BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
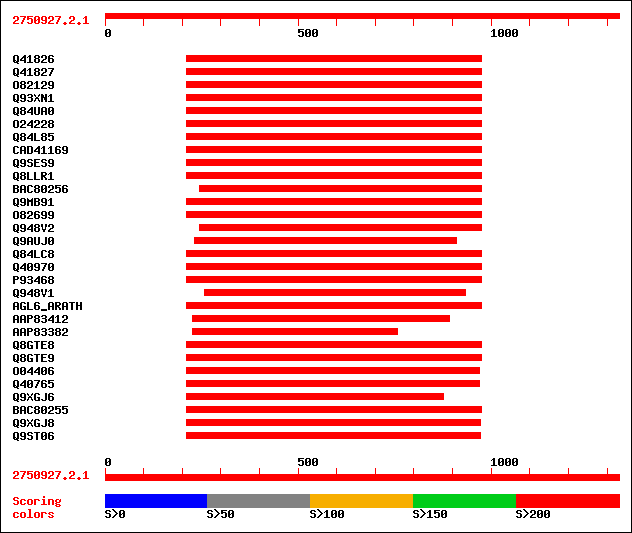
Query= 2750927.2.1
(1331 letters)
Database: /db/trembl-ebi/tmp/swall
1,242,019 sequences; 397,940,306 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q41826|Q41826 MADS box protein. 521 e-147
sptr|Q41827|Q41827 MADS box protein. 459 e-128
sptr|O82129|O82129 MADS box transcription factor. 411 e-114
sptr|Q93XN1|Q93XN1 Mads1. 408 e-113
sptr|Q84UA0|Q84UA0 MADS4. 406 e-112
sptr|O24228|O24228 MADS box protein. 399 e-110
sptr|Q84L85|Q84L85 MADS-box transcription factor SEP1. 335 1e-90
sptrnew|CAD41169|CAD41169 OSJNBa0064M23.14 protein. 327 3e-88
sptr|Q9SES9|Q9SES9 MADS box transcription factor MADS17. 327 3e-88
sptr|Q8LLR1|Q8LLR1 MADS-box protein 3. 318 1e-85
sptrnew|BAC80256|BAC80256 MADS-box transcription factor (Fragment). 308 1e-82
sptr|Q9MB91|Q9MB91 PMADS4 protein. 306 4e-82
sptr|O82699|O82699 MADS-box protein. 306 6e-82
sptr|Q948V2|Q948V2 Putative MADS-domain transcription factor MpM... 299 5e-80
sptr|Q9AUJ0|Q9AUJ0 MADS box protein nmads3 (Fragment). 288 2e-76
sptr|Q84LC8|Q84LC8 MADS-box transcription factor CDM104. 285 8e-76
sptr|Q40970|Q40970 Putative MADS-box family transcription factor. 283 4e-75
sptr|P93468|P93468 MADS-box family transcription factor. 281 1e-74
sptr|Q948V1|Q948V1 Putative MADS-domain transcription factor MpM... 280 4e-74
sw|P29386|AGL6_ARATH Agamous-like MADS box protein AGL6. 275 8e-73
sptrnew|AAP83412|AAP83412 AGL6-like MADS-box (Fragment). 275 1e-72
sptrnew|AAP83382|AAP83382 AGL6-like MADS-box (Fragment). 273 3e-72
sptr|Q8GTE8|Q8GTE8 MADS-box protein AGL6-a. 273 3e-72
sptr|Q8GTE9|Q8GTE9 MADS-box protein AGL6-a. 268 1e-70
sptr|O04406|O04406 MADS-box protein. 265 1e-69
sptr|Q40765|Q40765 Dal1 protein. 265 1e-69
sptr|Q9XGJ6|Q9XGJ6 Putative MADS domain transcription factor GGM11. 265 1e-69
sptrnew|BAC80255|BAC80255 MADS-box transcription factor. 252 1e-65
sptr|Q9XGJ8|Q9XGJ8 Putative MADS domain transcription factor GGM9. 250 3e-65
sptr|Q9ST06|Q9ST06 GpMADS3 protein. 250 4e-65
>sptr|Q41826|Q41826 MADS box protein.
Length = 255
Score = 521 bits (1342), Expect = e-147
Identities = 255/255 (100%), Positives = 255/255 (100%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV
Sbjct: 61 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 120
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYRT 751
KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYRT
Sbjct: 121 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYRT 180
Query: 752 LQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNIPR 931
LQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNIPR
Sbjct: 181 LQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNIPR 240
Query: 932 SPPGGENNFMLGWVL 976
SPPGGENNFMLGWVL
Sbjct: 241 SPPGGENNFMLGWVL 255
>sptr|Q41827|Q41827 MADS box protein.
Length = 255
Score = 459 bits (1182), Expect = e-128
Identities = 234/257 (91%), Positives = 239/257 (92%), Gaps = 2/257 (0%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS RGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSGRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNG-ALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLS 568
G+TKTLERYQHCCYNAQDSN ALSE+QSWYQEMSKLRAKFEALQRTQRHLLGE+LGPLS
Sbjct: 61 GVTKTLERYQHCCYNAQDSNNSALSESQSWYQEMSKLRAKFEALQRTQRHLLGEDLGPLS 120
Query: 569 VKELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYR 748
VKELQQLEKQLECALSQARQRKTQ+MMEQVEELRR ERHLGEMNRQLKHKLEAEGCSNY
Sbjct: 121 VKELQQLEKQLECALSQARQRKTQVMMEQVEELRRTERHLGEMNRQLKHKLEAEGCSNYT 180
Query: 749 TLQHAA-WPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNI 925
TLQHAA WPAPG T+VEHDGATY VHP A SVAMDCEPTLQIGY PHHQF P EA NNI
Sbjct: 181 TLQHAACWPAPGGTIVEHDGATYQVHP-PAHSVAMDCEPTLQIGY-PHHQFPPPEAVNNI 238
Query: 926 PRSPPGGENNFMLGWVL 976
PRS GENNFMLGWVL
Sbjct: 239 PRSAATGENNFMLGWVL 255
>sptr|O82129|O82129 MADS box transcription factor.
