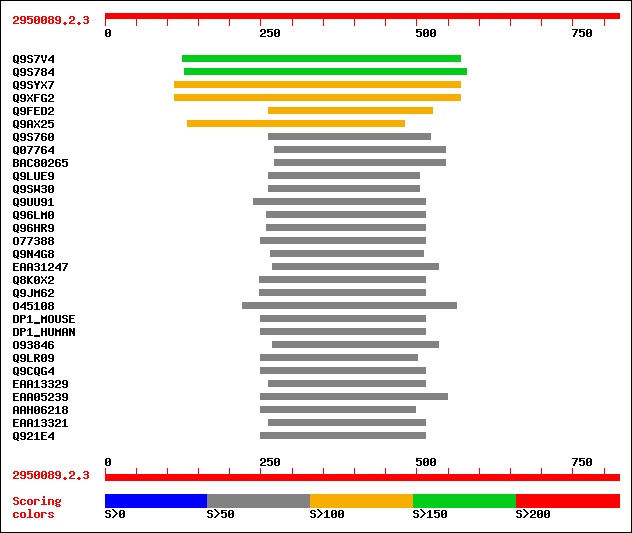
Query= 2950089.2.3
(824 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q9S7V4|Q9S7V4 ATHVA22A. 194 9e-49
sptr|Q9S784|Q9S784 ATHVA22C. 155 6e-37
sptr|Q9SYX7|Q9SYX7 AtHVA22b. 148 1e-34
sptr|Q9XFG2|Q9XFG2 ATHVA22B. 147 1e-34
sptr|Q9FED2|Q9FED2 AtHVA22e (Abscisic acid-induced-like protein). 104 2e-21
sptr|Q9AX25|Q9AX25 P0456A01.10 protein (P0042A10.34 protein). 102 8e-21
sptr|Q9S760|Q9S760 ATHVA22D (Putative abscisic acid-induced prot... 100 4e-20
sptr|Q07764|Q07764 Abscisic acid and stress inducible (A22) gene. 99 7e-20
sptrnew|BAC80265|BAC80265 Hypothetical protein Wrab15. 99 9e-20
sptr|Q9LUE9|Q9LUE9 Genomic DNA, chromosome 5, P1 clone:MFB16. 95 1e-18
sptr|Q9SW30|Q9SW30 Abscisic acid-induced-like protein. 92 1e-17
sptr|Q9UU91|Q9UU91 Putative transport protein. 71 2e-11
sptr|Q96LM0|Q96LM0 Hypothetical protein FLJ25383 (TB2 protein-li... 67 3e-10
sptr|Q96HR9|Q96HR9 Hypothetical protein. 67 3e-10
sptr|O77388|O77388 Hypothetical 26.0 kDa protein. 67 4e-10
sptr|Q9N4G8|Q9N4G8 Hypothetical 20.6 kDa protein. 66 6e-10
sptrnew|EAA31247|EAA31247 Hypothetical protein. 65 8e-10
sptr|Q8K0X2|Q8K0X2 Hypothetical protein. 65 1e-09
sptr|Q9JM62|Q9JM62 Polyposis locus protein 1-like 1 (TB2 protein... 65 1e-09
sptr|O45108|O45108 Hypothetical 22.8 kDa protein. 65 1e-09
sw|Q60870|DP1_MOUSE Polyposis locus protein 1 homolog (TB2 prote... 65 1e-09
sw|Q00765|DP1_HUMAN Polyposis locus protein 1 (TB2 protein). 64 2e-09
sptr|O93846|O93846 Pathogenicity protein. 64 3e-09
sptr|Q9LR09|Q9LR09 F10A5.11. 64 3e-09
sptr|Q9CQG4|Q9CQG4 Deleted in polyposis 1. 63 4e-09
sptrnew|EAA13329|EAA13329 ENSANGP00000003353 (Fragment). 63 5e-09
sptrnew|EAA05239|EAA05239 ENSANGP00000018512 (Fragment). 63 5e-09
sptrnew|AAH06218|AAH06218 Hypothetical protein. 63 5e-09
sptrnew|EAA13321|EAA13321 AgCP2771 (Fragment). 63 5e-09
sptr|Q921E4|Q921E4 Deleted in polyposis 1. 62 1e-08
>sptr|Q9S7V4|Q9S7V4 ATHVA22A.
