BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
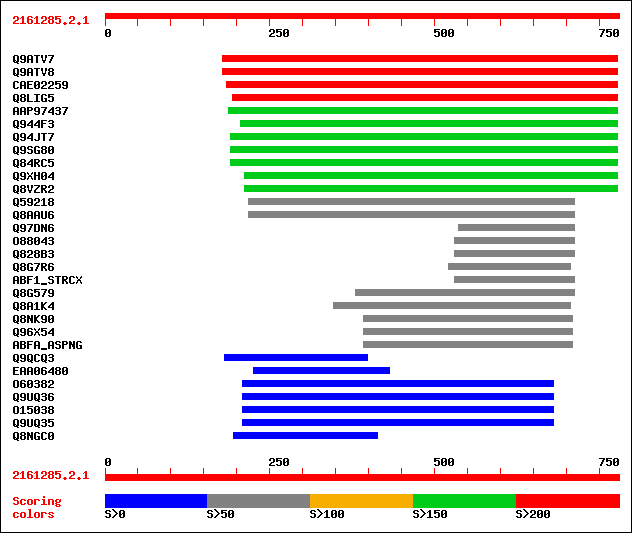
Query= 2161285.2.1
(780 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q9ATV7|Q9ATV7 Arabinoxylan arabinofuranohydrolase isoenzyme... 296 2e-79
sptr|Q9ATV8|Q9ATV8 Arabinoxylan arabinofuranohydrolase isoenzyme... 281 5e-75
sptrnew|CAE02259|CAE02259 OSJNBb0058J09.6 protein. 233 2e-60
sptr|Q8LIG5|Q8LIG5 Putative arabinoxylan narabinofuranohydrolase... 202 5e-51
sptrnew|AAP97437|AAP97437 Alpha-L-arabinofuranosidase. 192 3e-48
sptr|Q944F3|Q944F3 Arabinosidase ARA-1. 183 3e-45
sptr|Q94JT7|Q94JT7 AT3g10740/T7M13_18. 180 2e-44
sptr|Q9SG80|Q9SG80 Putative alpha-L-arabinofuranosidase. 180 2e-44
sptr|Q84RC5|Q84RC5 Alpha-L-arabinofuranosidase. 179 3e-44
sptr|Q9XH04|Q9XH04 T1N24.13 protein. 160 2e-38
sptr|Q8VZR2|Q8VZR2 Putative arabinosidase (Alpha-L-arabinofurano... 160 2e-38
sptr|Q59218|Q59218 Arabinosidase (EC 3.2.1.55). 69 5e-11
sptr|Q8AAU6|Q8AAU6 Alpha-L-arabinofuranosidase A precursor. 69 9e-11
sptr|Q97DN6|Q97DN6 Probable alpha-arabinofuranosidase. 68 2e-10
sptr|O88043|O88043 Putative secreted arabinosidase. 61 1e-08
sptr|Q828B3|Q828B3 Putative secreted arabinosidase. 59 7e-08
sptr|Q8G7R6|Q8G7R6 Similar to alpha-arabinofuranosidase I. 58 1e-07
sw|P82593|ABF1_STRCX Alpha-L-arabinofuranosidase I precursor (EC... 57 2e-07
sptr|Q8G579|Q8G579 Similar to alpha-L-arabinofuranosidase A. 57 3e-07
sptr|Q8A1K4|Q8A1K4 Alpha-L-arabinofuranosidase A precursor. 57 4e-07
sptr|Q8NK90|Q8NK90 Alpha-L-arabinofuranosidase A (EC 3.2.1.55). 54 3e-06
sptr|Q96X54|Q96X54 Alpha-L-arabinofuranosidase A (EC 3.2.1.55). 54 3e-06
sw|P42254|ABFA_ASPNG Alpha-L-arabinofuranosidase A precursor (EC... 54 3e-06
sptr|Q9QCQ3|Q9QCQ3 Thymidine kinase. 40 0.035
sptrnew|EAA06480|EAA06480 AgCP13260 (Fragment). 37 0.38
sptr|O60382|O60382 Hypothetical protein KIAA0324 (Fragment). 36 0.50
sptr|Q9UQ36|Q9UQ36 RNA binding protein (Fragment). 36 0.50
sptr|O15038|O15038 Hypothetical protein KIAA0324 (Fragment). 36 0.50
sptr|Q9UQ35|Q9UQ35 RNA binding protein. 36 0.50
sptr|Q8NGC0|Q8NGC0 Seven transmembrane helix receptor. 36 0.65
>sptr|Q9ATV7|Q9ATV7 Arabinoxylan arabinofuranohydrolase isoenzyme
AXAH-II.
Length = 656
Score = 296 bits (759), Expect = 2e-79
Identities = 148/200 (74%), Positives = 160/200 (80%)
Frame = -3
Query: 778 AVWRSDAGRGXXXXXXXXXXXLTGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQ 599
AVWR+DAGRG LTGLEKNSDIV MASYAPLFVNDNDQTWNPDAIVFNSWQ
Sbjct: 457 AVWRTDAGRGSLLGSLAEAAFLTGLEKNSDIVHMASYAPLFVNDNDQTWNPDAIVFNSWQ 516
Query: 598 HYGTPSYWMQTIFRESSGAMIHPVNIXXXXXXXXXXXAITWQDAENSFLRVKVVNFGSDP 419
YGTPSYWMQ FRESSGAMIHP+ I AITWQD+ N+FLRVK+VNFGSD
Sbjct: 517 QYGTPSYWMQKFFRESSGAMIHPITISSSYSGSLAASAITWQDSGNNFLRVKIVNFGSDT 576
Query: 418 VSLTVSTSGLGASVNALGSTATVLTSGNVMDENSFSNPNKVAPVKSQLSNAAEQMQVTLA 239
VSLT+S SGL AS+NALGS ATVLTS NV DENSFSNP KV PV SQL NAAEQMQVTLA
Sbjct: 577 VSLTISVSGLQASINALGSNATVLTSSNVKDENSFSNPTKVVPVTSQLHNAAEQMQVTLA 636
Query: 238 PHSFTAFDLALAEPKLVAEI 179
HSF++FDLALA+ +LVAE+
Sbjct: 637 AHSFSSFDLALAQSELVAEM 656
>sptr|Q9ATV8|Q9ATV8 Arabinoxylan arabinofuranohydrolase isoenzyme
AXAH-I.
