BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
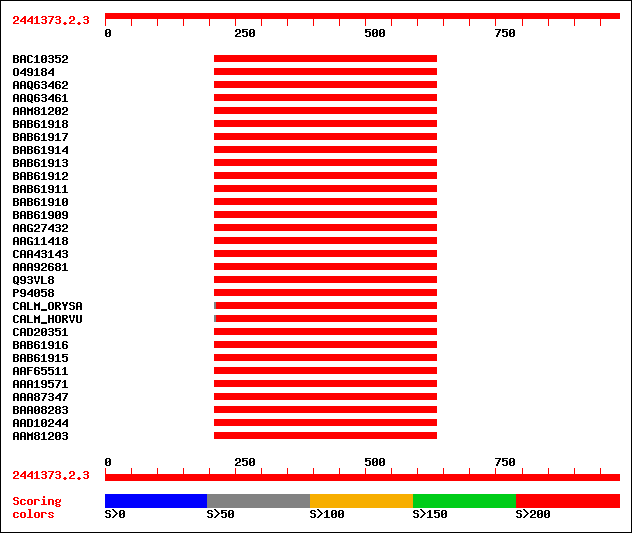
Query= 2441373.2.3
(986 letters)
Database: /db/trembl-ebi/tmp/swall
1,288,562 sequences; 411,996,502 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptrnew|BAC10352|BAC10352 Calmodulin. 285 5e-76
sptr|O49184|O49184 Calmodulin (ESTS AU081349) (Auxin-regulated c... 285 5e-76
sptrnew|AAQ63462|AAQ63462 Calmodulin 8 (Fragment). 284 1e-75
sptrnew|AAQ63461|AAQ63461 Calmodulin 4 (Fragment). 284 1e-75
sptrnew|AAM81202|AAM81202 Calmodulin 1. 284 1e-75
sptrnew|BAB61918|BAB61918 Calmodulin NtCaM12. 284 1e-75
sptrnew|BAB61917|BAB61917 Calmodulin NtCaM11. 284 1e-75
sptrnew|BAB61914|BAB61914 Calmodulin NtCaM8. 284 1e-75
sptrnew|BAB61913|BAB61913 Calmodulin NtCaM7. 284 1e-75
sptrnew|BAB61912|BAB61912 Calmodulin NtCaM6. 284 1e-75
sptrnew|BAB61911|BAB61911 Calmodulin NtCaM5. 284 1e-75
sptrnew|BAB61910|BAB61910 Calmodulin NtCaM4. 284 1e-75
sptrnew|BAB61909|BAB61909 Calmodulin NtCaM3. 284 1e-75
sptrnew|AAG27432|AAG27432 Calmodulin. 284 1e-75
sptrnew|AAG11418|AAG11418 Calmodulin. 284 1e-75
sptrnew|CAA43143|CAA43143 Calmodulin. 284 1e-75
sptrnew|AAA92681|AAA92681 Calmodulin. 284 1e-75
sptr|Q93VL8|Q93VL8 Calmodulin. 284 2e-75
sptr|P94058|P94058 Calmodulin TACAM2-3. 284 2e-75
sw|P29612|CALM_ORYSA Calmodulin. 283 2e-75
sw|P13565|CALM_HORVU Calmodulin. 283 2e-75
sptrnew|CAD20351|CAD20351 Calmodulin 2. 283 3e-75
sptrnew|BAB61916|BAB61916 Calmodulin NtCaM10. 283 3e-75
sptrnew|BAB61915|BAB61915 Calmodulin NtCaM9. 283 3e-75
sptrnew|AAF65511|AAF65511 Calmodulin. 283 3e-75
sptrnew|AAA19571|AAA19571 Calmodulin. 283 3e-75
sptrnew|AAA87347|AAA87347 Calmodulin. 283 3e-75
sptrnew|BAA08283|BAA08283 Calmodulin. 283 3e-75
sptrnew|AAD10244|AAD10244 Calmodulin. 283 3e-75
sptrnew|AAM81203|AAM81203 Calmodulin 2. 283 3e-75
>sptrnew|BAC10352|BAC10352 Calmodulin.