Length = 258
Score = 411 bits (1057), Expect = e-114
Identities = 217/264 (82%), Positives = 230/264 (87%), Gaps = 9/264 (3%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
G TKTLERYQHCCYNAQDSNGALSETQSWYQEMSKL+AKFEALQRTQRHLLGE+LGPLSV
Sbjct: 61 GTTKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLKAKFEALQRTQRHLLGEDLGPLSV 120
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEG--CSNY 745
KELQQLEKQLEC+LS ARQRKTQLMMEQVEELRRKER LG++NRQLKHKL+AEG +NY
Sbjct: 121 KELQQLEKQLECSLSLARQRKTQLMMEQVEELRRKERQLGDINRQLKHKLDAEGSNSNNY 180
Query: 746 RTLQHAAWPAPGSTMVEHDGATYHV-----HPTTAQSVAMDCEPTLQIGYPPHHQF-LPS 907
R +Q +W A T+V+ A YH+ HP S AMDCEPTLQIGY HHQF P
Sbjct: 181 RAMQQISWAA--GTVVDEGAAAYHMQQQQQHPN--HSAAMDCEPTLQIGY--HHQFAAPD 234
Query: 908 EAANNIPR-SPPGGENNFMLGWVL 976
+AANNIPR S PGGENNFMLGWVL
Sbjct: 235 QAANNIPRSSAPGGENNFMLGWVL 258
>sptr|Q93XN1|Q93XN1 Mads1.
Length = 259
Score = 408 bits (1049), Expect = e-113
Identities = 216/263 (82%), Positives = 227/263 (86%), Gaps = 8/263 (3%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
G TKTLERYQHCCYNAQDSN ALSETQSWYQEMSKL+AKFEALQRTQRHLLGE+LGPLSV
Sbjct: 61 GTTKTLERYQHCCYNAQDSNSALSETQSWYQEMSKLKAKFEALQRTQRHLLGEDLGPLSV 120
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCS--NY 745
KELQQLEKQLEC+LSQARQRKTQLM+EQVEELRRKER LGE+NRQLKHKL+AEG S NY
Sbjct: 121 KELQQLEKQLECSLSQARQRKTQLMVEQVEELRRKERQLGEINRQLKHKLDAEGSSSNNY 180
Query: 746 RTLQHAAWPAPGSTMVEHDGATYHVHPTTAQ----SVAMDCEPTLQIGYPPHHQF-LPSE 910
R +Q W A T+V+ A YH+ Q S AMDCEPTLQIGYP HQF P +
Sbjct: 181 RAMQQLTWAA--GTVVDEGAAAYHMQHQQQQQPNHSAAMDCEPTLQIGYP--HQFAAPEQ 236
Query: 911 AANNIPRSP-PGGENNFMLGWVL 976
AANNIPRS GGENNFMLGWVL
Sbjct: 237 AANNIPRSSGQGGENNFMLGWVL 259
>sptr|Q84UA0|Q84UA0 MADS4.
Length = 260
Score = 406 bits (1044), Expect = e-112
Identities = 214/261 (81%), Positives = 227/261 (86%), Gaps = 6/261 (2%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
G TKTLERYQHCCYNAQDS+ ALSETQSWYQEMSKL+AKFEALQRTQRHLLGE+LGPLSV
Sbjct: 61 GTTKTLERYQHCCYNAQDSSSALSETQSWYQEMSKLKAKFEALQRTQRHLLGEDLGPLSV 120
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCS--NY 745
KELQQLEKQLEC+LSQARQRKTQLMMEQVEELRRKER LGE+NRQLKHKL+AEG S ++
Sbjct: 121 KELQQLEKQLECSLSQARQRKTQLMMEQVEELRRKERQLGEINRQLKHKLDAEGSSSNSF 180
Query: 746 RTLQHAAWPAPGSTMVEHDGATYHVH--PTTAQSVAMDCEPTLQIGYPPHHQFLPSE-AA 916
R +Q W A T+V+ YH+H S AMDCEPTLQIGYP HQF +E AA
Sbjct: 181 RAMQQITWAA--GTVVDEGAGAYHMHHQQQPNHSAAMDCEPTLQIGYP--HQFAAAEQAA 236
Query: 917 NNIPR-SPPGGENNFMLGWVL 976
NNIPR S PGGENNFMLGWVL
Sbjct: 237 NNIPRSSAPGGENNFMLGWVL 257
>sptr|O24228|O24228 MADS box protein.
Length = 250
Score = 399 bits (1026), Expect = e-110
Identities = 216/258 (83%), Positives = 225/258 (87%), Gaps = 3/258 (1%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
GITKTLERYQHCCYNAQDSN ALSETQSWY EMSKL+AKFEALQRTQRHLLGE+LGPLSV
Sbjct: 61 GITKTLERYQHCCYNAQDSNNALSETQSWYHEMSKLKAKFEALQRTQRHLLGEDLGPLSV 120
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEG-CSNYR 748
KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKER LGE+NRQLKHKLE EG SNYR
Sbjct: 121 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERQLGEINRQLKHKLEVEGSTSNYR 180
Query: 749 TLQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNIP 928
+Q A+W A G+ V +GA Y P S AMD EPTLQIGYP HQF+P+E AN I
Sbjct: 181 AMQQASW-AQGA--VVENGAAYVQPP--PHSAAMDSEPTLQIGYP--HQFVPAE-ANTIQ 232
Query: 929 RS--PPGGENNFMLGWVL 976
RS P G ENNFMLGWVL
Sbjct: 233 RSTAPAGAENNFMLGWVL 250
>sptr|Q84L85|Q84L85 MADS-box transcription factor SEP1.