Length = 658
Score = 281 bits (720), Expect = 5e-75
Identities = 136/200 (68%), Positives = 157/200 (78%)
Frame = -3
Query: 778 AVWRSDAGRGXXXXXXXXXXXLTGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQ 599
AV +DAGRG LTGLEKNSD+VQMASYAPLF+NDND+TWNPDAIVFNSWQ
Sbjct: 459 AVTGNDAGRGSLLASLAEAAFLTGLEKNSDVVQMASYAPLFINDNDRTWNPDAIVFNSWQ 518
Query: 598 HYGTPSYWMQTIFRESSGAMIHPVNIXXXXXXXXXXXAITWQDAENSFLRVKVVNFGSDP 419
YGTPSYWMQT+FRESSGA +HP+++ AITW D ENSFLRVK+VNFG D
Sbjct: 519 QYGTPSYWMQTLFRESSGATVHPISLSSSCSGLLAASAITWHDDENSFLRVKIVNFGPDA 578
Query: 418 VSLTVSTSGLGASVNALGSTATVLTSGNVMDENSFSNPNKVAPVKSQLSNAAEQMQVTLA 239
V LT+S +GL S+NA GSTATVLTSG VMDENSF+NPNKV PV +L +AAE+MQVTL
Sbjct: 579 VGLTISATGLQGSINAFGSTATVLTSGGVMDENSFANPNKVVPVTVELRDAAEEMQVTLP 638
Query: 238 PHSFTAFDLALAEPKLVAEI 179
PHS TAFDLALA+ +LVA++
Sbjct: 639 PHSLTAFDLALAQSRLVADV 658
>sptrnew|CAE02259|CAE02259 OSJNBb0058J09.6 protein.
Length = 790
Score = 233 bits (595), Expect = 2e-60
Identities = 117/198 (59%), Positives = 141/198 (71%)
Frame = -3
Query: 778 AVWRSDAGRGXXXXXXXXXXXLTGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQ 599
AV + GRG LTGLEKNSD+VQMASYAPLF+NDND +WNP A+VFNSW+
Sbjct: 593 AVSSNGVGRGTLLASLAEAAFLTGLEKNSDVVQMASYAPLFMNDNDLSWNPAAVVFNSWK 652
Query: 598 HYGTPSYWMQTIFRESSGAMIHPVNIXXXXXXXXXXXAITWQDAENSFLRVKVVNFGSDP 419
YG+PSYWMQTIFRESSGA++HPV I AITW+ + +SFLRVK+VN GS+P
Sbjct: 653 QYGSPSYWMQTIFRESSGAVLHPVTINSMYSNSLAASAITWKASNSSFLRVKIVNIGSNP 712
Query: 418 VSLTVSTSGLGASVNALGSTATVLTSGNVMDENSFSNPNKVAPVKSQLSNAAEQMQVTLA 239
V+L VST+GL A VN ST T+LTS N+ DENSFS P V PV +L NA E+M L
Sbjct: 713 VNLIVSTTGLEALVNMRKSTITILTSKNLSDENSFSKPTNVVPVTRELPNAGEEMFAFLG 772
Query: 238 PHSFTAFDLALAEPKLVA 185
P+SFT+FDLAL + K V+
Sbjct: 773 PYSFTSFDLALGQQKHVS 790
>sptr|Q8LIG5|Q8LIG5 Putative arabinoxylan narabinofuranohydrolase
isoenzyme AXAH-I.
Length = 664
Score = 202 bits (513), Expect = 5e-51
Identities = 101/197 (51%), Positives = 131/197 (66%), Gaps = 2/197 (1%)
Frame = -3
Query: 778 AVWRSDAGRGXXXXXXXXXXXLTGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQ 599
AV DAG+G L GLE+NSD+V+MAS APLFVNDND+ W+PDAIVFNSWQ
Sbjct: 464 AVTGKDAGKGTLVEALAEAAFLVGLERNSDVVEMASCAPLFVNDNDRRWSPDAIVFNSWQ 523
Query: 598 HYGTPSYWMQTIFRESSGAMIHPVNIXXXXXXXXXXXAITWQDA--ENSFLRVKVVNFGS 425
+YG P+YWM F++S GA HP NI AITWQ++ ++++L++K+VNFG+
Sbjct: 524 NYGCPNYWMLHFFKDSCGATFHPSNIQISSYNQLVASAITWQNSKDKSTYLKIKLVNFGN 583
Query: 424 DPVSLTVSTSGLGASVNALGSTATVLTSGNVMDENSFSNPNKVAPVKSQLSNAAEQMQVT 245
V+L++S SGL + + GS TVLTS +DENSF P KVAPV S + NA EQM V
Sbjct: 584 QAVNLSISVSGLDEGIKSSGSKKTVLTSSGPLDENSFQQPQKVAPVSSPVDNANEQMGVL 643
Query: 244 LAPHSFTAFDLALAEPK 194
+ P+S T+FDL L K
Sbjct: 644 VDPYSLTSFDLLLQPSK 660
>sptrnew|AAP97437|AAP97437 Alpha-L-arabinofuranosidase.
