BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
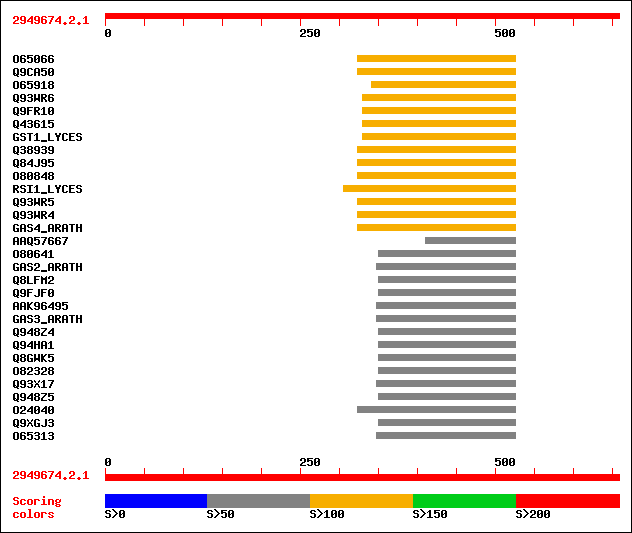
Query= 2949674.2.1
(658 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|O65066|O65066 GASA5-like protein. 134 1e-30
sptr|Q9CA50|Q9CA50 GAST1-like protein. 133 2e-30
sptr|O65918|O65918 GASA5-like protein (Fragment). 128 7e-29
sptr|Q93WR6|Q93WR6 Gibberellin induced protein 3. 126 2e-28
sptr|Q9FR10|Q9FR10 Gibberellin-induced protein 1 (Putative gibbe... 126 2e-28
sptr|Q43615|Q43615 Gip1 protein. 126 2e-28
sw|P27057|GST1_LYCES GAST1 protein precursor. 126 2e-28
sptr|Q38939|Q38939 GASA5. 125 3e-28
sptr|Q84J95|Q84J95 Hypothetical protein. 125 5e-28
sptr|O80848|O80848 Putative gibberellin-regulated protein. 121 7e-27
sw|P47926|RSI1_LYCES RSI-1 protein precursor (TR132). 121 9e-27
sptr|Q93WR5|Q93WR5 Gip1-like protein. 117 1e-25
sptr|Q93WR4|Q93WR4 Gip1-like protein. 116 2e-25
sw|P46690|GAS4_ARATH Gibberellin-regulated protein 4 precursor. 108 4e-23
sptrnew|AAQ57667|AAQ57667 GIP1-like protein (Fragment). 86 5e-16
sptr|O80641|O80641 GAST1/GASA-like protein. 82 6e-15
sw|P46688|GAS2_ARATH Gibberellin-regulated protein 2 precursor. 80 3e-14
sptr|Q8LFM2|Q8LFM2 Contains similarity to gibberellin-stimulated... 79 5e-14
sptr|Q9FJF0|Q9FJF0 Similarity to gibberellin-stimulated transcri... 79 5e-14
sptrnew|AAK96495|AAK96495 AT4g09600/T25P22_40. 76 3e-13
sw|P46687|GAS3_ARATH Gibberellin-regulated protein 3 precursor. 76 3e-13
sptr|Q948Z4|Q948Z4 Snakin-1. 75 9e-13
sptr|Q94HA1|Q94HA1 Hypothetical 9.9 kDa protein. 75 9e-13
sptr|Q8GWK5|Q8GWK5 Putative gibberellin-regulated protein. 74 2e-12
sptr|O82328|O82328 GAST1/GASA-like protein (Putative gibberellin... 74 2e-12
sptr|Q93X17|Q93X17 Snakin2 precursor. 73 3e-12
sptr|Q948Z5|Q948Z5 Snakin2 precursor. 72 6e-12
sptr|O24040|O24040 LTCOR11. 71 1e-11
sptr|Q9XGJ3|Q9XGJ3 GEG protein. 71 1e-11
sptr|O65313|O65313 Cold-regulated LTCOR12. 70 3e-11
>sptr|O65066|O65066 GASA5-like protein.
Length = 110
Score = 134 bits (337), Expect = 1e-30
Identities = 51/68 (75%), Positives = 61/68 (89%)
Frame = +2
Query: 323 GTGSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKT 502
G GSL+P ECG RC+ RCSAT+++KPC+FFC+KCCA CLCVPPGT+GNK CPCYNNWKT
Sbjct: 43 GEGSLRPSECGQRCSYRCSATSHKKPCMFFCQKCCAKCLCVPPGTFGNKQVCPCYNNWKT 102
Query: 503 KRGGPKCP 526
++GGPKCP
Sbjct: 103 QQGGPKCP 110
>sptr|Q9CA50|Q9CA50 GAST1-like protein.