Length = 243
Score = 335 bits (858), Expect = 1e-90
Identities = 177/255 (69%), Positives = 209/255 (81%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALI+FSSRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIVFSSRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
G +KTLERYQ CCY +QD+ A E Q+WYQE+++L+AKFE+LQ QRHLLGE+LGPLSV
Sbjct: 61 GTSKTLERYQRCCYTSQDATIADREKQNWYQEVARLKAKFESLQSAQRHLLGEDLGPLSV 120
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYRT 751
KELQQLE+QLE +LSQARQRKTQ+M +Q+EELR+KE HLGE+N+QLK KLEAEG N R
Sbjct: 121 KELQQLERQLEASLSQARQRKTQIMFDQMEELRKKEHHLGEINKQLKTKLEAEG-ENLRA 179
Query: 752 LQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNIPR 931
+Q +W + + + G + +HP + S AM+CEPTLQIGY HQ + E ++PR
Sbjct: 180 IQ-GSWESDATNV--GGGNVFSMHP--SHSSAMECEPTLQIGY---HQLVQPE--GSLPR 229
Query: 932 SPPGGENNFMLGWVL 976
+ GGENNFMLGWVL
Sbjct: 230 N-SGGENNFMLGWVL 243
>sptrnew|CAD41169|CAD41169 OSJNBa0064M23.14 protein.
Length = 249
Score = 327 bits (837), Expect = 3e-88
Identities = 181/261 (69%), Positives = 204/261 (78%), Gaps = 6/261 (2%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALS--ETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPL 565
GI KTLE+Y CCYNAQ SN AL+ E QSWYQEMS+L+ K E LQR+QRH+LGE+LGPL
Sbjct: 61 GINKTLEKYNSCCYNAQGSNSALAGGEHQSWYQEMSRLKTKLECLQRSQRHMLGEDLGPL 120
Query: 566 SVKELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEG-CSN 742
S+KELQQLEKQLE +LSQARQRKTQ+MMEQV++LRRKER LGE+N+QLK+KLEAE SN
Sbjct: 121 SIKELQQLEKQLEYSLSQARQRKTQIMMEQVDDLRRKERQLGELNKQLKNKLEAEADSSN 180
Query: 743 YRTLQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANN 922
R+ +W V G + P +DCEPTLQIGY +QF+ EAAN
Sbjct: 181 CRSAIQDSWV---HGTVVSGGRVLNAQPPP----DIDCEPTLQIGY---YQFVRPEAAN- 229
Query: 923 IPRSPPGG---ENNFMLGWVL 976
PRS GG NNF++GW L
Sbjct: 230 -PRSNGGGGDQNNNFVMGWPL 249
>sptr|Q9SES9|Q9SES9 MADS box transcription factor MADS17.
Length = 249
Score = 327 bits (837), Expect = 3e-88
Identities = 181/261 (69%), Positives = 204/261 (78%), Gaps = 6/261 (2%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALS--ETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPL 565
GI KTLE+Y CCYNAQ SN AL+ E QSWYQEMS+L+ K E LQR+QRH+LGE+LGPL
Sbjct: 61 GINKTLEKYNSCCYNAQGSNSALAGGEHQSWYQEMSRLKTKLECLQRSQRHMLGEDLGPL 120
Query: 566 SVKELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEG-CSN 742
S+KELQQLEKQLE +LSQARQRKTQ+MMEQV++LRRKER LGE+N+QLK+KLEAE SN
Sbjct: 121 SIKELQQLEKQLEYSLSQARQRKTQIMMEQVDDLRRKERQLGELNKQLKNKLEAEADSSN 180
Query: 743 YRTLQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANN 922
R+ +W V G + P +DCEPTLQIGY +QF+ EAAN
Sbjct: 181 CRSAIQDSWV---HGTVVSGGRVLNAQPPP----DIDCEPTLQIGY---YQFVRPEAAN- 229
Query: 923 IPRSPPGG---ENNFMLGWVL 976
PRS GG NNF++GW L
Sbjct: 230 -PRSNGGGGDQNNNFVMGWPL 249
>sptr|Q8LLR1|Q8LLR1 MADS-box protein 3.
Length = 244
Score = 318 bits (815), Expect = 1e-85
Identities = 176/257 (68%), Positives = 208/257 (80%), Gaps = 2/257 (0%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
G TKTLERYQ CY QD+N ETQSWYQE+SKL+AK+E+LQRTQRHLLGE+LGPLSV
Sbjct: 61 GTTKTLERYQRVCYTPQDNNMEC-ETQSWYQEVSKLKAKYESLQRTQRHLLGEDLGPLSV 119
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYRT 751
KELQ LEKQLE AL+QARQRKTQ+M+EQ+E+LRRKER LG++N+QLK KLEAEG + +
Sbjct: 120 KELQNLEKQLEGALAQARQRKTQMMIEQMEDLRRKERQLGDLNKQLKLKLEAEG-QSLKA 178
Query: 752 LQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDC--EPTLQIGYPPHHQFLPSEAANNI 925
+Q + P+ + +++ VHP +QS MDC EP LQIGY H ++P+E ++
Sbjct: 179 IQGSWNPSTATA----GNSSFPVHP--SQSNPMDCEPEPILQIGY---HHYVPAEGP-SV 228
Query: 926 PRSPPGGENNFMLGWVL 976
+S GE+NF+ GWVL
Sbjct: 229 SKS-MAGESNFIQGWVL 244
>sptrnew|BAC80256|BAC80256 MADS-box transcription factor (Fragment).