Length = 675
Score = 192 bits (489), Expect = 3e-48
Identities = 103/200 (51%), Positives = 132/200 (66%), Gaps = 3/200 (1%)
Frame = -3
Query: 778 AVWRSDAGRGXXXXXXXXXXXLTGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQ 599
AV DAGRG L GLEKNSDIV+MASYAPLFVN +D+ WNPDAIVF+S Q
Sbjct: 463 AVTGKDAGRGSLLAALAEAGFLIGLEKNSDIVEMASYAPLFVNTHDRRWNPDAIVFDSSQ 522
Query: 598 HYGTPSYWMQTIFRESSGAMIHPVNIXXXXXXXXXXXAITWQDA--ENSFLRVKVVNFGS 425
YGTPSYW+Q +F ESSGA ++ + AI+WQ++ EN++LR+KVVN G+
Sbjct: 523 LYGTPSYWVQCLFNESSGATLYNSTLQMNSSTSLLASAISWQNSENENTYLRIKVVNLGT 582
Query: 424 DPVSLTVSTSGLGA-SVNALGSTATVLTSGNVMDENSFSNPNKVAPVKSQLSNAAEQMQV 248
V+L V GL SV+ GST TVLTS N MDENSF+ P KV P +S L +A E+M+V
Sbjct: 583 YTVNLKVFVDGLEPNSVSLSGSTKTVLTSNNQMDENSFNEPKKVIPKRSLLESAGEEMEV 642
Query: 247 TLAPHSFTAFDLALAEPKLV 188
++P SFT+ DL + ++
Sbjct: 643 VISPRSFTSIDLLMESSDVI 662
>sptr|Q944F3|Q944F3 Arabinosidase ARA-1.
Length = 674
Score = 183 bits (464), Expect = 3e-45
Identities = 96/195 (49%), Positives = 123/195 (63%), Gaps = 4/195 (2%)
Frame = -3
Query: 778 AVWRSDAGRGXXXXXXXXXXXLTGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQ 599
AV +DAG+G L G+EKNS+ ++MASYAPLFVNDND+ WNPDAIVF S Q
Sbjct: 462 AVTGNDAGKGSLLAALGEAGFLIGVEKNSEAIEMASYAPLFVNDNDRRWNPDAIVFTSSQ 521
Query: 598 HYGTPSYWMQTIFRESSGAMIHPVNIXXXXXXXXXXXAITWQDA--ENSFLRVKVVNFGS 425
YGTPSYWMQ F+ES+GA + ++ AITW+++ N +LR+KVVNFG+
Sbjct: 522 MYGTPSYWMQHFFKESNGATLLSSSLQANPSNSLIASAITWRNSLDNNDYLRIKVVNFGT 581
Query: 424 DPVSLTVSTSGLGAS--VNALGSTATVLTSGNVMDENSFSNPNKVAPVKSQLSNAAEQMQ 251
V +S +GLG + G+ T LTS NVMDENSF PNKV PVK+Q+ ++ M
Sbjct: 582 TAVITKISLTGLGQNSLETLFGAVMTELTSNNVMDENSFREPNKVIPVKTQVEKVSDNMD 641
Query: 250 VTLAPHSFTAFDLAL 206
V LAP S + D L
Sbjct: 642 VVLAPRSLNSIDFLL 656
>sptr|Q94JT7|Q94JT7 AT3g10740/T7M13_18.
Length = 678
Score = 180 bits (457), Expect = 2e-44
Identities = 100/197 (50%), Positives = 121/197 (61%), Gaps = 1/197 (0%)
Frame = -3
Query: 778 AVWRSDAGRGXXXXXXXXXXXLTGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQ 599
AV DAG G L GLEKNSDIV+MASYAPLFVN ND+ WNPDAIVFNS
Sbjct: 467 AVTGKDAGTGSLLASLAEAAFLIGLEKNSDIVEMASYAPLFVNTNDRRWNPDAIVFNSSH 526
Query: 598 HYGTPSYWMQTIFRESSGAMIHPVNIXXXXXXXXXXXAITWQDAENSFLRVKVVNFGSDP 419
YGTPSYW+Q F ESSGA + + AI+W++ ++R+K VNFG++
Sbjct: 527 LYGTPSYWVQRFFAESSGATL-LTSTLKGNSTSLVASAISWKNNGKDYIRIKAVNFGANS 585
Query: 418 VSLTVSTSGLGASV-NALGSTATVLTSGNVMDENSFSNPNKVAPVKSQLSNAAEQMQVTL 242
++ V +GL +V GS TVLTS NVMDENSFS P KV P +S L A E M V L
Sbjct: 586 ENMQVLVTGLDPNVMRVSGSKKTVLTSTNVMDENSFSQPEKVVPHESLLELAEEDMTVVL 645
Query: 241 APHSFTAFDLALAEPKL 191
PHSF++FDL K+
Sbjct: 646 PPHSFSSFDLLKESAKI 662
>sptr|Q9SG80|Q9SG80 Putative alpha-L-arabinofuranosidase.
Length = 678
Score = 180 bits (457), Expect = 2e-44
Identities = 100/197 (50%), Positives = 121/197 (61%), Gaps = 1/197 (0%)
Frame = -3
Query: 778 AVWRSDAGRGXXXXXXXXXXXLTGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQ 599
AV DAG G L GLEKNSDIV+MASYAPLFVN ND+ WNPDAIVFNS
Sbjct: 467 AVTGKDAGTGSLLASLAEAAFLIGLEKNSDIVEMASYAPLFVNTNDRRWNPDAIVFNSSH 526
Query: 598 HYGTPSYWMQTIFRESSGAMIHPVNIXXXXXXXXXXXAITWQDAENSFLRVKVVNFGSDP 419
YGTPSYW+Q F ESSGA + + AI+W++ ++R+K VNFG++
Sbjct: 527 LYGTPSYWVQRFFAESSGATL-LTSTLKGNSTSLVASAISWKNNGKDYIRIKAVNFGANS 585
Query: 418 VSLTVSTSGLGASV-NALGSTATVLTSGNVMDENSFSNPNKVAPVKSQLSNAAEQMQVTL 242
++ V +GL +V GS TVLTS NVMDENSFS P KV P +S L A E M V L
Sbjct: 586 ENMQVLVTGLDPNVMRVSGSKKTVLTSTNVMDENSFSQPEKVVPHESLLELAEEDMTVVL 645
Query: 241 APHSFTAFDLALAEPKL 191
PHSF++FDL K+
Sbjct: 646 PPHSFSSFDLLKESAKI 662
>sptr|Q84RC5|Q84RC5 Alpha-L-arabinofuranosidase.
