BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
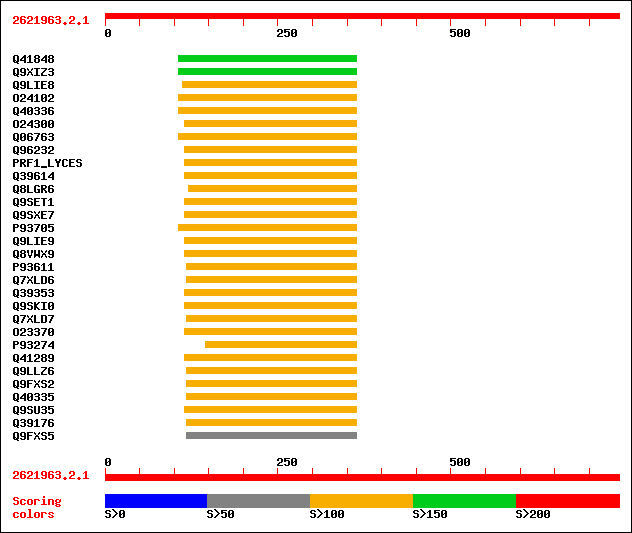
Query= 2621963.2.1
(743 letters)
Database: /db/trembl-ebi/tmp/swall
1,302,759 sequences; 415,658,970 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q41848|Q41848 Prolin rich protein. 178 8e-44
sptr|Q9XIZ3|Q9XIZ3 Similar to Zea mays PRP gene.(X60432). 161 1e-38
sptr|Q9LIE8|Q9LIE8 Similarity to cell wall-plasma membrane linke... 129 3e-29
sptr|O24102|O24102 MtN4 protein (Fragment). 124 1e-27
sptr|Q40336|Q40336 Proline-rich cell wall protein. 124 1e-27
sptr|O24300|O24300 PtxA protein precursor. 123 3e-27
sptr|Q06763|Q06763 ADR11 protein (Fragment). 121 8e-27
sptr|Q96232|Q96232 Proline-rich-like protein. 121 1e-26
sw|Q00451|PRF1_LYCES 36.4 kDa proline-rich protein. 120 2e-26
sptr|Q39614|Q39614 Proline-rich protein. 120 2e-26
sptr|Q8LGR6|Q8LGR6 Proline-rich protein 1. 119 5e-26
sptr|Q9SET1|Q9SET1 Cell wall-plasma membrane linker protein homo... 118 7e-26
sptr|Q9SXE7|Q9SXE7 T3P18.6 (Putative proline-rich cell wall prot... 118 7e-26
sptr|P93705|P93705 Hypocotil specific protein. 118 7e-26
sptr|Q9LIE9|Q9LIE9 Similarity to cell wall-plasma membrane linke... 118 7e-26
sptr|Q8VWX9|Q8VWX9 Extensin-like protein (Fragment). 118 9e-26
sptr|P93611|P93611 Cold acclimation protein WCOR518 (Fragment). 118 9e-26
sptr|Q7XLD6|Q7XLD6 OSJNBa0070C17.12 protein. 117 2e-25
sptr|Q39353|Q39353 Cell wall-plasma membrane linker protein. 117 2e-25
sptr|Q9SKI0|Q9SKI0 At2g10940 protein (Hypothetical protein) (At2... 115 8e-25
sptr|Q7XLD7|Q7XLD7 OSJNBa0070C17.11 protein. 115 8e-25
sptr|O23370|O23370 COLL wall protein homolog (Cell wall protein ... 112 4e-24
sptr|P93274|P93274 Proline rich protein (Fragment). 112 5e-24
sptr|Q41289|Q41289 Putative proline-rich protein. 111 9e-24
sptr|Q9LLZ6|Q9LLZ6 Proline-rich protein. 110 1e-23
sptr|Q9FXS2|Q9FXS2 P-rich protein Nt-SubC29. 109 4e-23
sptr|Q40335|Q40335 Bimodular protein. 107 2e-22
sptr|Q9SU35|Q9SU35 PEARLI 1-like protein. 102 7e-21
sptr|Q39176|Q39176 PEARLI 1 (AT4G12480/T1P17_70). 101 1e-20
sptr|Q9FXS5|Q9FXS5 P-rich protein EIG-I30. 100 2e-20
>sptr|Q41848|Q41848 Prolin rich protein.