Length = 177
Score = 194 bits (494), Expect = 9e-49
Identities = 89/150 (59%), Positives = 115/150 (76%), Gaps = 1/150 (0%)
Frame = -3
Query: 570 GSGSFLKVLANNFDVLAGPLISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFEL 391
G+G+FLKVL NFDVLAGP++SL YPLYASV+AIET++ DD+QWLTYWV S +TL EL
Sbjct: 4 GAGNFLKVLLRNFDVLAGPVVSLVYPLYASVQAIETQSHADDKQWLTYWVLYSLLTLIEL 63
Query: 390 TFAPIIEWLPFWSYAKLFFNCXLVLPWFNGXAYVYDHFXRPMFVNRQIVNIWYIXRN-EK 214
TFA +IEWLP WSY KL C LV+P+F+G AYVY+HF RP+FVN + +NIWY+ + +
Sbjct: 64 TFAKLIEWLPIWSYMKLILTCWLVIPYFSGAAYVYEHFVRPVFVNPRSINIWYVPKKMDI 123
Query: 213 LGKSDDVXXAAERYIEQNXXXAFXXLISKS 124
K DDV AAE+YI +N AF ++S++
Sbjct: 124 FRKPDDVLTAAEKYIAENGPDAFEKILSRA 153
>sptr|Q9S784|Q9S784 ATHVA22C.
Length = 184
Score = 155 bits (392), Expect = 6e-37
Identities = 77/157 (49%), Positives = 100/157 (63%), Gaps = 6/157 (3%)
Frame = -3
Query: 579 SKMGSGSFLKVLANNFDVLAGPLISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITL 400
S G + L+VL NFDVLA PL++L YPLYASV+AIET++ +D+QWLTYWV + I+L
Sbjct: 3 SNSGDDNVLQVLIKNFDVLALPLVTLVYPLYASVKAIETRSLPEDEQWLTYWVLYALISL 62
Query: 399 FELTFAPIIEWLPFWSYAKLFFNCXLVLPWFNGXAYVYDHFXRPMF--VNRQIVNIWYIX 226
FELTF+ +EW P W Y KLF C LVLP FNG ++Y HF RP + R IWY+
Sbjct: 63 FELTFSKPLEWFPIWPYMKLFGICWLVLPQFNGAEHIYKHFIRPFYRDPQRATTKIWYVP 122
Query: 225 RNE----KLGKSDDVXXAAERYIEQNXXXAFXXLISK 127
+ DD+ AAE+Y+EQ+ AF +I K
Sbjct: 123 HKKFNFFPKRDDDDILTAAEKYMEQHGTEAFERMIVK 159
>sptr|Q9SYX7|Q9SYX7 AtHVA22b.
Length = 167
Score = 148 bits (373), Expect = 1e-34
Identities = 73/153 (47%), Positives = 95/153 (62%)
Frame = -3
Query: 570 GSGSFLKVLANNFDVLAGPLISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFEL 391
G GS +KV+ NFDV+AGP+ISL YPLYASVRAIE+++ DD+QWLTYW S I LFEL
Sbjct: 4 GIGSLVKVIFKNFDVIAGPMISLVYPLYASVRAIESRSHGDDKQWLTYWALYSLIKLFEL 63
Query: 390 TFAPIIEWLPFWSYAKLFFNCXLVLPWFNGXAYVYDHFXRPMFVNRQIVNIWYIXRNEKL 211
TF ++EW+P + YAKL LVLP NG AY+Y+H+ R ++ VN+WY+
Sbjct: 64 TFFRLLEWIPLYPYAKLALTSWLVLPGMNGAAYLYEHYVRSFLLSPHTVNVWYVPAK--- 120
Query: 210 GKSDDVXXAAERYIEQNXXXAFXXLISKSTXXS 112
K DD+ A ++ N A I S S
Sbjct: 121 -KDDDLGATAGKFTPVNDSGAPQEKIVSSVDTS 152
>sptr|Q9XFG2|Q9XFG2 ATHVA22B.