Length = 227
Score = 308 bits (788), Expect = 1e-82
Identities = 164/244 (67%), Positives = 194/244 (79%)
Frame = +2
Query: 245 ENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSAGITKTLERYQH 424
ENKINRQVTFSKRRNGLLKKAYELS+LCDAE+ALIIFSSRGKLYEFGS+G+TKTLERYQ
Sbjct: 1 ENKINRQVTFSKRRNGLLKKAYELSILCDAEIALIIFSSRGKLYEFGSSGLTKTLERYQR 60
Query: 425 CCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSVKELQQLEKQLE 604
C Y Q++N A E Q W+QE+SKL+AK+E L R+QRHLLGE+LGPLSVKELQQLE+QLE
Sbjct: 61 CSYVPQENNPADREAQVWHQEISKLKAKYELLLRSQRHLLGEDLGPLSVKELQQLERQLE 120
Query: 605 CALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYRTLQHAAWPAPGS 784
ALSQARQRKTQ+MMEQ+EELR+KER LG++N+QLK KLEAEG + ++Q AW
Sbjct: 121 VALSQARQRKTQIMMEQMEELRKKERCLGDINKQLKGKLEAEGIGAFSSIQ-GAW----- 174
Query: 785 TMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNIPRSPPGGENNFML 964
A VHP +QS +DCEP+LQIGY HQF+P EAA +P GGE+NF+
Sbjct: 175 ----ESAAPVVVHP--SQSADVDCEPSLQIGY---HQFVPQEAA--MPCRSAGGESNFIQ 223
Query: 965 GWVL 976
GW+L
Sbjct: 224 GWML 227
>sptr|Q9MB91|Q9MB91 PMADS4 protein.
Length = 253
Score = 306 bits (784), Expect = 4e-82
Identities = 172/262 (65%), Positives = 192/262 (73%), Gaps = 7/262 (2%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFG+A
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGNA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
GITKTLERYQ CC N QD+ G ETQSWYQE+SKL+ KFEALQRTQRHLLGE+LGPLSV
Sbjct: 61 GITKTLERYQRCCLNPQDNCGE-RETQSWYQEVSKLKGKFEALQRTQRHLLGEDLGPLSV 119
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHK-------LEAE 730
KELQ LEKQLE AL+QARQRKTQ+MMEQ+EELRRKERHLG++N+QLK K LEAE
Sbjct: 120 KELQNLEKQLEGALAQARQRKTQIMMEQMEELRRKERHLGDVNKQLKVKVSLELSSLEAE 179
Query: 731 GCSNYRTLQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSE 910
G + W S +E + + VH +QS MDC P + Y HH
Sbjct: 180 GQAGLNRALPFLWT---SNALEAGNSNFPVH--HSQSNPMDCGPDPVLQYRDHHYMAADG 234
Query: 911 AANNIPRSPPGGENNFMLGWVL 976
+ + + E N M GW L
Sbjct: 235 PSGSRSMAV---ETNIMQGWGL 253
>sptr|O82699|O82699 MADS-box protein.
Length = 243
Score = 306 bits (783), Expect = 6e-82
Identities = 168/257 (65%), Positives = 204/257 (79%), Gaps = 2/257 (0%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEV LIIFSSRGKLYEF SA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVGLIIFSSRGKLYEFASA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
G++KTLERYQ C + + N ETQSWYQE++KL+AK+E+LQRTQRHLLGE+LGPLSV
Sbjct: 61 GMSKTLERYQRCSFTPPE-NSIERETQSWYQEVTKLKAKYESLQRTQRHLLGEDLGPLSV 119
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYRT 751
KELQ LEKQLE AL+Q RQRKTQLM+EQ+E+LR+KERHLG++N+QL+ KLEAEG N
Sbjct: 120 KELQNLEKQLEGALAQTRQRKTQLMIEQMEDLRKKERHLGDLNKQLRVKLEAEG-QNLNV 178
Query: 752 LQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDC--EPTLQIGYPPHHQFLPSEAANNI 925
+Q+ W + + + + +H ++Q+ MDC EP +Q+GY HQ+ P+E ++I
Sbjct: 179 IQN-MWSSDAAA----GSSNFSLH--SSQTNPMDCTPEPVIQMGY---HQYHPAE-GSSI 227
Query: 926 PRSPPGGENNFMLGWVL 976
PRS GE NF+ GWVL
Sbjct: 228 PRSLT-GETNFIQGWVL 243
>sptr|Q948V2|Q948V2 Putative MADS-domain transcription factor
MpMADS3 (Fragment).
Length = 231
Score = 299 bits (766), Expect = 5e-80
Identities = 160/244 (65%), Positives = 195/244 (79%)
Frame = +2
Query: 245 ENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSAGITKTLERYQH 424
ENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALI+FSSRGKLYEFGSAGI KTLERYQ
Sbjct: 1 ENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIVFSSRGKLYEFGSAGINKTLERYQR 60
Query: 425 CCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSVKELQQLEKQLE 604
CCY D+N +TQ WYQE+SKL AK ++LQR+QRHLLGE+LGPLSVKELQ+LE+QLE
Sbjct: 61 CCYTFHDANITDRDTQGWYQEVSKLNAKCDSLQRSQRHLLGEDLGPLSVKELQKLERQLE 120
Query: 605 CALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYRTLQHAAWPAPGS 784
AL+Q RQRKTQ+M+E +E LR KER LG++N++LK+KLEA+G +R +Q A+W +
Sbjct: 121 SALTQTRQRKTQIMLEHMEALREKERQLGDINKELKNKLEAKGQGAFRAMQ-ASWES--G 177
Query: 785 TMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNIPRSPPGGENNFML 964
+V ++G + +HP +QS A++CEPTLQIGY H F EA NIPR+ E+NFM
Sbjct: 178 PLVGNNG--FPMHP--SQSAAIECEPTLQIGY---HSFAAPEA--NIPRTVV-AESNFMH 227
Query: 965 GWVL 976
GW+L
Sbjct: 228 GWIL 231
>sptr|Q9AUJ0|Q9AUJ0 MADS box protein nmads3 (Fragment).