Length = 301
Score = 178 bits (451), Expect = 8e-44
Identities = 86/86 (100%), Positives = 86/86 (100%)
Frame = +1
Query: 106 AVRTCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLL 285
AVRTCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLL
Sbjct: 211 AVRTCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLL 270
Query: 286 NINIYLPIALNLLITCGKHAPSGFQC 363
NINIYLPIALNLLITCGKHAPSGFQC
Sbjct: 271 NINIYLPIALNLLITCGKHAPSGFQC 296
>sptr|Q9XIZ3|Q9XIZ3 Similar to Zea mays PRP gene.(X60432).
Length = 333
Score = 161 bits (407), Expect = 1e-38
Identities = 75/86 (87%), Positives = 80/86 (93%)
Frame = +1
Query: 106 AVRTCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLL 285
A +TCPID LKLNACVDVL GLIHLVIGQ+AR+KCCPLVQGVADLDAALCLCTTIRARLL
Sbjct: 242 ATKTCPIDALKLNACVDVLGGLIHLVIGQKARAKCCPLVQGVADLDAALCLCTTIRARLL 301
Query: 286 NINIYLPIALNLLITCGKHAPSGFQC 363
NINIYLP+AL LLITCGKH P GF+C
Sbjct: 302 NINIYLPVALELLITCGKHPPPGFKC 327
>sptr|Q9LIE8|Q9LIE8 Similarity to cell wall-plasma membrane linker
protein.
Length = 1480
Score = 129 bits (325), Expect = 3e-29
Identities = 54/84 (64%), Positives = 73/84 (86%)
Frame = +1
Query: 112 RTCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNI 291
+TCPIDTLKL +CVD+L GL+H+ IG+ A+ KCCP+V+G+ DLDAA+CLCTTI+A+LLNI
Sbjct: 1395 KTCPIDTLKLGSCVDLLGGLVHIGIGKSAKEKCCPVVEGLVDLDAAVCLCTTIKAKLLNI 1454
Query: 292 NIYLPIALNLLITCGKHAPSGFQC 363
++ LPIAL +L+ CGK+ P GF+C
Sbjct: 1455 DVILPIALEVLLNCGKNPPPGFKC 1478
>sptr|O24102|O24102 MtN4 protein (Fragment).
Length = 249
Score = 124 bits (312), Expect = 1e-27
Identities = 56/86 (65%), Positives = 68/86 (79%)
Frame = +1
Query: 106 AVRTCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLL 285
A +TC ID LKL ACVDVL GLIH+ IG A+ CCPL+QG+ DLDAA+CLCTTIR +LL
Sbjct: 161 AQQTCSIDALKLGACVDVLGGLIHIGIGGSAKQTCCPLLQGLVDLDAAICLCTTIRLKLL 220
Query: 286 NINIYLPIALNLLITCGKHAPSGFQC 363
NIN+ +P+AL +LI CGK P GF+C
Sbjct: 221 NINLVIPLALQVLIDCGKTPPEGFKC 246
>sptr|Q40336|Q40336 Proline-rich cell wall protein.
Length = 381
Score = 124 bits (312), Expect = 1e-27
Identities = 56/86 (65%), Positives = 68/86 (79%)
Frame = +1
Query: 106 AVRTCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLL 285
A +TC ID LKL ACVDVL GLIH+ IG A+ CCPL+QG+ DLDAA+CLCTTIR +LL
Sbjct: 293 AQQTCSIDALKLGACVDVLGGLIHIGIGGSAKQTCCPLLQGLVDLDAAICLCTTIRLKLL 352
Query: 286 NINIYLPIALNLLITCGKHAPSGFQC 363
NIN+ +P+AL +LI CGK P GF+C
Sbjct: 353 NINLVIPLALQVLIDCGKTPPEGFKC 378
>sptr|O24300|O24300 PtxA protein precursor.