Length = 167
Score = 147 bits (372), Expect = 1e-34
Identities = 73/153 (47%), Positives = 95/153 (62%)
Frame = -3
Query: 570 GSGSFLKVLANNFDVLAGPLISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFEL 391
G GS +KV+ NFDV+AGP+ISL YPLYASVRAIE+++ DD+QWLTYW S I LFEL
Sbjct: 4 GIGSLVKVIFKNFDVIAGPVISLVYPLYASVRAIESRSHGDDKQWLTYWALYSLIKLFEL 63
Query: 390 TFAPIIEWLPFWSYAKLFFNCXLVLPWFNGXAYVYDHFXRPMFVNRQIVNIWYIXRNEKL 211
TF ++EW+P + YAKL LVLP NG AY+Y+H+ R ++ VN+WY+
Sbjct: 64 TFFRLLEWIPLYPYAKLALTSWLVLPGMNGAAYLYEHYVRSFLLSPHTVNVWYVPAK--- 120
Query: 210 GKSDDVXXAAERYIEQNXXXAFXXLISKSTXXS 112
K DD+ A ++ N A I S S
Sbjct: 121 -KDDDLGATAGKFTPVNDSGAPQEKIVSSVDTS 152
>sptr|Q9FED2|Q9FED2 AtHVA22e (Abscisic acid-induced-like protein).
Length = 116
Score = 104 bits (259), Expect = 2e-21
Identities = 47/88 (53%), Positives = 60/88 (68%)
Frame = -3
Query: 525 LAGPLISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYA 346
LAGP++ L YPLYASV AIE+ + VDD+QWL YW+ SF+TL EL ++EW+P W A
Sbjct: 14 LAGPVVMLLYPLYASVIAIESPSKVDDEQWLAYWILYSFLTLSELILQSLLEWIPIWYTA 73
Query: 345 KLFFNCXLVLPWFNGXAYVYDHFXRPMF 262
KL F LVLP F G A++Y+ R F
Sbjct: 74 KLVFVAWLVLPQFRGAAFIYNKVVREQF 101
>sptr|Q9AX25|Q9AX25 P0456A01.10 protein (P0042A10.34 protein).
Length = 128
Score = 102 bits (253), Expect = 8e-21
Identities = 54/118 (45%), Positives = 70/118 (59%), Gaps = 2/118 (1%)
Frame = -3
Query: 480 VRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFFNCXLVLPWFNG 301
+RAIE+ + +DDQQWLTYWV S ITLFEL+ +++W P W Y KL F C LVLP FNG
Sbjct: 1 MRAIESPSTLDDQQWLTYWVLYSLITLFELSCWKVLQWFPLWPYMKLLFCCWLVLPIFNG 60
Query: 300 XAYVYDHFXRPMFVNRQIVNIWYIXRNEKLGK--SDDVXXAAERYIEQNXXXAFXXLI 133
AY+Y+ R F Q V+ Y R K + S D + ER+IE + A +I
Sbjct: 61 AAYIYETHVRRYFKIGQYVSPNYNERQRKALQMMSLDARKSVERFIESHGPDALDKII 118
>sptr|Q9S760|Q9S760 ATHVA22D (Putative abscisic acid-induced
protein).