Length = 80
Score = 133 bits (334), Expect = 2e-30
Identities = 50/68 (73%), Positives = 58/68 (85%)
Frame = +2
Query: 323 GTGSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKT 502
G GSL+ ++CGG+C RCS T Y KPC+FFC+KCCA CLCVPPGTYGNK CPCYNNWKT
Sbjct: 13 GPGSLKSYQCGGQCTRRCSNTKYHKPCMFFCQKCCAKCLCVPPGTYGNKQVCPCYNNWKT 72
Query: 503 KRGGPKCP 526
++GGPKCP
Sbjct: 73 QQGGPKCP 80
>sptr|O65918|O65918 GASA5-like protein (Fragment).
Length = 62
Score = 128 bits (321), Expect = 7e-29
Identities = 48/62 (77%), Positives = 56/62 (90%)
Frame = +2
Query: 341 PHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPK 520
P ECG RC+ RCSAT+++KPC+FFC+KCCA CLCVPPGTYGNK CPCYNNWKT++GGPK
Sbjct: 1 PSECGQRCSYRCSATSHKKPCMFFCQKCCAKCLCVPPGTYGNKQVCPCYNNWKTQQGGPK 60
Query: 521 CP 526
CP
Sbjct: 61 CP 62
>sptr|Q93WR6|Q93WR6 Gibberellin induced protein 3.
Length = 112
Score = 126 bits (317), Expect = 2e-28
Identities = 48/66 (72%), Positives = 55/66 (83%)
Frame = +2
Query: 329 GSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKR 508
GSL P +C +C RCS T+++KPC+FFC+KCCA CLCVP GTYGNK TCPCYNNWKTK
Sbjct: 47 GSLHPQDCQPKCTYRCSKTSFKKPCMFFCQKCCAKCLCVPAGTYGNKQTCPCYNNWKTKE 106
Query: 509 GGPKCP 526
GGPKCP
Sbjct: 107 GGPKCP 112
>sptr|Q9FR10|Q9FR10 Gibberellin-induced protein 1 (Putative
gibberellin induced protein 2).
Length = 112
Score = 126 bits (317), Expect = 2e-28
Identities = 48/66 (72%), Positives = 55/66 (83%)
Frame = +2
Query: 329 GSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKR 508
GSL P +C +C RCS T+++KPC+FFC+KCCA CLCVP GTYGNK TCPCYNNWKTK
Sbjct: 47 GSLHPQDCQPKCTYRCSKTSFKKPCMFFCQKCCAKCLCVPAGTYGNKQTCPCYNNWKTKE 106
Query: 509 GGPKCP 526
GGPKCP
Sbjct: 107 GGPKCP 112
>sptr|Q43615|Q43615 Gip1 protein.
Length = 112
Score = 126 bits (317), Expect = 2e-28
Identities = 48/66 (72%), Positives = 55/66 (83%)
Frame = +2
Query: 329 GSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKR 508
GSL P +C +C RCS T+++KPC+FFC+KCCA CLCVP GTYGNK TCPCYNNWKTK
Sbjct: 47 GSLHPQDCQPKCTYRCSKTSFKKPCMFFCQKCCAKCLCVPAGTYGNKQTCPCYNNWKTKE 106
Query: 509 GGPKCP 526
GGPKCP
Sbjct: 107 GGPKCP 112
>sw|P27057|GST1_LYCES GAST1 protein precursor.
Length = 112
Score = 126 bits (317), Expect = 2e-28
Identities = 48/66 (72%), Positives = 55/66 (83%)
Frame = +2
Query: 329 GSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKR 508
G L P +C +C RCS T+Y+KPC+FFC+KCCA CLCVP GTYGNK +CPCYNNWKTKR
Sbjct: 47 GRLHPQDCQPKCTYRCSKTSYKKPCMFFCQKCCAKCLCVPAGTYGNKQSCPCYNNWKTKR 106
Query: 509 GGPKCP 526
GGPKCP
Sbjct: 107 GGPKCP 112
>sptr|Q38939|Q38939 GASA5.
Length = 97
Score = 125 bits (315), Expect = 3e-28
Identities = 47/68 (69%), Positives = 56/68 (82%)
Frame = +2
Query: 323 GTGSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKT 502
G G L+P +C +C+ RCSAT+++KPC+FFC KCC CLCVPPGT+GNK TCPCYNNWKT
Sbjct: 30 GGGKLKPQQCNSKCSYRCSATSHKKPCMFFCLKCCKKCLCVPPGTFGNKQTCPCYNNWKT 89
Query: 503 KRGGPKCP 526
K G PKCP
Sbjct: 90 KEGRPKCP 97
>sptr|Q84J95|Q84J95 Hypothetical protein.