Length = 352
Score = 123 bits (308), Expect = 3e-27
Identities = 55/83 (66%), Positives = 66/83 (79%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNIN 294
TC ID LKL ACVDVL GLIH+ IG A+ CCPL+QG+ DLDAA+CLCTTIR +LLNIN
Sbjct: 267 TCSIDALKLGACVDVLGGLIHIGIGGSAKQTCCPLLQGLVDLDAAVCLCTTIRLKLLNIN 326
Query: 295 IYLPIALNLLITCGKHAPSGFQC 363
+ +P+AL +LI CGK P GF+C
Sbjct: 327 LVIPLALQVLIDCGKTPPEGFKC 349
>sptr|Q06763|Q06763 ADR11 protein (Fragment).
Length = 151
Score = 121 bits (304), Expect = 8e-27
Identities = 49/86 (56%), Positives = 70/86 (81%)
Frame = +1
Query: 106 AVRTCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLL 285
A TCPIDTLKL ACVD+L GL+H+ +G ++CCP++QG+ +++AA+CLCTT++ +LL
Sbjct: 64 AQATCPIDTLKLGACVDLLGGLVHIGLGDPVANQCCPVLQGLVEVEAAVCLCTTLKLKLL 123
Query: 286 NINIYLPIALNLLITCGKHAPSGFQC 363
N+NIY+P+AL LL+TCGK P G+ C
Sbjct: 124 NLNIYVPLALQLLVTCGKSPPPGYTC 149
>sptr|Q96232|Q96232 Proline-rich-like protein.
Length = 184
Score = 121 bits (303), Expect = 1e-26
Identities = 49/83 (59%), Positives = 68/83 (81%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNIN 294
TCP+D LKL ACVD+L GL+H+ +G ++CCPL++G+ +++AA+CLCTTIR +LLNIN
Sbjct: 100 TCPLDALKLGACVDLLGGLVHIGLGDPVVNQCCPLIEGLVEIEAAVCLCTTIRLKLLNIN 159
Query: 295 IYLPIALNLLITCGKHAPSGFQC 363
+YLP+AL LL+TCGK P G+ C
Sbjct: 160 LYLPLALQLLLTCGKTPPPGYTC 182
>sw|Q00451|PRF1_LYCES 36.4 kDa proline-rich protein.
Length = 346
Score = 120 bits (301), Expect = 2e-26
Identities = 57/84 (67%), Positives = 66/84 (78%), Gaps = 1/84 (1%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNIN 294
TCPID LKL ACVDVL GLIH+ IG A+ CCPL+ G+ DLDAA+CLCTTIR +LLNIN
Sbjct: 260 TCPIDALKLGACVDVLGGLIHIGIGGSAKQTCCPLLGGLVDLDAAICLCTTIRLKLLNIN 319
Query: 295 IYLPIALNLLI-TCGKHAPSGFQC 363
I LPIAL +LI CGK+ P F+C
Sbjct: 320 IILPIALQVLIDDCGKYPPKDFKC 343
>sptr|Q39614|Q39614 Proline-rich protein.
Length = 329
Score = 120 bits (300), Expect = 2e-26
Identities = 56/84 (66%), Positives = 68/84 (80%), Gaps = 1/84 (1%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNIN 294
TCPID LKLNACVD+L GLIH+ IG+ A+ CCP++ G+A LDA +CLCTTI+A+LLNIN
Sbjct: 242 TCPIDALKLNACVDLLGGLIHIGIGRSAKDTCCPVLGGLAGLDAGICLCTTIKAKLLNIN 301
Query: 295 IYLPIALNLLI-TCGKHAPSGFQC 363
I LPIAL +LI CG P+GFQC
Sbjct: 302 IILPIALQVLIDDCGMIPPAGFQC 325
>sptr|Q8LGR6|Q8LGR6 Proline-rich protein 1.
Length = 189
Score = 119 bits (297), Expect = 5e-26
Identities = 51/81 (62%), Positives = 65/81 (80%)
Frame = +1
Query: 121 PIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNINIY 300
P DTLKL ACVD+L GL+H+ IG A+ CCP++QG+ DLDAA+CLCT I+ +LLN+NI
Sbjct: 107 PHDTLKLGACVDLLGGLVHIGIGSSAKDTCCPVLQGLVDLDAAVCLCTAIKVKLLNVNII 166
Query: 301 LPIALNLLITCGKHAPSGFQC 363
+PIAL +L+ CGK PSGFQC
Sbjct: 167 IPIALQVLVGCGKTPPSGFQC 187
>sptr|Q9SET1|Q9SET1 Cell wall-plasma membrane linker protein
homolog.