Length = 135
Score = 99.8 bits (247), Expect = 4e-20
Identities = 44/87 (50%), Positives = 57/87 (65%)
Frame = -3
Query: 522 AGPLISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAK 343
AGP++ L YPLYASV A+E+ VDD+QWL YW+ SF++L EL +IEW+P W K
Sbjct: 15 AGPIVMLLYPLYASVIAMESTTKVDDEQWLAYWIIYSFLSLTELILQSLIEWIPIWYTVK 74
Query: 342 LFFNCXLVLPWFNGXAYVYDHFXRPMF 262
L F LVLP F G A++Y+ R F
Sbjct: 75 LVFVAWLVLPQFQGAAFIYNRVVREQF 101
>sptr|Q07764|Q07764 Abscisic acid and stress inducible (A22) gene.
Length = 130
Score = 99.0 bits (245), Expect = 7e-20
Identities = 44/92 (47%), Positives = 59/92 (64%)
Frame = -3
Query: 546 LANNFDVLAGPLISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEW 367
L + +AGP I+L YPLYASV A+E+ + VDD+QWL YW+ SFITL E+ P++ W
Sbjct: 7 LLTHLHSVAGPSITLLYPLYASVCAMESPSKVDDEQWLAYWILYSFITLLEMVAEPVLYW 66
Query: 366 LPFWSYAKLFFNCXLVLPWFNGXAYVYDHFXR 271
+P W KL F L LP F G +++YD R
Sbjct: 67 IPVWYPVKLLFVAWLALPQFKGASFIYDKVVR 98
>sptrnew|BAC80265|BAC80265 Hypothetical protein Wrab15.
Length = 130
Score = 98.6 bits (244), Expect = 9e-20
Identities = 44/92 (47%), Positives = 59/92 (64%)
Frame = -3
Query: 546 LANNFDVLAGPLISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEW 367
L + +AGP I+L YPLYASV A+E+ + VDD+QWL YW+ SFITL E+ P++ W
Sbjct: 7 LLTHLHSVAGPSITLLYPLYASVCAMESPSKVDDEQWLAYWILYSFITLMEMLAEPVLYW 66
Query: 366 LPFWSYAKLFFNCXLVLPWFNGXAYVYDHFXR 271
+P W KL F L LP F G +++YD R
Sbjct: 67 IPVWYPVKLLFVAWLALPQFKGASFIYDKVVR 98
>sptr|Q9LUE9|Q9LUE9 Genomic DNA, chromosome 5, P1 clone:MFB16.
Length = 107
Score = 95.1 bits (235), Expect = 1e-18
Identities = 43/81 (53%), Positives = 54/81 (66%)
Frame = -3
Query: 504 LAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFFNCX 325
L YPLYASV AIE+ + VDD+QWL YW+ SF+TL EL ++EW+P W AKL F
Sbjct: 2 LLYPLYASVIAIESPSKVDDEQWLAYWILYSFLTLSELILQSLLEWIPIWYTAKLVFVAW 61
Query: 324 LVLPWFNGXAYVYDHFXRPMF 262
LVLP F G A++Y+ R F
Sbjct: 62 LVLPQFRGAAFIYNKVVREQF 82
>sptr|Q9SW30|Q9SW30 Abscisic acid-induced-like protein.
Length = 116
Score = 91.7 bits (226), Expect = 1e-17
Identities = 41/81 (50%), Positives = 52/81 (64%)
Frame = -3
Query: 504 LAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFFNCX 325
L YPLYASV A+E+ VDD+QWL YW+ SF++L EL +IEW+P W KL F
Sbjct: 2 LLYPLYASVIAMESTTKVDDEQWLAYWIIYSFLSLTELILQSLIEWIPIWYTVKLVFVAW 61
Query: 324 LVLPWFNGXAYVYDHFXRPMF 262
LVLP F G A++Y+ R F
Sbjct: 62 LVLPQFQGAAFIYNRVVREQF 82
>sptr|Q9UU91|Q9UU91 Putative transport protein.