Length = 678
Score = 179 bits (455), Expect = 3e-44
Identities = 100/197 (50%), Positives = 120/197 (60%), Gaps = 1/197 (0%)
Frame = -3
Query: 778 AVWRSDAGRGXXXXXXXXXXXLTGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQ 599
AV DAG G L GLEKNSDIV MASYAPLFVN ND+ WNPDAIVFNS
Sbjct: 467 AVTGKDAGTGSLLASLAEAAFLIGLEKNSDIVDMASYAPLFVNTNDRRWNPDAIVFNSSH 526
Query: 598 HYGTPSYWMQTIFRESSGAMIHPVNIXXXXXXXXXXXAITWQDAENSFLRVKVVNFGSDP 419
YGTPSYW+Q F ESSGA + + AI+W++ ++R+K VNFG++
Sbjct: 527 LYGTPSYWVQRFFAESSGATL-LTSTLKGNSTSLVASAISWKNNGKDYIRIKAVNFGANS 585
Query: 418 VSLTVSTSGLGASV-NALGSTATVLTSGNVMDENSFSNPNKVAPVKSQLSNAAEQMQVTL 242
++ V +GL +V GS TVLTS NVMDENSFS P KV P +S L A E M V L
Sbjct: 586 ENMQVLVTGLDPNVMRVSGSKKTVLTSTNVMDENSFSQPEKVVPHESLLELAEEDMTVVL 645
Query: 241 APHSFTAFDLALAEPKL 191
PHSF++FDL K+
Sbjct: 646 PPHSFSSFDLLKESAKI 662
>sptr|Q9XH04|Q9XH04 T1N24.13 protein.
Length = 521
Score = 160 bits (404), Expect = 2e-38
Identities = 92/190 (48%), Positives = 114/190 (60%), Gaps = 1/190 (0%)
Frame = -3
Query: 778 AVWRSDAGRGXXXXXXXXXXXLTGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQ 599
AV ++DA G L GLEKNSDIV+M SYAPLFVN ND+ W PDAIVFNS
Sbjct: 313 AVNKADAKNGNLLAALGEAAFLLGLEKNSDIVEMVSYAPLFVNTNDRRWIPDAIVFNSSH 372
Query: 598 HYGTPSYWMQTIFRESSGAMIHPVNIXXXXXXXXXXXAITWQDAENSFLRVKVVNFGSDP 419
YGTPSYW+Q F ESSGA + + AI++Q ++++K VNFG
Sbjct: 373 LYGTPSYWVQHFFTESSGATLLNSTL-KGKTSSVEASAISFQTNGKDYIQIKAVNFGEQS 431
Query: 418 VSLTVSTSGLGASVNALGSTATVLTSGNVMDENSFSNPNKVAPVKSQLS-NAAEQMQVTL 242
V+L V+ +GL A GS VLTS +VMDENSFSNPN + P +S L E + L
Sbjct: 432 VNLKVAVTGLMAKF--YGSKKKVLTSASVMDENSFSNPNMIVPQESLLEMTEQEDLMFVL 489
Query: 241 APHSFTAFDL 212
PHSF++FDL
Sbjct: 490 PPHSFSSFDL 499
>sptr|Q8VZR2|Q8VZR2 Putative arabinosidase
(Alpha-L-arabinofuranosidase) (Hypothetical protein
At5g26120).
Length = 674
Score = 160 bits (404), Expect = 2e-38
Identities = 92/190 (48%), Positives = 114/190 (60%), Gaps = 1/190 (0%)
Frame = -3
Query: 778 AVWRSDAGRGXXXXXXXXXXXLTGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQ 599
AV ++DA G L GLEKNSDIV+M SYAPLFVN ND+ W PDAIVFNS
Sbjct: 466 AVNKADAKNGNLLAALGEAAFLLGLEKNSDIVEMVSYAPLFVNTNDRRWIPDAIVFNSSH 525
Query: 598 HYGTPSYWMQTIFRESSGAMIHPVNIXXXXXXXXXXXAITWQDAENSFLRVKVVNFGSDP 419
YGTPSYW+Q F ESSGA + + AI++Q ++++K VNFG
Sbjct: 526 LYGTPSYWVQHFFTESSGATLLNSTL-KGKTSSVEASAISFQTNGKDYIQIKAVNFGEQS 584
Query: 418 VSLTVSTSGLGASVNALGSTATVLTSGNVMDENSFSNPNKVAPVKSQLS-NAAEQMQVTL 242
V+L V+ +GL A GS VLTS +VMDENSFSNPN + P +S L E + L
Sbjct: 585 VNLKVAVTGLMAKF--YGSKKKVLTSASVMDENSFSNPNMIVPQESLLEMTEQEDLMFVL 642
Query: 241 APHSFTAFDL 212
PHSF++FDL
Sbjct: 643 PPHSFSSFDL 652
>sptr|Q59218|Q59218 Arabinosidase (EC 3.2.1.55).