Length = 149
Score = 285 bits (730), Expect = 5e-76
Identities = 142/142 (100%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptr|O49184|O49184 Calmodulin (ESTS AU081349) (Auxin-regulated
calmodulin) (Calmodulin TACAM1-1) (Calmodulin TACAM1-2)
(Calmodulin TACAM1-3) (Calmodulin TACAM3-1) (Calmodulin
TACAM3-2) (Calmodulin TACAM3-3) (Calmodulin TACAM4-1)
(CaM protein).
Length = 149
Score = 285 bits (730), Expect = 5e-76
Identities = 142/142 (100%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|AAQ63462|AAQ63462 Calmodulin 8 (Fragment).
Length = 150
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|AAQ63461|AAQ63461 Calmodulin 4 (Fragment).
Length = 150
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|AAM81202|AAM81202 Calmodulin 1.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|BAB61918|BAB61918 Calmodulin NtCaM12.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|BAB61917|BAB61917 Calmodulin NtCaM11.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|BAB61914|BAB61914 Calmodulin NtCaM8.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|BAB61913|BAB61913 Calmodulin NtCaM7.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|BAB61912|BAB61912 Calmodulin NtCaM6.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|BAB61911|BAB61911 Calmodulin NtCaM5.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|BAB61910|BAB61910 Calmodulin NtCaM4.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|BAB61909|BAB61909 Calmodulin NtCaM3.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|AAG27432|AAG27432 Calmodulin.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|AAG11418|AAG11418 Calmodulin.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|CAA43143|CAA43143 Calmodulin.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|AAA92681|AAA92681 Calmodulin.
Length = 149
Score = 284 bits (727), Expect = 1e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptr|Q93VL8|Q93VL8 Calmodulin.
Length = 149
Score = 284 bits (726), Expect = 2e-75
Identities = 141/142 (99%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTD+QIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDEQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptr|P94058|P94058 Calmodulin TACAM2-3.
Length = 149
Score = 284 bits (726), Expect = 2e-75
Identities = 140/142 (98%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEM+READVDGDGQINY+EF
Sbjct: 121 EVDEMVREADVDGDGQINYDEF 142
Score = 67.0 bits (162), Expect = 4e-10
Identities = 34/77 (44%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +M+ E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMVREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYDEFVKVMMAK 149
>sw|P29612|CALM_ORYSA Calmodulin.
Length = 148
Score = 283 bits (725), Expect = 2e-75
Identities = 141/141 (100%), Positives = 141/141 (100%)
Frame = +3
Query: 213 ADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGN 392
ADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGN
Sbjct: 1 ADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGN 60
Query: 393 GTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEE 572
GTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEE
Sbjct: 61 GTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEE 120
Query: 573 VDEMIREADVDGDGQINYEEF 635
VDEMIREADVDGDGQINYEEF
Sbjct: 121 VDEMIREADVDGDGQINYEEF 141
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 72 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 131
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 132 GDGQINYEEFVKVMMAK 148
>sw|P13565|CALM_HORVU Calmodulin.
Length = 148
Score = 283 bits (725), Expect = 2e-75
Identities = 141/141 (100%), Positives = 141/141 (100%)
Frame = +3
Query: 213 ADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGN 392
ADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGN
Sbjct: 1 ADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADGN 60
Query: 393 GTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEE 572
GTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEE
Sbjct: 61 GTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEE 120
Query: 573 VDEMIREADVDGDGQINYEEF 635
VDEMIREADVDGDGQINYEEF
Sbjct: 121 VDEMIREADVDGDGQINYEEF 141
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 72 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 131
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 132 GDGQINYEEFVKVMMAK 148
>sptrnew|CAD20351|CAD20351 Calmodulin 2.