Length = 182
Score = 70.9 bits (172), Expect = 2e-11
Identities = 34/92 (36%), Positives = 51/92 (55%)
Frame = -3
Query: 513 LISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFF 334
L++ A P + S+ AIET N DD QWLTY++ SF+ + E I+ ++P + K F
Sbjct: 62 LLAFAMPAFFSINAIETTNKADDTQWLTYYLVTSFLNVIEYWSQLILYYVPVYWLLKAIF 121
Query: 333 NCXLVLPWFNGXAYVYDHFXRPMFVNRQIVNI 238
L LP FNG +Y H RP ++ ++ I
Sbjct: 122 LIWLALPKFNGATIIYRHLIRP-YITPHVIRI 152
>sptr|Q96LM0|Q96LM0 Hypothetical protein FLJ25383 (TB2 protein-like
1).
Length = 184
Score = 67.0 bits (162), Expect = 3e-10
Identities = 34/87 (39%), Positives = 45/87 (51%), Gaps = 2/87 (2%)
Frame = -3
Query: 513 LISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFF 334
LI YP YAS++AIE+ + DD WLTYWV + L E ++ W PF+ K F
Sbjct: 60 LIGFVYPAYASIKAIESPSKDDDTVWLTYWVVYALFGLAEFFSDLLLSWFPFYYVGKCAF 119
Query: 333 --NCXLVLPWFNGXAYVYDHFXRPMFV 259
C PW NG +Y RP+F+
Sbjct: 120 LLFCMAPRPW-NGALMLYQRVVRPLFL 145
>sptr|Q96HR9|Q96HR9 Hypothetical protein.
Length = 184
Score = 67.0 bits (162), Expect = 3e-10
Identities = 34/87 (39%), Positives = 45/87 (51%), Gaps = 2/87 (2%)
Frame = -3
Query: 513 LISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFF 334
LI YP YAS++AIE+ + DD WLTYWV + L E ++ W PF+ K F
Sbjct: 60 LIGFVYPAYASIKAIESPSKDDDTVWLTYWVVYALFGLAEFFSDLLLSWFPFYYVGKCAF 119
Query: 333 --NCXLVLPWFNGXAYVYDHFXRPMFV 259
C PW NG +Y RP+F+
Sbjct: 120 LLFCMAPRPW-NGALMLYQRVVRPLFL 145
>sptr|O77388|O77388 Hypothetical 26.0 kDa protein.
Length = 221
Score = 66.6 bits (161), Expect = 4e-10
Identities = 29/88 (32%), Positives = 47/88 (53%)
Frame = -3
Query: 513 LISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFF 334
++ AYP Y S +A+E+++ + + WLTYWV S E I+ W+PF+ KL F
Sbjct: 95 VVGFAYPAYQSFKAVESQSRDETKLWLTYWVVFSLFFFIEYLIDIILFWVPFYYLIKLLF 154
Query: 333 NCXLVLPWFNGXAYVYDHFXRPMFVNRQ 250
L +P G VY++ RP+ + +
Sbjct: 155 LLYLYMPQVRGAVMVYNYIIRPILLKHE 182
>sptr|Q9N4G8|Q9N4G8 Hypothetical 20.6 kDa protein.
Length = 183
Score = 65.9 bits (159), Expect = 6e-10
Identities = 31/82 (37%), Positives = 44/82 (53%)
Frame = -3
Query: 510 ISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFFN 331
+ YP Y S++AIE+ N DD QWLTYWV + +++ E I+ P + K F
Sbjct: 67 MGFVYPAYMSIKAIESSNKEDDTQWLTYWVIFAILSVVEFFSVQIVAVFPVYWLFKSIFL 126
Query: 330 CXLVLPWFNGXAYVYDHFXRPM 265
L LP F G A +Y F +P+
Sbjct: 127 LYLYLPSFLGAAKLYHRFVKPV 148
>sptrnew|EAA31247|EAA31247 Hypothetical protein.