Length = 236
Score = 288 bits (736), Expect = 2e-76
Identities = 157/229 (68%), Positives = 179/229 (78%), Gaps = 3/229 (1%)
Frame = +2
Query: 233 LKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSAGITKTLE 412
+K IENKI RQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSAGI KTLE
Sbjct: 1 IKSIENKITRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSAGINKTLE 60
Query: 413 RYQHCCYNAQDSNGALS--ETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSVKELQQ 586
+Y CCYNAQ SN AL+ E QSWYQEMS+L+ K E LQR+QRH+LGE+LGPLS+KELQQ
Sbjct: 61 KYNSCCYNAQGSNSALAGGEHQSWYQEMSRLKTKLECLQRSQRHMLGEDLGPLSIKELQQ 120
Query: 587 LEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEG-CSNYRTLQHA 763
LEKQLE +LSQARQRKTQ+MMEQV++LRRKER LGE+N+QLK+KLEAE SN R+
Sbjct: 121 LEKQLEYSLSQARQRKTQIMMEQVDDLRRKERQLGELNKQLKNKLEAEADSSNCRSAIQD 180
Query: 764 AWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSE 910
+W V G + P +DCEPTLQIGY +QF+ E
Sbjct: 181 SWV---HGTVVSGGRVLNAQPPP----DIDCEPTLQIGY---YQFVRPE 219
>sptr|Q84LC8|Q84LC8 MADS-box transcription factor CDM104.
Length = 250
Score = 285 bits (730), Expect = 8e-76
Identities = 166/265 (62%), Positives = 194/265 (73%), Gaps = 10/265 (3%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEV LIIFSSR KLYEFGS
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVGLIIFSSRDKLYEFGSV 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
G+ KTLERYQ CC+N QD+N ETQSWYQE+SKL+AKFE+LQRTQRHLLGE+LGPLSV
Sbjct: 61 GVMKTLERYQRCCFNPQDNNNE-RETQSWYQEVSKLKAKFESLQRTQRHLLGEDLGPLSV 119
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKL-------EAE 730
KEL LEKQLE AL+QARQRKTQ+++EQ+EELR KER LG+MN+ LK K+ E E
Sbjct: 120 KELHNLEKQLEGALTQARQRKTQILVEQMEELRCKERELGDMNKHLKIKVSHELSTFETE 179
Query: 731 GCSNYRTLQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDC---EPTLQIGYPPHHQFL 901
G YRT Q P P ++ + T+ +HP +QS M C +P LQIG +QF+
Sbjct: 180 G-QGYRTHQ---LPCPWNS---SNNNTFLMHP--SQSNPMGCQQEQPILQIG---DNQFM 227
Query: 902 PSEAANNIPRSPPGGENNFMLGWVL 976
E ++ + E N WV+
Sbjct: 228 QGEGSSG--QRNMVDETNIHGNWVI 250
>sptr|Q40970|Q40970 Putative MADS-box family transcription factor.
Length = 242
Score = 283 bits (724), Expect = 4e-75
Identities = 159/256 (62%), Positives = 192/256 (75%), Gaps = 1/256 (0%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
G+ KTLERYQ C Y QD+ + E Q+W+QE+ KL+A+ E LQR+QRHLLGE+LGPLS+
Sbjct: 61 GMLKTLERYQKCSYVLQDATVSDREAQNWHQEVGKLKARVELLQRSQRHLLGEDLGPLSI 120
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKL-EAEGCSNYR 748
KELQQLE+QLE AL+ R RKTQ+M+E ++ELRRKER L E+N+ L+ KL EAEG +
Sbjct: 121 KELQQLERQLEVALTHVRSRKTQVMLEMMDELRRKERILQEVNKSLRKKLQEAEG-QAFN 179
Query: 749 TLQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNIP 928
+Q P + + A HP S A+DCEPTLQIGY Q+ P E +++P
Sbjct: 180 AMQPP--PHAWDSHAVANNAYAMQHP----SNAVDCEPTLQIGY----QYAPPE--SSMP 227
Query: 929 RSPPGGENNFMLGWVL 976
R +NN+M GW++
Sbjct: 228 RHEQ-AQNNYMQGWMV 242
>sptr|P93468|P93468 MADS-box family transcription factor.
Length = 242
Score = 281 bits (719), Expect = 1e-74
Identities = 158/256 (61%), Positives = 191/256 (74%), Gaps = 1/256 (0%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
G+ KTLERYQ C Y QD+ + E Q+W+QE+ KL+A+ E LQR+QRHLLGE+LGPLS+
Sbjct: 61 GMLKTLERYQKCSYVLQDATVSDREAQNWHQEVGKLKARVELLQRSQRHLLGEDLGPLSI 120
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKL-EAEGCSNYR 748
KELQQLE+QLE AL+ R RKTQ+M+E ++ELRRKER L E+N+ L+ KL EAEG +
Sbjct: 121 KELQQLERQLEVALTHVRSRKTQVMLEMMDELRRKERILQEVNKSLRKKLQEAEG-QAFN 179
Query: 749 TLQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNIP 928
+Q P + + A HP S A+DCEPTLQ GY Q+ P E +++P
Sbjct: 180 AMQPP--PHAWDSHAVANNAYAMQHP----SNAVDCEPTLQTGY----QYAPPE--SSMP 227
Query: 929 RSPPGGENNFMLGWVL 976
R +NN+M GW++
Sbjct: 228 RHEQ-AQNNYMQGWMV 242
>sptr|Q948V1|Q948V1 Putative MADS-domain transcription factor
MpMADS4 (Fragment).
Length = 248
Score = 280 bits (715), Expect = 4e-74
Identities = 151/226 (66%), Positives = 178/226 (78%)
Frame = +2
Query: 257 NRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSAGITKTLERYQHCCYN 436
NRQVTF KRRNGLLKKAYELSVLCDAEVALIIFSSRGKLY FGSAG KT ERYQ CCY
Sbjct: 1 NRQVTFCKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYGFGSAGTNKTPERYQRCCYT 60
Query: 437 AQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSVKELQQLEKQLECALS 616
QD + ETQ WYQE+S+L+AK+E+LQR+QRHLLGE+LGPLSVKELQ LE+QLE ALS
Sbjct: 61 PQDVVVSDRETQGWYQEVSRLKAKYESLQRSQRHLLGEDLGPLSVKELQHLERQLEVALS 120
Query: 617 QARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYRTLQHAAWPAPGSTMVE 796
QARQRKTQ+M+EQ+EELR+KER LG++N+QLK KLEAEG +R +Q +W + MV
Sbjct: 121 QARQRKTQIMIEQMEELRKKERQLGDINKQLKIKLEAEGQGPFRCIQ-GSWES--GAMVG 177
Query: 797 HDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNIPRS 934
++ + + Q+ M+CEPTLQIGY H F+P EA NIPRS
Sbjct: 178 NNNFSMN----APQAAPMECEPTLQIGY---HHFVPPEA--NIPRS 214
>sw|P29386|AGL6_ARATH Agamous-like MADS box protein AGL6.