Length = 660
Score = 69.3 bits (168), Expect = 5e-11
Identities = 43/170 (25%), Positives = 77/170 (45%), Gaps = 5/170 (2%)
Frame = -3
Query: 712 TGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQHYGTPSYWMQTIFRESSGAMIH 533
TGLE+N+DIV MA+YAPLF + W PD I F++ T SY++Q ++ ++ G +
Sbjct: 487 TGLERNADIVHMATYAPLFAHVEGWQWRPDMIWFDNLNSVRTTSYYVQQLYAQNKGTNVL 546
Query: 532 PV-----NIXXXXXXXXXXXAITWQDAENSFLRVKVVNFGSDPVSLTVSTSGLGASVNAL 368
P+ N+ + + +N + VKV N + ++++ GL
Sbjct: 547 PLTMNKKNVTGAEGQNGLFASAVYDKGKNELI-VKVANTSATIQPISLNFEGLKKQDVLS 605
Query: 367 GSTATVLTSGNVMDENSFSNPNKVAPVKSQLSNAAEQMQVTLAPHSFTAF 218
L S ++ +N+ P + P ++ +S L P +F +
Sbjct: 606 NGRCIKLRSLDLDKDNTLEQPFGIVPQETPVSIEGNVFTTELEPTTFAVY 655
>sptr|Q8AAU6|Q8AAU6 Alpha-L-arabinofuranosidase A precursor.
Length = 660
Score = 68.6 bits (166), Expect = 9e-11
Identities = 48/171 (28%), Positives = 79/171 (46%), Gaps = 6/171 (3%)
Frame = -3
Query: 712 TGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQHYGTPSYWMQTIFRESSGAMIH 533
TGLE+N+DIV MA+YAPLF + W PD I F++ T SY++Q ++ ++ G +
Sbjct: 487 TGLERNADIVHMATYAPLFAHVEGWQWRPDMIWFDNLNSVRTVSYYVQQLYAQNKGTNVL 546
Query: 532 PVNI----XXXXXXXXXXXAITWQDAENSFLRVKVVNFG--SDPVSLTVSTSGLGASVNA 371
P+ + A D + + L VKV N + PVSLT GL
Sbjct: 547 PLTMNKKPVTGAEGQNGLFASAVYDKDKNELIVKVANTSDKTQPVSLTF--EGLKKQDVL 604
Query: 370 LGSTATVLTSGNVMDENSFSNPNKVAPVKSQLSNAAEQMQVTLAPHSFTAF 218
L+S + +N+ P + P ++ ++ + L P++F +
Sbjct: 605 SEGRCITLSSLDQDKDNTLEQPFAITPQETPVTINGHALTTELGPNTFAVY 655
>sptr|Q97DN6|Q97DN6 Probable alpha-arabinofuranosidase.
Length = 835
Score = 67.8 bits (164), Expect = 2e-10
Identities = 32/59 (54%), Positives = 41/59 (69%)
Frame = -3
Query: 712 TGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQHYGTPSYWMQTIFRESSGAMI 536
TGLE+NSDIV+MASYAPLF +D W+PD I FN +Y TP+Y +Q +F + G I
Sbjct: 490 TGLERNSDIVKMASYAPLFAKADDYQWSPDMIWFNGNTNYVTPNYNVQKLFSTNLGTQI 548
>sptr|O88043|O88043 Putative secreted arabinosidase.
Length = 824
Score = 61.2 bits (147), Expect = 1e-08
Identities = 26/61 (42%), Positives = 43/61 (70%)
Frame = -3
Query: 712 TGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQHYGTPSYWMQTIFRESSGAMIH 533
TGLE+N+D+V++ASYAPL N++ W+PD I FN+ +G+ +Y +Q +F ++G +
Sbjct: 485 TGLERNADVVKLASYAPLLSNEDYVQWSPDLIWFNNHASWGSANYEVQKLFMNNTGDRVV 544
Query: 532 P 530
P
Sbjct: 545 P 545
>sptr|Q828B3|Q828B3 Putative secreted arabinosidase.
Length = 710
Score = 58.9 bits (141), Expect = 7e-08
Identities = 25/61 (40%), Positives = 41/61 (67%)
Frame = -3
Query: 712 TGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQHYGTPSYWMQTIFRESSGAMIH 533
TGLE+N+D+V++ASYAPL N++ W PD I F++ +G+ Y +Q +F ++G +
Sbjct: 485 TGLERNADVVKLASYAPLLANEDYVQWRPDMIWFDNHASWGSADYEVQKLFMTNTGDRVV 544
Query: 532 P 530
P
Sbjct: 545 P 545
>sptr|Q8G7R6|Q8G7R6 Similar to alpha-arabinofuranosidase I.
Length = 823
Score = 58.2 bits (139), Expect = 1e-07
Identities = 25/62 (40%), Positives = 38/62 (61%)
Frame = -3
Query: 706 LEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQHYGTPSYWMQTIFRESSGAMIHPV 527
+E N D+V MASYAPLF + +WNPD I F++ Y SYW+Q ++ ++ PV
Sbjct: 503 MELNGDVVHMASYAPLFAKNGHTSWNPDLIYFDNENVYRPYSYWVQQMYATTTADTAWPV 562
Query: 526 NI 521
++
Sbjct: 563 SL 564
>sw|P82593|ABF1_STRCX Alpha-L-arabinofuranosidase I precursor (EC
3.2.1.55) (Arabinosidase I) (Alpha-L-AFase I).
Length = 825
Score = 57.4 bits (137), Expect = 2e-07
Identities = 24/61 (39%), Positives = 40/61 (65%)
Frame = -3
Query: 712 TGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQHYGTPSYWMQTIFRESSGAMIH 533
TGLE+N+D+V++ASYAPL N++ W PD + FN+ + + +Y +Q +F + G +
Sbjct: 486 TGLERNADVVKLASYAPLLANEDYVQWRPDLVWFNNRASWNSANYEVQKLFMNNVGDRVV 545
Query: 532 P 530
P
Sbjct: 546 P 546
>sptr|Q8G579|Q8G579 Similar to alpha-L-arabinofuranosidase A.