Length = 385
Score = 65.5 bits (158), Expect = 8e-10
Identities = 32/92 (34%), Positives = 48/92 (52%), Gaps = 3/92 (3%)
Frame = -3
Query: 534 FDVLAGPLISLA---YPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWL 364
FD+L L S+A +P+YAS +A+ + +P WL YWV SF+ L E + W+
Sbjct: 2 FDILPNLLSSVASFLFPVYASYKALRSSDPAQLTPWLMYWVVISFVVLVESWVGWFLVWV 61
Query: 363 PFWSYAKLFFNCXLVLPWFNGXAYVYDHFXRP 268
PF+++ + F LVLP G +Y P
Sbjct: 62 PFYAFMRFVFLLYLVLPQTQGARVIYQTHIEP 93
>sptr|Q8K0X2|Q8K0X2 Hypothetical protein.
Length = 174
Score = 64.7 bits (156), Expect = 1e-09
Identities = 34/91 (37%), Positives = 46/91 (50%), Gaps = 2/91 (2%)
Frame = -3
Query: 513 LISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFF 334
+I YP YASV+AIE+ + DD WLTYWV + L E ++ W PF+ K F
Sbjct: 60 VIGFVYPAYASVKAIESPSKEDDTVWLTYWVVYALFGLVEFFSDLLLFWFPFYYAGKCAF 119
Query: 333 --NCXLVLPWFNGXAYVYDHFXRPMFVNRQI 247
C PW NG +Y RP+F+ +
Sbjct: 120 LLFCMTPGPW-NGALLLYHRVIRPLFLKHHM 149
>sptr|Q9JM62|Q9JM62 Polyposis locus protein 1-like 1 (TB2
protein-like 1).
Length = 201
Score = 64.7 bits (156), Expect = 1e-09
Identities = 34/91 (37%), Positives = 46/91 (50%), Gaps = 2/91 (2%)
Frame = -3
Query: 513 LISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFF 334
+I YP YASV+AIE+ + DD WLTYWV + L E ++ W PF+ K F
Sbjct: 60 VIGFVYPAYASVKAIESPSKEDDTVWLTYWVVYALFGLVEFFSDLLLFWFPFYYAGKCAF 119
Query: 333 --NCXLVLPWFNGXAYVYDHFXRPMFVNRQI 247
C PW NG +Y RP+F+ +
Sbjct: 120 LLFCMTPGPW-NGALLLYHRVIRPLFLKHHM 149
>sptr|O45108|O45108 Hypothetical 22.8 kDa protein.
Length = 205
Score = 64.7 bits (156), Expect = 1e-09
Identities = 38/117 (32%), Positives = 53/117 (45%), Gaps = 2/117 (1%)
Frame = -3
Query: 564 GSFLKVLANNFDVLAGPLISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTF 385
GS ++L N LI YP YASV+AI + DD WL YW + + L +
Sbjct: 97 GSVARLLCN--------LIGFGYPTYASVKAIRSPGGDDDTVWLIYWTCFAVLYLVDFFS 148
Query: 384 APIIEWLPFWSYAKLFFNCXLVLPWFNGXAYVYDHFXRPM--FVNRQIVNIWYIXRN 220
I+ W PF+ AK F L LP G Y+ P+ FV++ + +Y N
Sbjct: 149 EAILSWFPFYYIAKACFLVYLYLPQTQGSVMFYETIVDPLVIFVDKNLDKFYYNRSN 205
>sw|Q60870|DP1_MOUSE Polyposis locus protein 1 homolog (TB2 protein
homolog) (GP106).
Length = 185
Score = 64.7 bits (156), Expect = 1e-09
Identities = 33/89 (37%), Positives = 45/89 (50%), Gaps = 1/89 (1%)
Frame = -3
Query: 513 LISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFF 334
LI YP Y S++AIE+ N DD QWLTYWV ++ E + WLPF+ K F
Sbjct: 57 LIGFGYPAYISMKAIESPNKDDDTQWLTYWVVYGVFSIAEFFSDLFLSWLPFYYMLKCGF 116
Query: 333 NCXLVLPW-FNGXAYVYDHFXRPMFVNRQ 250
+ P NG +Y RP+F+ +
Sbjct: 117 LLWCMAPSPANGAEMLYRRIIRPIFLRHE 145
>sw|Q00765|DP1_HUMAN Polyposis locus protein 1 (TB2 protein).
