BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= 2907809.2.1
(614 letters)
Database: /db/trembl-ebi/tmp/swall
1,288,562 sequences; 411,996,502 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
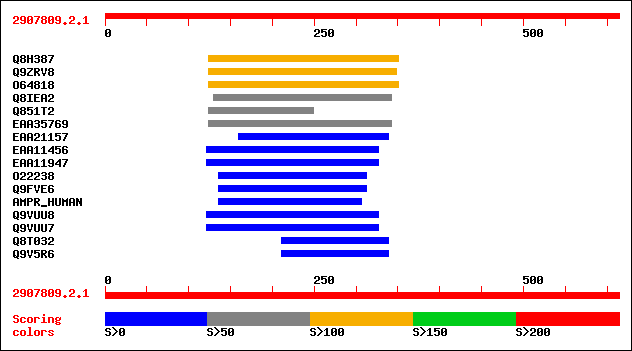
sptr|Q8H387|Q8H387 OJ1513_F02.31 protein. 147 1e-34
sptr|Q9ZRV8|Q9ZRV8 Hypothetical protein. 131 7e-30
sptr|O64818|O64818 At2g23090 protein (At2g23090/F21P24.15) (Hypo... 130 2e-29
sptr|Q8IEA2|Q8IEA2 Hypothetical protein, conserved. 69 4e-11
sptr|Q851T2|Q851T2 Hypothetical protein. 69 6e-11
sptrnew|EAA35769|EAA35769 Predicted protein. 67 2e-10
sptrnew|EAA21157|EAA21157 Hypothetical protein. 50 3e-05
sptrnew|EAA11456|EAA11456 EbiP7255 (Fragment). 38 0.085
sptrnew|EAA11947|EAA11947 ENSANGP00000018311 (Fragment). 36 0.32
sptr|O22238|O22238 Putative histone deacetylase. 33 3.6
sptr|Q9FVE6|Q9FVE6 Putative histone deacetylase. 33 3.6
sw|P15514|AMPR_HUMAN Amphiregulin precursor (AR) (Colorectum cel... 32 6.1
sptr|Q9VUU8|Q9VUU8 CG15715 protein. 32 7.9
sptr|Q9VUU7|Q9VUU7 CG18081 protein (RE63284p). 32 7.9
sptr|Q8T032|Q8T032 LD35343p. 32 7.9
sptr|Q9V5R6|Q9V5R6 CG30020 protein (LD40262p). 32 7.9
>sptr|Q8H387|Q8H387 OJ1513_F02.31 protein.
Length = 77
Score = 147 bits (371), Expect = 1e-34
Identities = 68/76 (89%), Positives = 70/76 (92%)
Frame = +1
Query: 124 MGGGNAQKSKMARERNAEKNKGGKGSQLEANKKAMNIQCKICMQTFICTTSEAKCKEHAE 303
MGGGN QKSKMARERN EKNKG KGSQLEANKKAMNIQCKICMQTFICTTSE KCKEHAE
Sbjct: 1 MGGGNGQKSKMARERNMEKNKGAKGSQLEANKKAMNIQCKICMQTFICTTSETKCKEHAE 60
Query: 304 ARHPKNDLYQCFPHLK 351
A+HPK+DL CFPHLK
Sbjct: 61 AKHPKSDLTACFPHLK 76
>sptr|Q9ZRV8|Q9ZRV8 Hypothetical protein.
Length = 78
Score = 131 bits (329), Expect = 7e-30
Identities = 60/76 (78%), Positives = 67/76 (88%), Gaps = 1/76 (1%)
Frame = +1
Query: 124 MGGGNAQKSKMARERNAEKNKG-GKGSQLEANKKAMNIQCKICMQTFICTTSEAKCKEHA 300
MGGGN QK+KMARERN EK K GKGSQLE NKKAM+IQCK+CMQTFICTTSE KC+EHA
Sbjct: 1 MGGGNGQKAKMARERNLEKQKNAGKGSQLETNKKAMSIQCKVCMQTFICTTSEVKCREHA 60
Query: 301 EARHPKNDLYQCFPHL 348
EA+HPK+D+ CFPHL
Sbjct: 61 EAKHPKSDVLVCFPHL 76
>sptr|O64818|O64818 At2g23090 protein (At2g23090/F21P24.15)
(Hypothetical protein).
Length = 78
Score = 130 bits (326), Expect = 2e-29
Identities = 61/77 (79%), Positives = 67/77 (87%), Gaps = 1/77 (1%)
Frame = +1
Query: 124 MGGGNAQKSKMARERNAEKNKG-GKGSQLEANKKAMNIQCKICMQTFICTTSEAKCKEHA 300
MGGGNAQKS MAR +N EK K GKGSQLEANKKAM+IQCK+CMQTFICTTSE KC+EHA
Sbjct: 1 MGGGNAQKSAMARAKNLEKAKAAGKGSQLEANKKAMSIQCKVCMQTFICTTSEVKCREHA 60
Query: 301 EARHPKNDLYQCFPHLK 351
EA+HPK D+ CFPHLK
Sbjct: 61 EAKHPKADVVACFPHLK 77
>sptr|Q8IEA2|Q8IEA2 Hypothetical protein, conserved.