Length = 149
Score = 283 bits (724), Expect = 3e-75
Identities = 140/142 (98%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINY+EF
Sbjct: 121 EVDEMIREADVDGDGQINYDEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYDEFVKVMMAK 149
>sptrnew|BAB61916|BAB61916 Calmodulin NtCaM10.
Length = 149
Score = 283 bits (724), Expect = 3e-75
Identities = 140/142 (98%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINY+EF
Sbjct: 121 EVDEMIREADVDGDGQINYDEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYDEFVKVMMAK 149
>sptrnew|BAB61915|BAB61915 Calmodulin NtCaM9.
Length = 149
Score = 283 bits (724), Expect = 3e-75
Identities = 140/142 (98%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINY+EF
Sbjct: 121 EVDEMIREADVDGDGQINYDEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYDEFVKVMMAK 149
>sptrnew|AAF65511|AAF65511 Calmodulin.
Length = 149
Score = 283 bits (724), Expect = 3e-75
Identities = 140/142 (98%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINY+EF
Sbjct: 121 EVDEMIREADVDGDGQINYDEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYDEFVKVMMAK 149
>sptrnew|AAA19571|AAA19571 Calmodulin.
Length = 149
Score = 283 bits (724), Expect = 3e-75
Identities = 140/142 (98%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMI+EADVDGDGQINYEEF
Sbjct: 121 EVDEMIKEADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIKEADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|AAA87347|AAA87347 Calmodulin.
Length = 149
Score = 283 bits (724), Expect = 3e-75
Identities = 140/142 (98%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMI+EADVDGDGQINYEEF
Sbjct: 121 EVDEMIKEADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIKEADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|BAA08283|BAA08283 Calmodulin.
Length = 149
Score = 283 bits (724), Expect = 3e-75
Identities = 140/142 (98%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTDDQI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDDQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMI+EADVDGDGQINYEEF
Sbjct: 121 EVDEMIKEADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIKEADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|AAD10244|AAD10244 Calmodulin.
Length = 149
Score = 283 bits (723), Expect = 3e-75
Identities = 140/142 (98%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTD+QI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDEQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
>sptrnew|AAM81203|AAM81203 Calmodulin 2.
Length = 149
Score = 283 bits (723), Expect = 3e-75
Identities = 140/142 (98%), Positives = 142/142 (100%)
Frame = +3
Query: 210 MADQLTDDQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 389
MADQLTD+QI+EFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG
Sbjct: 1 MADQLTDEQISEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDADG 60
Query: 390 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 569
NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE
Sbjct: 61 NGTIDFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDE 120
Query: 570 EVDEMIREADVDGDGQINYEEF 635
EVDEMIREADVDGDGQINYEEF
Sbjct: 121 EVDEMIREADVDGDGQINYEEF 142
Score = 67.4 bits (163), Expect = 3e-10
Identities = 35/77 (45%), Positives = 50/77 (64%), Gaps = 1/77 (1%)
Frame = +3
Query: 210 MADQLTD-DQIAEFKEAFSLFDKDGDGCITTKELGTVMRSLGQNPTEAELQDMINEVDAD 386
MA ++ D D E KEAF +FDKD +G I+ EL VM +LG+ T+ E+ +MI E D D
Sbjct: 73 MARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLGEKLTDEEVDEMIREADVD 132
Query: 387 GNGTIDFPEFLNLMARK 437
G+G I++ EF+ +M K
Sbjct: 133 GDGQINYEEFVKVMMAK 149
Database: /db/trembl-ebi/tmp/swall
Posted date: Oct 24, 2003 8:22 PM
Number of letters in database: 411,996,502
Number of sequences in database: 1,288,562
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 659,044,752
Number of Sequences: 1288562
Number of extensions: 13204663
Number of successful extensions: 52808
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 44077
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 49326
length of database: 411,996,502
effective HSP length: 123
effective length of database: 253,503,376
effective search space used: 51968192080
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