Length = 306
Score = 118 bits (296), Expect = 7e-26
Identities = 54/84 (64%), Positives = 70/84 (83%), Gaps = 1/84 (1%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIGQE-ARSKCCPLVQGVADLDAALCLCTTIRARLLNI 291
TCPIDTLKL ACVDVL GLIH+ +G+ A+++CCP++ G+ DLDAA+CLCTTI+ +LLNI
Sbjct: 221 TCPIDTLKLGACVDVLGGLIHIGLGKSHAKAECCPVLGGLLDLDAAVCLCTTIKLKLLNI 280
Query: 292 NIYLPIALNLLITCGKHAPSGFQC 363
++ LPIAL LL+ CGK PS F+C
Sbjct: 281 DLVLPIALELLLDCGKTPPSDFKC 304
>sptr|Q9SXE7|Q9SXE7 T3P18.6 (Putative proline-rich cell wall
protein).
Length = 297
Score = 118 bits (296), Expect = 7e-26
Identities = 51/83 (61%), Positives = 67/83 (80%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNIN 294
TCPI+ LKL ACVDVL GLIH+ +G + CCP++QG+ +L+AA+CLCTTIR +LLN+N
Sbjct: 211 TCPINALKLGACVDVLGGLIHIGLGNPVENVCCPVLQGLLELEAAVCLCTTIRLKLLNLN 270
Query: 295 IYLPIALNLLITCGKHAPSGFQC 363
I++P+AL LITCG + PSGF C
Sbjct: 271 IFIPLALQALITCGINPPSGFVC 293
>sptr|P93705|P93705 Hypocotil specific protein.
Length = 347
Score = 118 bits (296), Expect = 7e-26
Identities = 53/86 (61%), Positives = 66/86 (76%)
Frame = +1
Query: 106 AVRTCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLL 285
A TCP+D LKL ACVDVL GL+H+ +G +KCCP++ G+ +L+AA+CLCTTI+ LL
Sbjct: 260 APETCPLDALKLGACVDVLGGLVHVGLGDPNVNKCCPVLAGLVELEAAVCLCTTIKLSLL 319
Query: 286 NINIYLPIALNLLITCGKHAPSGFQC 363
NINI LP+AL LLITCGK P GF C
Sbjct: 320 NINIALPVALQLLITCGKTPPPGFTC 345
>sptr|Q9LIE9|Q9LIE9 Similarity to cell wall-plasma membrane linker
protein.
Length = 334
Score = 118 bits (296), Expect = 7e-26
Identities = 54/84 (64%), Positives = 70/84 (83%), Gaps = 1/84 (1%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIGQE-ARSKCCPLVQGVADLDAALCLCTTIRARLLNI 291
TCPIDTLKL ACVDVL GLIH+ +G+ A+++CCP++ G+ DLDAA+CLCTTI+ +LLNI
Sbjct: 249 TCPIDTLKLGACVDVLGGLIHIGLGKSHAKAECCPVLGGLLDLDAAVCLCTTIKLKLLNI 308
Query: 292 NIYLPIALNLLITCGKHAPSGFQC 363
++ LPIAL LL+ CGK PS F+C
Sbjct: 309 DLVLPIALELLLDCGKTPPSDFKC 332
>sptr|Q8VWX9|Q8VWX9 Extensin-like protein (Fragment).
Length = 219
Score = 118 bits (295), Expect = 9e-26
Identities = 49/83 (59%), Positives = 67/83 (80%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNIN 294
TCPIDTLKL CV++L GL+H+ IG A + CCP++ G+A+L+AA+CLCTT++ + L++N
Sbjct: 135 TCPIDTLKLGGCVNLLGGLVHIGIGDPAANACCPIISGLAELEAAVCLCTTLKIKALDLN 194
Query: 295 IYLPIALNLLITCGKHAPSGFQC 363
IY+PIAL LLITCGK P G+ C
Sbjct: 195 IYVPIALQLLITCGKTPPPGYTC 217
>sptr|P93611|P93611 Cold acclimation protein WCOR518 (Fragment).