Length = 78
Score = 68.9 bits (167), Expect = 4e-11
Identities = 36/71 (50%), Positives = 48/71 (67%)
Frame = +1
Query: 130 GGNAQKSKMARERNAEKNKGGKGSQLEANKKAMNIQCKICMQTFICTTSEAKCKEHAEAR 309
G NA K ARER + KGGK SQL+ANK+A+N+ CK+C F+ T S ++ EHAE +
Sbjct: 4 GSNACKRNQARERKNVEVKGGK-SQLKANKEALNVTCKVCYTVFMQTQSISQLAEHAENK 62
Query: 310 HPKNDLYQCFP 342
H K D+ +CFP
Sbjct: 63 HNK-DVKECFP 72
>sptr|Q851T2|Q851T2 Hypothetical protein.
Length = 42
Score = 68.6 bits (166), Expect = 6e-11
Identities = 33/42 (78%), Positives = 34/42 (80%)
Frame = +1
Query: 124 MGGGNAQKSKMARERNAEKNKGGKGSQLEANKKAMNIQCKIC 249
MGGGN QKS+MARERN EK KG KGSQLE NKKAMNIQ C
Sbjct: 1 MGGGNGQKSRMARERNMEKAKGAKGSQLETNKKAMNIQVGSC 42
>sptrnew|EAA35769|EAA35769 Predicted protein.
Length = 80
Score = 67.0 bits (162), Expect = 2e-10
Identities = 34/73 (46%), Positives = 42/73 (57%)
Frame = +1
Query: 124 MGGGNAQKSKMARERNAEKNKGGKGSQLEANKKAMNIQCKICMQTFICTTSEAKCKEHAE 303
MGGGN K+ RERNA+ G SQL+ N AMNI C+ C TF+ T+ EHA+
Sbjct: 1 MGGGNGAKAAQKRERNAKNAAAGPKSQLKTNAAAMNIICQTCRATFLSTSRAKALDEHAQ 60
Query: 304 ARHPKNDLYQCFP 342
+H K L CFP
Sbjct: 61 NKHNKT-LADCFP 72
>sptrnew|EAA21157|EAA21157 Hypothetical protein.
Length = 71
Score = 49.7 bits (117), Expect = 3e-05
Identities = 22/61 (36%), Positives = 38/61 (62%), Gaps = 1/61 (1%)
Frame = +1
Query: 160 RERNAEKNKGGK-GSQLEANKKAMNIQCKICMQTFICTTSEAKCKEHAEARHPKNDLYQC 336
R R++++N+ K S ++ K +N+ CK+C TF+CT +++ KEH E +H K+ C
Sbjct: 11 RIRSSKRNEKAKPNSTVKDASKNLNVICKLCRHTFMCTVNQSILKEHHEKKHSKHAYEDC 70
Query: 337 F 339
F
Sbjct: 71 F 71
>sptrnew|EAA11456|EAA11456 EbiP7255 (Fragment).
Length = 77
Score = 38.1 bits (87), Expect = 0.085
Identities = 23/72 (31%), Positives = 34/72 (47%), Gaps = 3/72 (4%)
Frame = +1
Query: 121 AMGGGNAQKSKMARERNAEKNKG---GKGSQLEANKKAMNIQCKICMQTFICTTSEAKCK 291
A G Q + A+E+ A+ K G Q++A +KA+ C +C K
Sbjct: 2 ARGHQKIQSQQKAQEKAAKMKKQQGHGLSDQMKAAQKALVFSCAVCKSQM---PDPKTYK 58
Query: 292 EHAEARHPKNDL 327
+H E +HPKNDL
Sbjct: 59 QHFENKHPKNDL 70
>sptrnew|EAA11947|EAA11947 ENSANGP00000018311 (Fragment).
Length = 78
Score = 36.2 bits (82), Expect = 0.32
Identities = 22/72 (30%), Positives = 34/72 (47%), Gaps = 3/72 (4%)
Frame = +1
Query: 121 AMGGGNAQKSKMARERNAEKNKG---GKGSQLEANKKAMNIQCKICMQTFICTTSEAKCK 291
A G Q + A+E+ A+ K G Q++A +KA+ C +C K
Sbjct: 3 ARGHQKIQSQQKAQEKAAKLKKQQGHGLSDQMKAAQKALVFSCAVCKSQM---PDPKTYK 59
Query: 292 EHAEARHPKNDL 327
+H E +HPKN+L
Sbjct: 60 QHFENKHPKNEL 71
>sptr|O22238|O22238 Putative histone deacetylase.