Length = 252
Score = 275 bits (704), Expect = 8e-73
Identities = 149/256 (58%), Positives = 190/256 (74%), Gaps = 1/256 (0%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVE+KRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGS
Sbjct: 1 MGRGRVEMKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSV 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSE-TQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLS 568
GI T+ERY CYN SN E TQSW QE++KL++K+E+L RT R+LLGE+LG +
Sbjct: 61 GIESTIERYNR-CYNCSLSNNKPEETTQSWCQEVTKLKSKYESLVRTNRNLLGEDLGEMG 119
Query: 569 VKELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYR 748
VKELQ LE+QLE AL+ RQRKTQ+MME++E+LR+KER LG++N+QLK K E EG + ++
Sbjct: 120 VKELQALERQLEAALTATRQRKTQVMMEEMEDLRKKERQLGDINKQLKIKFETEGHA-FK 178
Query: 749 TLQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNIP 928
T Q + S + + + + V P+ + + EP LQIG+ H+ ++ E +++
Sbjct: 179 TFQDLWANSAASVAGDPNNSEFPVEPSHPNVLDCNTEPFLQIGFQQHY-YVQGE-GSSVS 236
Query: 929 RSPPGGENNFMLGWVL 976
+S GE NF+ GWVL
Sbjct: 237 KSNVAGETNFVQGWVL 252
>sptrnew|AAP83412|AAP83412 AGL6-like MADS-box (Fragment).
Length = 242
Score = 275 bits (702), Expect = 1e-72
Identities = 148/232 (63%), Positives = 176/232 (75%), Gaps = 10/232 (4%)
Frame = +2
Query: 227 VELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSAGITKT 406
V+L+R+ENKINRQVTFSKRRNGLLKKAYELSVLCDAEV LIIFSSRGKLYEFGSAG+T T
Sbjct: 2 VQLRRMENKINRQVTFSKRRNGLLKKAYELSVLCDAEVGLIIFSSRGKLYEFGSAGMTAT 61
Query: 407 LERYQHCCYNAQDSN-GALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSVKELQ 583
LERYQ CC+N Q++ GA ETQSWYQE+SKL+AK+E+LQRTQRHLLGE+LGPL+VKEL+
Sbjct: 62 LERYQRCCFNPQNAGAGAERETQSWYQEVSKLKAKYESLQRTQRHLLGEDLGPLNVKELE 121
Query: 584 QLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLK---------HKLEAEGC 736
LEKQLE +LSQARQRKT++MMEQ+E+LRRKER LGEMN+QLK H+ E +G
Sbjct: 122 NLEKQLEGSLSQARQRKTKIMMEQMEDLRRKERQLGEMNKQLKIRVSLELSSHETEGQGL 181
Query: 737 SNYRTLQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHH 892
+ +AA A S + G + + EP LQ+GY HH
Sbjct: 182 RGFPCQXNAAASAGTSNFMAQPG---------TNPMEFEPEPVLQMGY--HH 222
>sptrnew|AAP83382|AAP83382 AGL6-like MADS-box (Fragment).
Length = 206
Score = 273 bits (699), Expect = 3e-72
Identities = 138/177 (77%), Positives = 157/177 (88%)
Frame = +2
Query: 227 VELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSAGITKT 406
V+LKR+ENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSAG KT
Sbjct: 1 VQLKRMENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSAGTNKT 60
Query: 407 LERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSVKELQQ 586
LERYQ CCY QD + ETQ WYQE+SKL+AK+E+LQR+QRHLL E+LGPLSVKELQ
Sbjct: 61 LERYQRCCYTPQDVVVSDRETQGWYQEVSKLKAKYESLQRSQRHLLXEDLGPLSVKELQH 120
Query: 587 LEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYRTLQ 757
LE+QLE ALSQARQRKTQ+M+EQ+EELR+KER LG++N+QLK KLEAEG +R +Q
Sbjct: 121 LERQLEVALSQARQRKTQIMIEQMEELRKKERQLGDINKQLKIKLEAEGQGPFRCIQ 177
>sptr|Q8GTE8|Q8GTE8 MADS-box protein AGL6-a.
Length = 252
Score = 273 bits (699), Expect = 3e-72
Identities = 146/257 (56%), Positives = 194/257 (75%), Gaps = 2/257 (0%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVE+KRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALI+FSSRGKLYEFGS
Sbjct: 1 MGRGRVEMKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIVFSSRGKLYEFGSV 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSET-QSWYQEMSKLRAKFEALQRTQRHLLGEELGPLS 568
G+ +T+ERY H CYN +N E+ Q+W QE++KL+AK+E+L RT RHLLGE++G +
Sbjct: 61 GVERTIERY-HRCYNCSVTNNRPEESKQNWCQEVAKLKAKYESLVRTNRHLLGEDIGEMG 119
Query: 569 VKELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYR 748
VK+LQ LE+QLE AL+ RQRKTQ+MME++E+LR+KER LG++N+QLK K EA G + ++
Sbjct: 120 VKQLQALERQLEAALTATRQRKTQVMMEEMEDLRKKERQLGDINKQLKIKFEAGGHA-FK 178
Query: 749 TLQHAAWPAPGSTMV-EHDGATYHVHPTTAQSVAMDCEPTLQIGYPPHHQFLPSEAANNI 925
+ Q WP ++M + + + + V P+ SV + EP LQIG+ H+ ++ E +++
Sbjct: 179 SFQD-FWPNSAASMAGDPNNSKFPVQPSHPDSVDCNTEPFLQIGFQQHY-YVQGE-GSSV 235
Query: 926 PRSPPGGENNFMLGWVL 976
P+S E NF+ W L
Sbjct: 236 PKSNVACETNFVQDWFL 252
>sptr|Q8GTE9|Q8GTE9 MADS-box protein AGL6-a.