Length = 97
Score = 125 bits (314), Expect = 5e-28
Identities = 47/68 (69%), Positives = 56/68 (82%)
Frame = +2
Query: 323 GTGSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKT 502
G G L+P +C +C+ RCSAT+++KPC+FFC KCC CLCVPPGT+GNK TCPCYNNWKT
Sbjct: 30 GGGKLKPQQCNSKCSFRCSATSHKKPCMFFCLKCCKKCLCVPPGTFGNKQTCPCYNNWKT 89
Query: 503 KRGGPKCP 526
K G PKCP
Sbjct: 90 KEGRPKCP 97
>sptr|O80848|O80848 Putative gibberellin-regulated protein.
Length = 103
Score = 121 bits (304), Expect = 7e-27
Identities = 48/68 (70%), Positives = 53/68 (77%)
Frame = +2
Query: 323 GTGSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKT 502
G GSL+P EC C RCSAT+++KPCLFFC KCC CLCVP GTYG+K CPCYNNW T
Sbjct: 36 GEGSLKPEECPKACEYRCSATSHRKPCLFFCNKCCNKCLCVPSGTYGHKEECPCYNNWTT 95
Query: 503 KRGGPKCP 526
K GGPKCP
Sbjct: 96 KEGGPKCP 103
>sw|P47926|RSI1_LYCES RSI-1 protein precursor (TR132).
Length = 96
Score = 121 bits (303), Expect = 9e-27
Identities = 46/74 (62%), Positives = 56/74 (75%)
Frame = +2
Query: 305 YGGVTPGTGSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPC 484
+ V G L+P +C RC RCSAT+++KPC+FFC+KCCA CLCVP G YGNK +CPC
Sbjct: 23 FSNVVEGYNKLRPTDCKPRCTYRCSATSHKKPCMFFCQKCCATCLCVPKGVYGNKQSCPC 82
Query: 485 YNNWKTKRGGPKCP 526
YNNWKT+ G PKCP
Sbjct: 83 YNNWKTQEGKPKCP 96
>sptr|Q93WR5|Q93WR5 Gip1-like protein.
Length = 105
Score = 117 bits (293), Expect = 1e-25
Identities = 45/68 (66%), Positives = 50/68 (73%)
Frame = +2
Query: 323 GTGSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKT 502
G GSL+P +C +C RCS T Y C+ FC+KCC CLCVPPG YGNK CPCYNNWKT
Sbjct: 38 GPGSLKPSQCLPQCTRRCSQTQYHNACMLFCQKCCNKCLCVPPGFYGNKGVCPCYNNWKT 97
Query: 503 KRGGPKCP 526
K GGPKCP
Sbjct: 98 KEGGPKCP 105
>sptr|Q93WR4|Q93WR4 Gip1-like protein.
Length = 104
Score = 116 bits (291), Expect = 2e-25
Identities = 45/68 (66%), Positives = 50/68 (73%)
Frame = +2
Query: 323 GTGSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKT 502
G GSL+P +C +C RCS T Y C+ FC+KCC CLCVPPG YGNK CPCYNNWKT
Sbjct: 37 GPGSLKPTQCLPQCIRRCSHTQYHNACMLFCQKCCKKCLCVPPGFYGNKGVCPCYNNWKT 96
Query: 503 KRGGPKCP 526
K GGPKCP
Sbjct: 97 KEGGPKCP 104
>sw|P46690|GAS4_ARATH Gibberellin-regulated protein 4 precursor.
Length = 106
Score = 108 bits (271), Expect = 4e-23
Identities = 42/68 (61%), Positives = 46/68 (67%)
Frame = +2
Query: 323 GTGSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKT 502
G GSL+ +C C RC T Y K C+ FC KCC CLCVPPG YGNK C CYNNWKT
Sbjct: 39 GPGSLKRTQCPSECDRRCKKTQYHKACITFCNKCCRKCLCVPPGYYGNKQVCSCYNNWKT 98
Query: 503 KRGGPKCP 526
+ GGPKCP
Sbjct: 99 QEGGPKCP 106
>sptrnew|AAQ57667|AAQ57667 GIP1-like protein (Fragment).