Length = 257
Score = 32.7 bits (73), Expect = 3.6
Identities = 20/61 (32%), Positives = 34/61 (55%), Gaps = 2/61 (3%)
Frame = +1
Query: 136 NAQKSKMA--RERNAEKNKGGKGSQLEANKKAMNIQCKICMQTFICTTSEAKCKEHAEAR 309
+A+K+K+A ++ EK KGGK + ++ K A + C C +TF S + H +A+
Sbjct: 185 SAKKAKVAVTPQKTDEKKKGGKAAN-QSPKSASQVSCGSCKKTF---NSGNALESHNKAK 240
Query: 310 H 312
H
Sbjct: 241 H 241
>sptr|Q9FVE6|Q9FVE6 Putative histone deacetylase.
Length = 245
Score = 32.7 bits (73), Expect = 3.6
Identities = 20/61 (32%), Positives = 34/61 (55%), Gaps = 2/61 (3%)
Frame = +1
Query: 136 NAQKSKMA--RERNAEKNKGGKGSQLEANKKAMNIQCKICMQTFICTTSEAKCKEHAEAR 309
+A+K+K+A ++ EK KGGK + ++ K A + C C +TF S + H +A+
Sbjct: 185 SAKKAKVAVTPQKTDEKKKGGKAAN-QSPKSASQVSCGSCKKTF---NSGNALESHNKAK 240
Query: 310 H 312
H
Sbjct: 241 H 241
>sw|P15514|AMPR_HUMAN Amphiregulin precursor (AR) (Colorectum
cell-derived growth factor) (CRDGF).
Length = 252
Score = 32.0 bits (71), Expect = 6.1
Identities = 20/57 (35%), Positives = 26/57 (45%)
Frame = +1
Query: 136 NAQKSKMARERNAEKNKGGKGSQLEANKKAMNIQCKICMQTFICTTSEAKCKEHAEA 306
N +S+ ++ K KGGK + N+K N C Q F C E K EH EA
Sbjct: 113 NKTESENTSDKPKRKKKGGKNGKNRRNRKKKN-PCNAEFQNF-CIHGECKYIEHLEA 167
>sptr|Q9VUU8|Q9VUU8 CG15715 protein.
Length = 77
Score = 31.6 bits (70), Expect = 7.9
Identities = 21/72 (29%), Positives = 30/72 (41%), Gaps = 3/72 (4%)
Frame = +1
Query: 121 AMGGGNAQKSKMARERNAEKNKG---GKGSQLEANKKAMNIQCKICMQTFICTTSEAKCK 291
A G Q A E+ A+ K Q +A +KA+ C +C K
Sbjct: 2 ARGHQKIQSQAKASEKQAKLKKQQGHSANDQKKAAQKALVYVCAVCKSQM---PDPKTYK 58
Query: 292 EHAEARHPKNDL 327
+H E +HPKND+
Sbjct: 59 QHFENKHPKNDI 70
>sptr|Q9VUU7|Q9VUU7 CG18081 protein (RE63284p).
Length = 77
Score = 31.6 bits (70), Expect = 7.9
Identities = 21/72 (29%), Positives = 30/72 (41%), Gaps = 3/72 (4%)
Frame = +1
Query: 121 AMGGGNAQKSKMARERNAEKNKG---GKGSQLEANKKAMNIQCKICMQTFICTTSEAKCK 291
A G Q A E+ A+ K Q +A +KA+ C +C K
Sbjct: 2 ARGHQKIQSQAKASEKQAKLKKQQGHSANDQKKAAQKALVYVCAVCKSQM---PDPKTYK 58
Query: 292 EHAEARHPKNDL 327
+H E +HPKND+
Sbjct: 59 QHFENKHPKNDM 70
>sptr|Q8T032|Q8T032 LD35343p.
Length = 563
Score = 31.6 bits (70), Expect = 7.9
Identities = 14/43 (32%), Positives = 27/43 (62%)
Frame = +1
Query: 211 ANKKAMNIQCKICMQTFICTTSEAKCKEHAEARHPKNDLYQCF 339
+N+K+ QCK+C ++F+ A+ + H +A+H K+ +CF
Sbjct: 399 SNRKS---QCKLCPKSFVTI---AELERHTKAKHSKDKTLRCF 435
>sptr|Q9V5R6|Q9V5R6 CG30020 protein (LD40262p).
Length = 1309
Score = 31.6 bits (70), Expect = 7.9
Identities = 14/43 (32%), Positives = 27/43 (62%)
Frame = +1
Query: 211 ANKKAMNIQCKICMQTFICTTSEAKCKEHAEARHPKNDLYQCF 339
+N+K+ QCK+C ++F+ A+ + H +A+H K+ +CF
Sbjct: 1145 SNRKS---QCKLCPKSFVTI---AELERHTKAKHSKDKTLRCF 1181
Database: /db/trembl-ebi/tmp/swall
Posted date: Oct 24, 2003 8:22 PM
Number of letters in database: 411,996,502
Number of sequences in database: 1,288,562
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 329,424,839
Number of Sequences: 1288562
Number of extensions: 5148309
Number of successful extensions: 10914
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 10457
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 10897
length of database: 411,996,502
effective HSP length: 117
effective length of database: 261,234,748
effective search space used: 22727423076
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