Length = 315
Score = 118 bits (295), Expect = 9e-26
Identities = 55/83 (66%), Positives = 64/83 (77%), Gaps = 1/83 (1%)
Frame = +1
Query: 118 CPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNINI 297
CPIDTLKL CVD L+GL+H VIG A CCPL+ GVADLDAALCLCTTI+ + LNIN+
Sbjct: 231 CPIDTLKLLGCVDGLNGLVHAVIGSSASDSCCPLLSGVADLDAALCLCTTIKLKALNINL 290
Query: 298 YLPIALNLLIT-CGKHAPSGFQC 363
LPIA++LL+ CGK P FQC
Sbjct: 291 VLPIAIDLLVNQCGKTVPKDFQC 313
Score = 115 bits (289), Expect = 5e-25
Identities = 53/83 (63%), Positives = 62/83 (74%), Gaps = 1/83 (1%)
Frame = +1
Query: 118 CPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNINI 297
CP+DTLKL CVD L+GL+H VIG A KCCPL+ GV LDAALCLCTTI + LNIN+
Sbjct: 84 CPVDTLKLLGCVDALNGLVHAVIGSSASDKCCPLLSGVTGLDAALCLCTTIELKALNINL 143
Query: 298 YLPIALNLLIT-CGKHAPSGFQC 363
LPIA+ +L+ CGK PS FQC
Sbjct: 144 VLPIAIQVLVNQCGKTVPSDFQC 166
>sptr|Q7XLD6|Q7XLD6 OSJNBa0070C17.12 protein.
Length = 154
Score = 117 bits (293), Expect = 2e-25
Identities = 50/83 (60%), Positives = 67/83 (80%), Gaps = 1/83 (1%)
Frame = +1
Query: 118 CPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNINI 297
CP++TLKL ACVD L+GL+H V+G +A CCPL+ GVADLDAALCLCT I+A+ L +++
Sbjct: 70 CPVNTLKLLACVDALNGLVHAVVGAKASDTCCPLLSGVADLDAALCLCTAIKAKALGVSL 129
Query: 298 YLPIALNLLIT-CGKHAPSGFQC 363
LP+A+++L+ CGKH PS FQC
Sbjct: 130 VLPVAISVLVNECGKHVPSSFQC 152
>sptr|Q39353|Q39353 Cell wall-plasma membrane linker protein.
Length = 376
Score = 117 bits (292), Expect = 2e-25
Identities = 53/84 (63%), Positives = 68/84 (80%), Gaps = 1/84 (1%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIG-QEARSKCCPLVQGVADLDAALCLCTTIRARLLNI 291
TCPIDTLKL ACVDVL GLIH+ +G A+ +CCP++ G+ DLDAA+CLCTTI+A+LL +
Sbjct: 291 TCPIDTLKLGACVDVLGGLIHIGLGGSSAKKECCPVLGGLVDLDAAVCLCTTIKAKLLIV 350
Query: 292 NIYLPIALNLLITCGKHAPSGFQC 363
++ +PIAL LLI CGK P GF+C
Sbjct: 351 DLIIPIALELLIDCGKTPPPGFKC 374
>sptr|Q9SKI0|Q9SKI0 At2g10940 protein (Hypothetical protein)
(At2g10940/F15K19.1).
Length = 291
Score = 115 bits (287), Expect = 8e-25
Identities = 46/83 (55%), Positives = 68/83 (81%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNIN 294
TCPIDTLKL ACVD+L GL+ + +G A +KCCPL++G+ +++AA CLCTT++ + L++N
Sbjct: 207 TCPIDTLKLGACVDLLGGLVKIGLGDPAVNKCCPLLKGLVEVEAAACLCTTLKLKALDLN 266
Query: 295 IYLPIALNLLITCGKHAPSGFQC 363
+Y+P+AL LL+TCGK+ P G+ C
Sbjct: 267 LYVPVALQLLLTCGKNPPPGYTC 289
>sptr|Q7XLD7|Q7XLD7 OSJNBa0070C17.11 protein.