Length = 259
Score = 268 bits (686), Expect = 1e-70
Identities = 146/263 (55%), Positives = 192/263 (73%), Gaps = 8/263 (3%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVE+KRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALI+FSSRGKLYEFGS
Sbjct: 1 MGRGRVEMKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIVFSSRGKLYEFGSV 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSET-QSWYQEMSKLRAKFEALQRTQRHLLGEELGPLS 568
G+ +T+ERY H CYN +N E+ Q+W QE++KL+AK+E+L RT RHLLGE++G +
Sbjct: 61 GVERTIERY-HRCYNCSVTNNRPEESKQNWCQEVAKLKAKYESLVRTNRHLLGEDIGEMG 119
Query: 569 VKELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYR 748
VK+LQ LE+QLE AL+ RQRKTQ+MME++E+LR+KER LG++N+QLK K EA G + ++
Sbjct: 120 VKQLQALERQLEAALTATRQRKTQVMMEEMEDLRKKERQLGDINKQLKIKFEAGGHA-FK 178
Query: 749 TLQHAAWPAPGSTMV-EHDGATYHVHPTTAQSVAMDCEPTLQIGYP------PHHQFLPS 907
+ Q WP ++M + + + + V P+ SV + EP LQIG H ++
Sbjct: 179 SFQD-FWPNSAASMAGDPNNSKFPVQPSHPDSVDCNTEPFLQIGLVLVYIRFQQHYYVQG 237
Query: 908 EAANNIPRSPPGGENNFMLGWVL 976
E +++P+S E NF+ W L
Sbjct: 238 E-GSSVPKSNVACETNFVQDWFL 259
>sptr|O04406|O04406 MADS-box protein.
Length = 261
Score = 265 bits (677), Expect = 1e-69
Identities = 151/269 (56%), Positives = 186/269 (69%), Gaps = 16/269 (5%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRV+L+RIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS+RGKLYEF S+
Sbjct: 1 MGRGRVQLRRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSTRGKLYEFASS 60
Query: 392 GITKTLERYQHCCYNAQDSNGALS-ETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLS 568
+ KTLERY+ C Y QD+ G E Q+W+QE++KL+ K E LQR+QRHLLGE+LGPL+
Sbjct: 61 SMNKTLERYEKCSYAMQDTTGVSDREAQNWHQEVTKLKGKVELLQRSQRHLLGEDLGPLN 120
Query: 569 VKELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKL-EAEG---- 733
VKELQQLE+QLE AL+ R RKTQ+M++Q+EELR++ER L E+N+ L+ KL E EG
Sbjct: 121 VKELQQLERQLEVALTHLRSRKTQVMLDQIEELRQRERLLHEVNKSLQKKLSETEGRDVI 180
Query: 734 -----CSNYRTLQHAAWPAPGSTMVEHDGATYHVHPTTAQSV-AMDCEPTLQIGYPP--H 889
SN T + W S++ A H + S+ +DCEPTLQIGY P
Sbjct: 181 TGIEQTSNTNTGTNGPW---DSSITNTAYALSHPQQDSNSSLHHVDCEPTLQIGYQPVAP 237
Query: 890 HQFLPSEAANNIPRSPPGGE--NNFMLGW 970
+P P PP + N +M GW
Sbjct: 238 ESIVP-------PHQPPHNQTPNQYMQGW 259
>sptr|Q40765|Q40765 Dal1 protein.
Length = 261
Score = 265 bits (677), Expect = 1e-69
Identities = 151/267 (56%), Positives = 187/267 (70%), Gaps = 14/267 (5%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRV+L+RIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS+RGKLYEF S+
Sbjct: 1 MGRGRVQLRRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSTRGKLYEFASS 60
Query: 392 GITKTLERYQHCCYNAQDSNGALS-ETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLS 568
+ KTLERY+ C Y QD+ G E Q+W+QE++KL+ K E LQR+QRHLLGE+LGPL+
Sbjct: 61 SMNKTLERYEKCSYAMQDTTGVSDREAQNWHQEVTKLKGKVELLQRSQRHLLGEDLGPLN 120
Query: 569 VKELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKL-EAEG---- 733
VKELQQLE+QLE AL+ R RKTQ+M++Q+EELR++ER L E+N+ L+ KL E EG
Sbjct: 121 VKELQQLERQLEVALAHLRSRKTQVMLDQIEELRQRERLLHEVNKSLQKKLSETEGRDVI 180
Query: 734 -----CSNYRTLQHAAWPAPGSTMVEHDGATYHVHPTTAQSV-AMDCEPTLQIGYPPHHQ 895
SN T + W S++ A H + S+ +DCEPTLQIGY P
Sbjct: 181 TGIEQTSNTNTGTNGPW---DSSITNTAYALSHPQQNSNASLHHVDCEPTLQIGYQP--- 234
Query: 896 FLPSEAANNIPRSPPGG--ENNFMLGW 970
+P E+ P P +N +M GW
Sbjct: 235 -VPPESIGP-PHQPQHNQTQNQYMQGW 259
>sptr|Q9XGJ6|Q9XGJ6 Putative MADS domain transcription factor GGM11.
Length = 254
Score = 265 bits (676), Expect = 1e-69
Identities = 138/222 (62%), Positives = 167/222 (75%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
G KTLERYQ C Y Q+SN + + Q+W+ E+SKL+ K E LQR+QRHLLGE+LGPLS+
Sbjct: 61 GTLKTLERYQKCSYALQESNNSDRDAQTWHHEVSKLKTKVEILQRSQRHLLGEDLGPLSI 120
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYRT 751
+ELQ LE+Q+E AL+Q R RKTQ+MM+ +++L++KER L E+N+ L+ KL+ Y
Sbjct: 121 RELQTLERQIEVALTQVRARKTQVMMDMMDDLKKKERLLQEVNKSLRKKLDETEGQVYSN 180
Query: 752 LQHAAWPAPGSTMVEHDGATYHVHPTTAQSVAMDCEPTLQIG 877
Q A P P Y + PT A+DCEPT ++G
Sbjct: 181 AQLQAAPPPEWDSNAIANPVYALPPTPQN--AVDCEPTCKLG 220
>sptrnew|BAC80255|BAC80255 MADS-box transcription factor.