Length = 769
Score = 57.0 bits (136), Expect = 3e-07
Identities = 36/117 (30%), Positives = 57/117 (48%), Gaps = 6/117 (5%)
Frame = -3
Query: 712 TGLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQHYGTPSYWMQTIFRESSGAMIH 533
TG E+NSD+V+M SYAPL + + W + I FN T +Y ++ +F G +
Sbjct: 590 TGCERNSDVVRMTSYAPLLAHIPAKGWAQNLIEFNPAHVNPTVNYEVERLFSTHLGDTTY 649
Query: 532 PVNIXXXXXXXXXXXAI--TWQDAENSFLRVKVVNFGSDPVSLTV----STSGLGAS 380
V+I + T D +++ +K+VN PV +T+ +GLGAS
Sbjct: 650 AVSIEQTASRPAKHLYVSATGHDRDDACRYIKIVNISDSPVDVTLEIARGLAGLGAS 706
>sptr|Q8A1K4|Q8A1K4 Alpha-L-arabinofuranosidase A precursor.
Length = 825
Score = 56.6 bits (135), Expect = 4e-07
Identities = 38/123 (30%), Positives = 58/123 (47%), Gaps = 3/123 (2%)
Frame = -3
Query: 706 LEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQHYGTPSYWMQTIFRESSGAMIHPV 527
LE+N D+V M+SYAPL D WNPD I FN+ + T Y++Q ++ +++G P
Sbjct: 661 LERNGDVVHMSSYAPLLAKDGRTQWNPDLIYFNNREVRPTTGYYVQKLYGQNAGDHYIPS 720
Query: 526 NIXXXXXXXXXXXAI---TWQDAENSFLRVKVVNFGSDPVSLTVSTSGLGASVNALGSTA 356
I + +D++ + VK+VN PVS+ G +T
Sbjct: 721 QINLDNQDGRVKLRVGSSIVRDSKTGDVIVKLVNL--LPVSIETDVRLPGMDGIQSSATR 778
Query: 355 TVL 347
TVL
Sbjct: 779 TVL 781
>sptr|Q8NK90|Q8NK90 Alpha-L-arabinofuranosidase A (EC 3.2.1.55).
Length = 628
Score = 53.5 bits (127), Expect = 3e-06
Identities = 37/110 (33%), Positives = 54/110 (49%), Gaps = 4/110 (3%)
Frame = -3
Query: 709 GLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQH--YGTPSYWMQTIFRESSGAMI 536
G E+NSD+V+MA+YAPL N W PD I + + + SY++Q +F + G I
Sbjct: 472 GFERNSDVVKMAAYAPLLQLVNSTQWTPDLIGYTQSPDDIFLSTSYYVQEMFSRNRGDTI 531
Query: 535 HPVNIXXXXXXXXXXXAITW--QDAENSFLRVKVVNFGSDPVSLTVSTSG 392
V + W A +S+ VK+ N+GS+ LTVS G
Sbjct: 532 KEV------TSDSDFGPLYWVASSAGDSYY-VKLANYGSETQDLTVSIPG 574
>sptr|Q96X54|Q96X54 Alpha-L-arabinofuranosidase A (EC 3.2.1.55).
Length = 628
Score = 53.5 bits (127), Expect = 3e-06
Identities = 37/110 (33%), Positives = 54/110 (49%), Gaps = 4/110 (3%)
Frame = -3
Query: 709 GLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVFNSWQH--YGTPSYWMQTIFRESSGAMI 536
G E+NSD+V+MA+YAPL N W PD I + + + SY++Q +F + G I
Sbjct: 472 GFERNSDVVKMAAYAPLLQLVNSTQWTPDLIGYTQSPDDIFLSTSYYVQEMFSRNRGDTI 531
Query: 535 HPVNIXXXXXXXXXXXAITW--QDAENSFLRVKVVNFGSDPVSLTVSTSG 392
V + W A +S+ VK+ N+GS+ LTVS G
Sbjct: 532 KEV------TSDSDFGPLYWVASSAGDSYY-VKLANYGSETQDLTVSIPG 574
>sw|P42254|ABFA_ASPNG Alpha-L-arabinofuranosidase A precursor (EC
3.2.1.55) (Arabinosidase A) (ABF A).
Length = 628
Score = 53.5 bits (127), Expect = 3e-06
Identities = 37/110 (33%), Positives = 55/110 (50%), Gaps = 4/110 (3%)
Frame = -3
Query: 709 GLEKNSDIVQMASYAPLFVNDNDQTWNPDAIVF--NSWQHYGTPSYWMQTIFRESSGAMI 536
G E+NSD+V+MA+YAPL N W PD I + + + + SY++Q +F + G I
Sbjct: 472 GFERNSDVVKMAAYAPLLQLINSTQWTPDLIGYTQSPGDIFLSTSYYVQEMFSRNRGDTI 531
Query: 535 HPVNIXXXXXXXXXXXAITW--QDAENSFLRVKVVNFGSDPVSLTVSTSG 392
V + W A +S+ VK+ N+GS+ LTVS G
Sbjct: 532 KEV------TSDSDFGPLYWVASSAGDSYY-VKLANYGSETQDLTVSIPG 574
>sptr|Q9QCQ3|Q9QCQ3 Thymidine kinase.
Length = 297
Score = 40.0 bits (92), Expect = 0.035
Identities = 32/84 (38%), Positives = 40/84 (47%), Gaps = 11/84 (13%)
Frame = -1
Query: 399 RAGSGPASTRWDQPPPSSRLATSWTRTPSVTRTRLRR*KAS*ATPRSRC-RSRWLLT--- 232
RA + P +T ++R +RTPS RTR R A+ A RSRC R RW +
Sbjct: 209 RASTRPCATASRSTSSAARATAPSSRTPSSARTR-RPSSATGAGARSRCTRGRWTRSWPS 267
Query: 231 ----PSPPSIW---RWLSPSLWRR 181
SPPS W R +P WRR
Sbjct: 268 CCRCASPPSTWGLRRASAPRPWRR 291
>sptrnew|EAA06480|EAA06480 AgCP13260 (Fragment).