Length = 202
Score = 115 bits (287), Expect = 8e-25
Identities = 50/83 (60%), Positives = 66/83 (79%), Gaps = 1/83 (1%)
Frame = +1
Query: 118 CPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNINI 297
CP+DTLKL ACVD L+GL+H V+G A CCPL+ GVADLDAALCLCT I+A+ L +++
Sbjct: 112 CPVDTLKLLACVDALNGLVHAVVGATAGDTCCPLLSGVADLDAALCLCTAIKAKALGLSL 171
Query: 298 YLPIALNLLIT-CGKHAPSGFQC 363
LP+A+++L+ CGK+ PS FQC
Sbjct: 172 VLPVAISVLVNDCGKYVPSDFQC 194
>sptr|O23370|O23370 COLL wall protein homolog (Cell wall protein
like).
Length = 428
Score = 112 bits (281), Expect = 4e-24
Identities = 53/85 (62%), Positives = 68/85 (80%), Gaps = 2/85 (2%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIGQE-ARSKCCPLVQGVADLDAALCLCTTIRARLLNI 291
TCPID LKL ACVDVL GLIH+ +G+ A++KCCPL+ + LDAA+CLCTTIRA+LLNI
Sbjct: 180 TCPIDALKLGACVDVLGGLIHIGLGKSYAKAKCCPLLDDLVGLDAAVCLCTTIRAKLLNI 239
Query: 292 NIYLPIALNLLITCGK-HAPSGFQC 363
++ +PIAL +L+ CGK P GF+C
Sbjct: 240 DLIIPIALEVLVDCGKTPPPRGFKC 264
>sptr|P93274|P93274 Proline rich protein (Fragment).
Length = 75
Score = 112 bits (280), Expect = 5e-24
Identities = 49/73 (67%), Positives = 60/73 (82%)
Frame = +1
Query: 145 ACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNINIYLPIALNLL 324
ACVDVL GLIH+ IG A+ CCP++QG+ DLDAA+CLCTTI+A+LLNIN+ +PI L +L
Sbjct: 1 ACVDVLGGLIHIGIGSSAKDACCPVLQGLVDLDAAICLCTTIKAKLLNINLIIPIVLQVL 60
Query: 325 ITCGKHAPSGFQC 363
I CGK PSGFQC
Sbjct: 61 IDCGKTPPSGFQC 73
>sptr|Q41289|Q41289 Putative proline-rich protein.
Length = 407
Score = 111 bits (278), Expect = 9e-24
Identities = 54/84 (64%), Positives = 64/84 (76%), Gaps = 1/84 (1%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNIN 294
TCPID L + ACVDVL GLIH+ G A+ CCPL+ G+ DLDAA+CLCTTIR +LLNIN
Sbjct: 322 TCPIDALNVGACVDVLGGLIHIGTGGSAKQTCCPLL-GLVDLDAAICLCTTIRLKLLNIN 380
Query: 295 IYLPIALNLLI-TCGKHAPSGFQC 363
I LPIAL +LI CGK+ P F+C
Sbjct: 381 IILPIALQVLIDDCGKYPPKDFKC 404
>sptr|Q9LLZ6|Q9LLZ6 Proline-rich protein.
Length = 139
Score = 110 bits (276), Expect = 1e-23
Identities = 47/82 (57%), Positives = 66/82 (80%)
Frame = +1
Query: 118 CPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNINI 297
CP++ LKL ACVD+L GL+H+ +G ++CCPL+QGVA L+AALCLCTTIRA++L++N+
Sbjct: 55 CPLNALKLGACVDLLQGLVHVGLGDPVVNQCCPLIQGVAALEAALCLCTTIRAKVLSLNV 114
Query: 298 YLPIALNLLITCGKHAPSGFQC 363
LPIAL+L+ +CG P F+C
Sbjct: 115 LLPIALSLVASCGLTVPPDFKC 136
>sptr|Q9FXS2|Q9FXS2 P-rich protein Nt-SubC29.
Length = 130
Score = 109 bits (272), Expect = 4e-23
Identities = 49/83 (59%), Positives = 63/83 (75%), Gaps = 1/83 (1%)
Frame = +1
Query: 118 CPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNINI 297
CPIDTLKL C +VL L+ LVIG + CC L+QGVADL+AA+CLCT I+A +L IN+
Sbjct: 47 CPIDTLKLGVCANVLGNLLGLVIGNPPKKPCCTLIQGVADLEAAICLCTAIKANILGINL 106
Query: 298 YLPIALNLLI-TCGKHAPSGFQC 363
+P++L+LL+ CGK PSGFQC
Sbjct: 107 NVPLSLSLLLNVCGKQVPSGFQC 129
>sptr|Q40335|Q40335 Bimodular protein.