Length = 39
Score = 85.5 bits (210), Expect = 5e-16
Identities = 31/39 (79%), Positives = 34/39 (87%)
Frame = +2
Query: 410 FCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
FC+KCC CLCVPPG YGNK CPCYNNWKT++GGPKCP
Sbjct: 1 FCQKCCRKCLCVPPGFYGNKAVCPCYNNWKTQQGGPKCP 39
>sptr|O80641|O80641 GAST1/GASA-like protein.
Length = 87
Score = 82.0 bits (201), Expect = 6e-15
Identities = 30/59 (50%), Positives = 38/59 (64%)
Frame = +2
Query: 350 CGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
CGG+C RCS + CL +C CC C CVP GT+G+K+ CPCY + K +GG KCP
Sbjct: 29 CGGKCNVRCSKAGQHEECLKYCNICCQKCNCVPSGTFGHKDECPCYRDMKNSKGGSKCP 87
>sw|P46688|GAS2_ARATH Gibberellin-regulated protein 2 precursor.
Length = 99
Score = 79.7 bits (195), Expect = 3e-14
Identities = 32/60 (53%), Positives = 39/60 (65%)
Frame = +2
Query: 347 ECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
+CGGRC +RCS ++ K CL C CC+ C CVPPGT GN + CPCY + T G KCP
Sbjct: 40 DCGGRCKDRCSKSSRTKLCLRACNSCCSRCNCVPPGTSGNTHLCPCYASITTHGGRLKCP 99
>sptr|Q8LFM2|Q8LFM2 Contains similarity to gibberellin-stimulated
transcript 1 like protein (Hypothetical protein).
Length = 89
Score = 79.0 bits (193), Expect = 5e-14
Identities = 32/60 (53%), Positives = 37/60 (61%), Gaps = 1/60 (1%)
Frame = +2
Query: 350 CGGRCAERCSATAYQKPCLFFCRKCCAAC-LCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
CGG+C RCS Q CL +C CC C CVP GTYGNK+ CPCY + K +G KCP
Sbjct: 30 CGGKCNVRCSKAGRQDRCLKYCNICCEKCNYCVPSGTYGNKDECPCYRDMKNSKGTSKCP 89
>sptr|Q9FJF0|Q9FJF0 Similarity to gibberellin-stimulated transcript
1 like protein.
Length = 88
Score = 79.0 bits (193), Expect = 5e-14
Identities = 32/60 (53%), Positives = 37/60 (61%), Gaps = 1/60 (1%)
Frame = +2
Query: 350 CGGRCAERCSATAYQKPCLFFCRKCCAAC-LCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
CGG+C RCS Q CL +C CC C CVP GTYGNK+ CPCY + K +G KCP
Sbjct: 29 CGGKCNVRCSKAGRQDRCLKYCNICCEKCNYCVPSGTYGNKDECPCYRDMKNSKGTSKCP 88
>sptrnew|AAK96495|AAK96495 AT4g09600/T25P22_40.
Length = 99
Score = 76.3 bits (186), Expect = 3e-13
Identities = 31/60 (51%), Positives = 37/60 (61%)
Frame = +2
Query: 347 ECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
+CGGRC RCS ++ CL C CC C CVPPGT GN + CPCY + T+ G KCP
Sbjct: 40 DCGGRCKGRCSKSSRPNLCLRACNSCCYRCNCVPPGTAGNHHLCPCYASITTRGGRLKCP 99
>sw|P46687|GAS3_ARATH Gibberellin-regulated protein 3 precursor.
Length = 99
Score = 76.3 bits (186), Expect = 3e-13
Identities = 31/60 (51%), Positives = 37/60 (61%)
Frame = +2
Query: 347 ECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
+CGGRC RCS ++ CL C CC C CVPPGT GN + CPCY + T+ G KCP
Sbjct: 40 DCGGRCKGRCSKSSRPNLCLRACNSCCYRCNCVPPGTAGNHHLCPCYASITTRGGRLKCP 99
>sptr|Q948Z4|Q948Z4 Snakin-1.
Length = 88
Score = 74.7 bits (182), Expect = 9e-13
Identities = 29/59 (49%), Positives = 34/59 (57%)
Frame = +2
Query: 350 CGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
C +C RCS CL +C CC C CVP GTYGNK+ CPCY + K +G KCP
Sbjct: 30 CDSKCKLRCSKAGLADRCLKYCGICCEECKCVPSGTYGNKHECPCYRDKKNSKGKSKCP 88
>sptr|Q94HA1|Q94HA1 Hypothetical 9.9 kDa protein.