Length = 1320
Score = 36.6 bits (83), Expect = 0.38
Identities = 24/71 (33%), Positives = 27/71 (38%), Gaps = 2/71 (2%)
Frame = -1
Query: 432 SGRTPLASRSPRAGSGPASTRWDQPPPSSRLAT--SWTRTPSVTRTRLRR*KAS*ATPRS 259
SG R A RWD+ P + R SW TP R TP S
Sbjct: 267 SGAATTPGRETPAEKSTRRNRWDETPKTERETPGHSWAETPRADRVSGDGVLLEGTTPAS 326
Query: 258 RCRSRWLLTPS 226
+ RSRW TPS
Sbjct: 327 KRRSRWDETPS 337
>sptr|O60382|O60382 Hypothetical protein KIAA0324 (Fragment).
Length = 1791
Score = 36.2 bits (82), Expect = 0.50
Identities = 46/155 (29%), Positives = 71/155 (45%), Gaps = 9/155 (5%)
Frame = -1
Query: 645 MTKRGTRTLSSSTPGSTMELLATGCRRFSASQAAP*SIRLISVPGTRVRWPRLRSRGRMQ 466
+T+R +R+ + +T + RR S S+ P + R + + R RSR R
Sbjct: 962 VTRRRSRSRTPTTRRRSRSRTPPVTRRRSRSRTPPVTRRRSRSRTSPIT--RRRSRSRTS 1019
Query: 465 RIAS*E*RS*TSGRTPLASRSPRAGSGPASTR--WDQPPPSSRLATSWTRTPSVTRTRLR 292
+ RS TS P+ R R+ + P + R + PP+ R S +RTP + R R R
Sbjct: 1020 PVTRRRSRSRTS---PVTRRRSRSRTSPVTRRRSRSRTPPAIR-RRSRSRTPLLPRKRSR 1075
Query: 291 -------R*KAS*ATPRSRCRSRWLLTPSPPSIWR 208
R ++ TPR+ R + LT SPP+I R
Sbjct: 1076 SRSPLAIRRRSRSRTPRT-ARGKRSLTRSPPAIRR 1109
Score = 33.9 bits (76), Expect = 2.5
Identities = 41/114 (35%), Positives = 53/114 (46%)
Frame = -1
Query: 570 RRFSASQAAP*SIRLISVPGTRVRWPRLRSRGRMQRIAS*E*RS*TSGRTPLASRSPRAG 391
RR S S+ +P S R +R R RSR R ++ RS T T SRS R
Sbjct: 917 RRRSRSRTSPVSRRRSR---SRTSVTRRRSRSRASPVSRRRSRSRTPPVTRRRSRS-RTP 972
Query: 390 SGPASTRWDQPPPSSRLATSWTRTPSVTRTRLRR*KAS*ATPRSRCRSRWLLTP 229
+ +R PP + R + S RTP VTR R R S +P +R RSR +P
Sbjct: 973 TTRRRSRSRTPPVTRRRSRS--RTPPVTRRRSR----SRTSPITRRRSRSRTSP 1020
Score = 33.1 bits (74), Expect = 4.2
Identities = 51/181 (28%), Positives = 67/181 (37%), Gaps = 30/181 (16%)
Frame = -1
Query: 681 RWQVTHHSS*TTMTKRGTRTLSSSTPGSTMELLATGCRRFSASQAAP*SIRLISVPGTRV 502
R Q + S T +RG S +P + RR S+ P S + +
Sbjct: 832 RRQRSRSRSRVTRRRRGGSGYHSRSPARQESSRTSSRRRRGRSRTPPTSRKRSRSRTSPA 891
Query: 501 RWPRLRSRG----------RMQRIAS*E*RS*TS--------GRTPLASRSPRAGSGPAS 376
W R RSR R I+ RS TS RT + R R+ + P S
Sbjct: 892 PWKRSRSRASPATHRRSRSRTPLISRRRSRSRTSPVSRRRSRSRTSVTRRRSRSRASPVS 951
Query: 375 TRWDQ---PPPSSRLATSWT---------RTPSVTRTRLRR*KAS*ATPRSRCRSRWLLT 232
R + PP + R + S T RTP VTR R R S P +R RSR +
Sbjct: 952 RRRSRSRTPPVTRRRSRSRTPTTRRRSRSRTPPVTRRRSR----SRTPPVTRRRSRSRTS 1007
Query: 231 P 229
P
Sbjct: 1008 P 1008
>sptr|Q9UQ36|Q9UQ36 RNA binding protein (Fragment).