Length = 185
Score = 64.3 bits (155), Expect = 2e-09
Identities = 32/89 (35%), Positives = 43/89 (48%), Gaps = 1/89 (1%)
Frame = -3
Query: 513 LISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFF 334
LI YP Y S++AIE+ N DD QWLTYWV ++ E + W PF+ K F
Sbjct: 57 LIGFGYPAYISIKAIESPNKEDDTQWLTYWVVYGVFSIAEFFSDIFLSWFPFYYMLKCGF 116
Query: 333 NCXLVLPW-FNGXAYVYDHFXRPMFVNRQ 250
+ P NG +Y RP F+ +
Sbjct: 117 LLWCMAPSPSNGAELLYKRIIRPFFLKHE 145
>sptr|O93846|O93846 Pathogenicity protein.
Length = 369
Score = 63.5 bits (153), Expect = 3e-09
Identities = 32/92 (34%), Positives = 49/92 (53%), Gaps = 3/92 (3%)
Frame = -3
Query: 534 FDVLAGPLISLA---YPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWL 364
FD+ A L S+A +PL+AS +A++T +P WL YWV + L E + W+
Sbjct: 2 FDIFAKLLSSIASFLFPLFASYKALKTTDPAQLTPWLMYWVVLACALLVESWTEWFLCWI 61
Query: 363 PFWSYAKLFFNCXLVLPWFNGXAYVYDHFXRP 268
PF++Y +L F L+LP G +Y + P
Sbjct: 62 PFYAYIRLLFLLYLILPQTQGARVLYQDYIHP 93
>sptr|Q9LR09|Q9LR09 F10A5.11.
Length = 166
Score = 63.5 bits (153), Expect = 3e-09
Identities = 31/86 (36%), Positives = 41/86 (47%), Gaps = 2/86 (2%)
Frame = -3
Query: 501 AYPLYASVRAIETKNPVDDQQ--WLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFFNC 328
AYP Y + +E P Q W YW+ + +T+FE ++ WLP +S AKL F
Sbjct: 6 AYPAYECFKTVELNKPEIQQLQFWCQYWIIVAALTIFERIGDALVSWLPMYSEAKLAFFI 65
Query: 327 XLVLPWFNGXAYVYDHFXRPMFVNRQ 250
L P G YVYD F RP +
Sbjct: 66 YLWFPKTKGTTYVYDSFFRPYIAKHE 91
>sptr|Q9CQG4|Q9CQG4 Deleted in polyposis 1.
Length = 185
Score = 63.2 bits (152), Expect = 4e-09
Identities = 32/89 (35%), Positives = 44/89 (49%), Gaps = 1/89 (1%)
Frame = -3
Query: 513 LISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFF 334
LI YP Y S++AIE+ N DD QWLTYWV ++ E + W PF+ K F
Sbjct: 57 LIGFGYPAYISMKAIESPNKDDDTQWLTYWVVYGVFSIAEFFSDLFLSWFPFYYMLKCGF 116
Query: 333 NCXLVLPW-FNGXAYVYDHFXRPMFVNRQ 250
+ P NG +Y RP+F+ +
Sbjct: 117 LLWCMAPSPANGAEMLYRRIIRPIFLKHE 145
>sptrnew|EAA13329|EAA13329 ENSANGP00000003353 (Fragment).
Length = 143
Score = 62.8 bits (151), Expect = 5e-09
Identities = 30/85 (35%), Positives = 49/85 (57%), Gaps = 1/85 (1%)
Frame = -3
Query: 513 LISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFF 334
+I +AYP Y S++AIET+ DD +WLTYWV +++FE +++ +PF+ K F
Sbjct: 54 VIGVAYPAYISMKAIETRTKEDDTKWLTYWVTFGVLSVFEHFSFFLVQIIPFYWLLKCLF 113
Query: 333 NCXLVLPW-FNGXAYVYDHFXRPMF 262
+ ++P NG +Y +P F
Sbjct: 114 HIWCMVPMENNGSTIMYHKVIQPYF 138
>sptrnew|EAA05239|EAA05239 ENSANGP00000018512 (Fragment).