Length = 93
Score = 74.7 bits (182), Expect = 9e-13
Identities = 31/62 (50%), Positives = 37/62 (59%), Gaps = 3/62 (4%)
Frame = +2
Query: 350 CGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGG---PK 520
C G+C RCS + CL +C CCA+C CVP GT GNK+ CPCY + T G PK
Sbjct: 32 CDGKCKVRCSKASRHDDCLKYCGVCCASCNCVPSGTAGNKDECPCYRDMTTGHGARKRPK 91
Query: 521 CP 526
CP
Sbjct: 92 CP 93
>sptr|Q8GWK5|Q8GWK5 Putative gibberellin-regulated protein.
Length = 119
Score = 73.9 bits (180), Expect = 2e-12
Identities = 31/59 (52%), Positives = 36/59 (61%)
Frame = +2
Query: 350 CGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
CG CA RCS T+ +K C C CCA C CVPPGT GN +CPCY + +T KCP
Sbjct: 61 CGHACARRCSKTSRKKVCHRACGSCCAKCQCVPPGTSGNTASCPCYASIRTHGNKLKCP 119
>sptr|O82328|O82328 GAST1/GASA-like protein (Putative
gibberellin-regulated protein).
Length = 108
Score = 73.6 bits (179), Expect = 2e-12
Identities = 28/59 (47%), Positives = 35/59 (59%)
Frame = +2
Query: 350 CGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
CG +C RC + CL +C CC C CVP GTYGNK+ C CY + + +G PKCP
Sbjct: 50 CGQKCEGRCKEAGMKDRCLKYCGICCKDCQCVPSGTYGNKHECACYRDKLSSKGTPKCP 108
>sptr|Q93X17|Q93X17 Snakin2 precursor.
Length = 104
Score = 72.8 bits (177), Expect = 3e-12
Identities = 30/60 (50%), Positives = 35/60 (58%)
Frame = +2
Query: 347 ECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
+CGG CA RC ++ + C C CCA C CVPPGT GN TCPCY + T KCP
Sbjct: 45 DCGGACAARCRLSSRPRLCNRACGTCCARCNCVPPGTSGNTETCPCYASLTTHGNKRKCP 104
>sptr|Q948Z5|Q948Z5 Snakin2 precursor.
Length = 104
Score = 72.0 bits (175), Expect = 6e-12
Identities = 30/59 (50%), Positives = 34/59 (57%)
Frame = +2
Query: 350 CGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
CGG CA RC ++ + C C CCA C CVPPGT GN TCPCY + T KCP
Sbjct: 46 CGGACAARCRLSSRPRLCNRACGTCCARCNCVPPGTSGNTETCPCYASLTTHGNKRKCP 104
>sptr|O24040|O24040 LTCOR11.
Length = 102
Score = 70.9 bits (172), Expect = 1e-11
Identities = 32/69 (46%), Positives = 38/69 (55%), Gaps = 1/69 (1%)
Frame = +2
Query: 323 GTGSLQPHECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCP-CYNNWK 499
G S + +CGG CA RC ++ C C CCA C CVPPGT GN+ CP CY +
Sbjct: 34 GNSSPKKIDCGGACAARCQLSSRPHLCKRACGTCCARCACVPPGTAGNQEMCPKCYASLT 93
Query: 500 TKRGGPKCP 526
T G KCP
Sbjct: 94 THGGKRKCP 102
>sptr|Q9XGJ3|Q9XGJ3 GEG protein.
Length = 101
Score = 70.9 bits (172), Expect = 1e-11
Identities = 29/59 (49%), Positives = 32/59 (54%)
Frame = +2
Query: 350 CGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
CG C RC ++ C C CCA C CVPPGT GN+ CPCY N T G KCP
Sbjct: 43 CGAACKARCRLSSRPNLCHRACGTCCARCRCVPPGTSGNQKVCPCYYNMTTHGGRRKCP 101
>sptr|O65313|O65313 Cold-regulated LTCOR12.
Length = 101
Score = 69.7 bits (169), Expect = 3e-11
Identities = 29/60 (48%), Positives = 34/60 (56%)
Frame = +2
Query: 347 ECGGRCAERCSATAYQKPCLFFCRKCCAACLCVPPGTYGNKNTCPCYNNWKTKRGGPKCP 526
+CGG CA RC ++ C C CCA CVPPGT GN+ CPCY + T G KCP
Sbjct: 42 DCGGACAARCQLSSRPHLCKRACGTCCARSRCVPPGTAGNQEMCPCYASLTTHGGKRKCP 101
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 392,736,201
Number of Sequences: 1272877
Number of extensions: 7446520
Number of successful extensions: 24688
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 22877
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 24499
length of database: 406,886,216
effective HSP length: 118
effective length of database: 256,686,730
effective search space used: 25668673000
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