Length = 247
Score = 252 bits (643), Expect = 1e-65
Identities = 145/258 (56%), Positives = 176/258 (68%), Gaps = 3/258 (1%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEF-GS 388
MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS+RGKLYEF S
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSNRGKLYEFCSS 60
Query: 389 AGITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLS 568
+ + TLERYQ C Y+ ++ G ET+ YQE KL+ K E LQRTQR+LLGE+LGPLS
Sbjct: 61 SSMMTTLERYQECSYSMPEATGPTRETEKSYQEYLKLKGKVELLQRTQRNLLGEDLGPLS 120
Query: 569 VKELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKLEAEGCSNYR 748
KEL+QLE QLE +L Q R KTQ +++Q+ +LRRKE+ + E N+ LK KL G N
Sbjct: 121 SKELEQLENQLEHSLRQIRSTKTQALLDQLSDLRRKEQQMLESNKILKKKLAEHGPEN-- 178
Query: 749 TLQHAAWPAPGSTMVEHDGATYHVHPTTAQSV--AMDCEPTLQIGYPPHHQFLPSEAANN 922
L AW + G + Y P +++ +DC PTLQIGY P Q + AA
Sbjct: 179 -LLQLAWQSCGQS------NPYSRQPAHSEAFFQPLDCNPTLQIGYHPVGQEEITMAAPA 231
Query: 923 IPRSPPGGENNFMLGWVL 976
I +PP N F+ GW++
Sbjct: 232 I--APPQNVNGFIPGWMV 247
>sptr|Q9XGJ8|Q9XGJ8 Putative MADS domain transcription factor GGM9.
Length = 253
Score = 250 bits (639), Expect = 3e-65
Identities = 139/259 (53%), Positives = 179/259 (69%), Gaps = 5/259 (1%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRV+L+RIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS+RGKLYEF S+
Sbjct: 1 MGRGRVQLRRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSTRGKLYEFASS 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
++KTLERY+ C Y+ Q++ + Q+W+ E++KL+AK E+L + QR L+GE+LGPL++
Sbjct: 61 SMSKTLERYEKCSYSMQENASTDRDAQNWHHEVTKLKAKLESLHKAQRSLMGEDLGPLNI 120
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKL-EAEGCSNYR 748
KELQ LE+QLE AL R RKTQL+++ ++ELR KER L E+N+ L+ KL E EG +
Sbjct: 121 KELQSLEQQLEVALGHVRNRKTQLLIQTIDELRDKERTLQEVNKSLQKKLSETEGRNVIP 180
Query: 749 TLQHAAWPAPGSTMVEHDGATYHVHPTTAQSVA----MDCEPTLQIGYPPHHQFLPSEAA 916
L HA GS E T + Q + +DCEPTL IG P+HQ + EA
Sbjct: 181 ALPHA---PNGSGEWESSTLTTTTYALQTQQTSNIHHVDCEPTLHIG--PYHQAVHHEAI 235
Query: 917 NNIPRSPPGGENNFMLGWV 973
P + N + + WV
Sbjct: 236 TAPPATHSEPHNQY-IWWV 253
>sptr|Q9ST06|Q9ST06 GpMADS3 protein.
Length = 252
Score = 250 bits (638), Expect = 4e-65
Identities = 140/258 (54%), Positives = 179/258 (69%), Gaps = 4/258 (1%)
Frame = +2
Query: 212 MGRGRVELKRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSSRGKLYEFGSA 391
MGRGRV+L+RIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFS+RGKLYEF S+
Sbjct: 1 MGRGRVQLRRIENKINRQVTFSKRRNGLLKKAYELSVLCDAEVALIIFSTRGKLYEFASS 60
Query: 392 GITKTLERYQHCCYNAQDSNGALSETQSWYQEMSKLRAKFEALQRTQRHLLGEELGPLSV 571
++KTLERY+ C Y+ Q++ + Q+W+ E++KL+AK E+L + QR+L+GE+LGPL++
Sbjct: 61 SMSKTLERYEKCSYSMQENASTDRDAQNWHHEVTKLKAKLESLHKAQRNLMGEDLGPLNI 120
Query: 572 KELQQLEKQLECALSQARQRKTQLMMEQVEELRRKERHLGEMNRQLKHKL-EAEGCSNYR 748
KELQ LE+QLE AL R RKTQL+++ ++ELR KER L E+N+ L+ KL E EG +
Sbjct: 121 KELQSLEQQLEVALGHVRNRKTQLLIQTIDELRDKERTLQEVNKSLQKKLSETEGRNVIP 180
Query: 749 TLQHAAWPAPGSTMVEHDGATYHVHPTTAQSVA---MDCEPTLQIGYPPHHQFLPSEAAN 919
L HA GS E T T Q +DCEPTL IG P+HQ + EA
Sbjct: 181 ALPHA---TTGSGEWESSTLTTTYALQTQQPSGIHHVDCEPTLHIG--PYHQAVHHEAIP 235
Query: 920 NIPRSPPGGENNFMLGWV 973
P + N + + WV
Sbjct: 236 GPPATHSEPHNQY-IWWV 252
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 12, 2003 8:28 PM
Number of letters in database: 397,940,306
Number of sequences in database: 1,242,019
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 983,538,076
Number of Sequences: 1242019
Number of extensions: 22069994
Number of successful extensions: 69330
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 63185
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 68259
length of database: 397,940,306
effective HSP length: 126
effective length of database: 241,445,912
effective search space used: 76538354104
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