Length = 166
Score = 107 bits (266), Expect = 2e-22
Identities = 49/83 (59%), Positives = 65/83 (78%), Gaps = 1/83 (1%)
Frame = +1
Query: 118 CPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNINI 297
CP DTLKL C DVL GL+++++G A SKCC L+QG+ADLDAA+CLCT I+A +L IN+
Sbjct: 84 CPTDTLKLGVCADVL-GLVNVIVGSPASSKCCTLIQGLADLDAAVCLCTAIKANILGINL 142
Query: 298 YLPIALNLLIT-CGKHAPSGFQC 363
+PI L+LL++ C K P+GFQC
Sbjct: 143 NVPITLSLLLSACEKSIPNGFQC 165
>sptr|Q9SU35|Q9SU35 PEARLI 1-like protein.
Length = 161
Score = 102 bits (253), Expect = 7e-21
Identities = 43/84 (51%), Positives = 64/84 (76%), Gaps = 1/84 (1%)
Frame = +1
Query: 115 TCPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNIN 294
+CPID LKL C +VLS L+++ +GQ + +CC L+QG+ D+DAA+CLCT +RA +L IN
Sbjct: 77 SCPIDALKLGVCANVLSSLLNIQLGQPSSQQCCSLIQGLVDVDAAICLCTALRANVLGIN 136
Query: 295 IYLPIALNLLI-TCGKHAPSGFQC 363
+ +PI+L++L+ C + PSGFQC
Sbjct: 137 LNVPISLSVLLNVCNRKLPSGFQC 160
>sptr|Q39176|Q39176 PEARLI 1 (AT4G12480/T1P17_70).
Length = 168
Score = 101 bits (251), Expect = 1e-20
Identities = 43/83 (51%), Positives = 62/83 (74%), Gaps = 1/83 (1%)
Frame = +1
Query: 118 CPIDTLKLNACVDVLSGLIHLVIGQEARSKCCPLVQGVADLDAALCLCTTIRARLLNINI 297
CPID L+L C +VLS L+++ +GQ + CC L+QG+ DLDAA+CLCT +RA +L IN+
Sbjct: 85 CPIDALRLGVCANVLSSLLNIQLGQPSAQPCCSLIQGLVDLDAAICLCTALRANVLGINL 144
Query: 298 YLPIALNLLI-TCGKHAPSGFQC 363
+PI+L++L+ C + PSGFQC
Sbjct: 145 NVPISLSVLLNVCNRKVPSGFQC 167
>sptr|Q9FXS5|Q9FXS5 P-rich protein EIG-I30.
Length = 148
Score = 100 bits (250), Expect = 2e-20
Identities = 46/84 (54%), Positives = 64/84 (76%), Gaps = 2/84 (2%)
Frame = +1
Query: 118 CPIDTLKLNACVDVLSGLIHLVIGQE-ARSKCCPLVQGVADLDAALCLCTTIRARLLNIN 294
C DTLKL C ++L+ L+HLVIG + A++ CC L+ G+ADLDAA+CLCT I+A LL IN
Sbjct: 64 CSKDTLKLKVCANLLNDLVHLVIGSDPAKTPCCSLIHGLADLDAAVCLCTAIKANLLGIN 123
Query: 295 IYLPIALNLLI-TCGKHAPSGFQC 363
+ +P++L+LL+ CGK+ P FQC
Sbjct: 124 LNVPLSLSLLLNNCGKYVPKDFQC 147
Database: /db/trembl-ebi/tmp/swall
Posted date: Nov 7, 2003 8:22 PM
Number of letters in database: 415,658,970
Number of sequences in database: 1,302,759
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 414,228,175
Number of Sequences: 1302759
Number of extensions: 6386351
Number of successful extensions: 20379
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 18696
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 20100
length of database: 415,658,970
effective HSP length: 120
effective length of database: 259,327,890
effective search space used: 32934642030
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