Length = 321
Score = 62.8 bits (151), Expect = 5e-09
Identities = 32/100 (32%), Positives = 49/100 (49%)
Frame = -3
Query: 549 VLANNFDVLAGPLISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIE 370
+L F +L G L YP YAS +A+ TKN + +W+ YW+ +F T E ++
Sbjct: 69 ILFPGFRLLFGTL----YPAYASYKAVRTKNVKEYVKWMMYWIVFAFFTFIETFTDILLS 124
Query: 369 WLPFWSYAKLFFNCXLVLPWFNGXAYVYDHFXRPMFVNRQ 250
W PF+ K+ L+ P G + +Y F PM R+
Sbjct: 125 WFPFYYEIKVIVVLWLLSPATRGSSTLYRKFVHPMLTRRE 164
>sptrnew|AAH06218|AAH06218 Hypothetical protein.
Length = 252
Score = 62.8 bits (151), Expect = 5e-09
Identities = 26/83 (31%), Positives = 45/83 (54%)
Frame = -3
Query: 498 YPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFFNCXLV 319
YP Y+S +A++TKN + +W+ YW+ +F T E ++ W PF+ K+ F L+
Sbjct: 18 YPAYSSYKAVKTKNVKEYVKWMMYWIVFAFFTTAETLTDIVLSWFPFYFELKIAFVIWLL 77
Query: 318 LPWFNGXAYVYDHFXRPMFVNRQ 250
P+ G + +Y F P N++
Sbjct: 78 SPYTKGSSVLYRKFVHPTLSNKE 100
>sptrnew|EAA13321|EAA13321 AgCP2771 (Fragment).
Length = 164
Score = 62.8 bits (151), Expect = 5e-09
Identities = 30/85 (35%), Positives = 49/85 (57%), Gaps = 1/85 (1%)
Frame = -3
Query: 513 LISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFF 334
+I +AYP Y S++AIET+ DD +WLTYWV +++FE +++ +PF+ K F
Sbjct: 48 VIGVAYPAYISMKAIETRTKEDDTKWLTYWVTFGVLSVFEHFSFFLVQIIPFYWLLKCLF 107
Query: 333 NCXLVLPW-FNGXAYVYDHFXRPMF 262
+ ++P NG +Y +P F
Sbjct: 108 HIWCMVPMENNGSTIMYHKVIQPYF 132
>sptr|Q921E4|Q921E4 Deleted in polyposis 1.
Length = 185
Score = 61.6 bits (148), Expect = 1e-08
Identities = 32/89 (35%), Positives = 43/89 (48%), Gaps = 1/89 (1%)
Frame = -3
Query: 513 LISLAYPLYASVRAIETKNPVDDQQWLTYWVXXSFITLFELTFAPIIEWLPFWSYAKLFF 334
LI YP Y S++AIE+ N DD QWLTYWV ++ E + W PF+ K F
Sbjct: 57 LIGFGYPAYISMKAIESPNKDDDTQWLTYWVVYGVFSIAEFFSDLFLSWFPFYYMLKCGF 116
Query: 333 NCXLVLPW-FNGXAYVYDHFXRPMFVNRQ 250
+ P NG Y RP+F+ +
Sbjct: 117 LLWCMAPSPANGAEMRYRRIIRPIFLKHE 145
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 464,394,353
Number of Sequences: 1272877
Number of extensions: 9126071
Number of successful extensions: 38069
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 34505
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 37757
length of database: 406,886,216
effective HSP length: 121
effective length of database: 252,868,099
effective search space used: 38688819147
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