Length = 1275
Score = 36.2 bits (82), Expect = 0.50
Identities = 46/155 (29%), Positives = 71/155 (45%), Gaps = 9/155 (5%)
Frame = -1
Query: 645 MTKRGTRTLSSSTPGSTMELLATGCRRFSASQAAP*SIRLISVPGTRVRWPRLRSRGRMQ 466
+T+R +R+ + +T + RR S S+ P + R + + R RSR R
Sbjct: 453 VTRRRSRSRTPTTRRRSRSRTPPVTRRRSRSRTPPVTRRRSRSRTSPIT--RRRSRSRTS 510
Query: 465 RIAS*E*RS*TSGRTPLASRSPRAGSGPASTR--WDQPPPSSRLATSWTRTPSVTRTRLR 292
+ RS TS P+ R R+ + P + R + PP+ R S +RTP + R R R
Sbjct: 511 PVTRRRSRSRTS---PVTRRRSRSRTSPVTRRRSRSRTPPAIR-RRSRSRTPLLPRKRSR 566
Query: 291 -------R*KAS*ATPRSRCRSRWLLTPSPPSIWR 208
R ++ TPR+ R + LT SPP+I R
Sbjct: 567 SRSPLAIRRRSRSRTPRT-ARGKRSLTRSPPAIRR 600
Score = 33.9 bits (76), Expect = 2.5
Identities = 41/114 (35%), Positives = 53/114 (46%)
Frame = -1
Query: 570 RRFSASQAAP*SIRLISVPGTRVRWPRLRSRGRMQRIAS*E*RS*TSGRTPLASRSPRAG 391
RR S S+ +P S R +R R RSR R ++ RS T T SRS R
Sbjct: 408 RRRSRSRTSPVSRRRSR---SRTSVTRRRSRSRASPVSRRRSRSRTPPVTRRRSRS-RTP 463
Query: 390 SGPASTRWDQPPPSSRLATSWTRTPSVTRTRLRR*KAS*ATPRSRCRSRWLLTP 229
+ +R PP + R + S RTP VTR R R S +P +R RSR +P
Sbjct: 464 TTRRRSRSRTPPVTRRRSRS--RTPPVTRRRSR----SRTSPITRRRSRSRTSP 511
Score = 33.1 bits (74), Expect = 4.2
Identities = 51/181 (28%), Positives = 67/181 (37%), Gaps = 30/181 (16%)
Frame = -1
Query: 681 RWQVTHHSS*TTMTKRGTRTLSSSTPGSTMELLATGCRRFSASQAAP*SIRLISVPGTRV 502
R Q + S T +RG S +P + RR S+ P S + +
Sbjct: 323 RRQRSRSRSRVTRRRRGGSGYHSRSPARQESSRTSSRRRRGRSRTPPTSRKRSRSRTSPA 382
Query: 501 RWPRLRSRG----------RMQRIAS*E*RS*TS--------GRTPLASRSPRAGSGPAS 376
W R RSR R I+ RS TS RT + R R+ + P S
Sbjct: 383 PWKRSRSRASPATHRRSRSRTPLISRRRSRSRTSPVSRRRSRSRTSVTRRRSRSRASPVS 442
Query: 375 TRWDQ---PPPSSRLATSWT---------RTPSVTRTRLRR*KAS*ATPRSRCRSRWLLT 232
R + PP + R + S T RTP VTR R R S P +R RSR +
Sbjct: 443 RRRSRSRTPPVTRRRSRSRTPTTRRRSRSRTPPVTRRRSR----SRTPPVTRRRSRSRTS 498
Query: 231 P 229
P
Sbjct: 499 P 499
>sptr|O15038|O15038 Hypothetical protein KIAA0324 (Fragment).
Length = 1783
Score = 36.2 bits (82), Expect = 0.50
Identities = 46/155 (29%), Positives = 71/155 (45%), Gaps = 9/155 (5%)
Frame = -1
Query: 645 MTKRGTRTLSSSTPGSTMELLATGCRRFSASQAAP*SIRLISVPGTRVRWPRLRSRGRMQ 466
+T+R +R+ + +T + RR S S+ P + R + + R RSR R
Sbjct: 961 VTRRRSRSRTPTTRRRSRSRTPPVTRRRSRSRTPPVTRRRSRSRTSPIT--RRRSRSRTS 1018
Query: 465 RIAS*E*RS*TSGRTPLASRSPRAGSGPASTR--WDQPPPSSRLATSWTRTPSVTRTRLR 292
+ RS TS P+ R R+ + P + R + PP+ R S +RTP + R R R
Sbjct: 1019 PVTRRRSRSRTS---PVTRRRSRSRTSPVTRRRSRSRTPPAIR-RRSRSRTPLLPRKRSR 1074
Query: 291 -------R*KAS*ATPRSRCRSRWLLTPSPPSIWR 208
R ++ TPR+ R + LT SPP+I R
Sbjct: 1075 SRSPLAIRRRSRSRTPRT-ARGKRSLTRSPPAIRR 1108
Score = 33.9 bits (76), Expect = 2.5
Identities = 41/114 (35%), Positives = 53/114 (46%)
Frame = -1
Query: 570 RRFSASQAAP*SIRLISVPGTRVRWPRLRSRGRMQRIAS*E*RS*TSGRTPLASRSPRAG 391
RR S S+ +P S R +R R RSR R ++ RS T T SRS R
Sbjct: 916 RRRSRSRTSPVSRRRSR---SRTSVTRRRSRSRASPVSRRRSRSRTPPVTRRRSRS-RTP 971
Query: 390 SGPASTRWDQPPPSSRLATSWTRTPSVTRTRLRR*KAS*ATPRSRCRSRWLLTP 229
+ +R PP + R + S RTP VTR R R S +P +R RSR +P
Sbjct: 972 TTRRRSRSRTPPVTRRRSRS--RTPPVTRRRSR----SRTSPITRRRSRSRTSP 1019
Score = 33.1 bits (74), Expect = 4.2
Identities = 51/181 (28%), Positives = 67/181 (37%), Gaps = 30/181 (16%)
Frame = -1
Query: 681 RWQVTHHSS*TTMTKRGTRTLSSSTPGSTMELLATGCRRFSASQAAP*SIRLISVPGTRV 502
R Q + S T +RG S +P + RR S+ P S + +
Sbjct: 831 RRQRSRSRSRVTRRRRGGSGYHSRSPARQESSRTSSRRRRGRSRTPPTSRKRSRSRTSPA 890
Query: 501 RWPRLRSRG----------RMQRIAS*E*RS*TS--------GRTPLASRSPRAGSGPAS 376
W R RSR R I+ RS TS RT + R R+ + P S
Sbjct: 891 PWKRSRSRASPATHRRSRSRTPLISRRRSRSRTSPVSRRRSRSRTSVTRRRSRSRASPVS 950
Query: 375 TRWDQ---PPPSSRLATSWT---------RTPSVTRTRLRR*KAS*ATPRSRCRSRWLLT 232
R + PP + R + S T RTP VTR R R S P +R RSR +
Sbjct: 951 RRRSRSRTPPVTRRRSRSRTPTTRRRSRSRTPPVTRRRSR----SRTPPVTRRRSRSRTS 1006
Query: 231 P 229
P
Sbjct: 1007 P 1007
>sptr|Q9UQ35|Q9UQ35 RNA binding protein.
Length = 2752
Score = 36.2 bits (82), Expect = 0.50
Identities = 46/155 (29%), Positives = 71/155 (45%), Gaps = 9/155 (5%)
Frame = -1
Query: 645 MTKRGTRTLSSSTPGSTMELLATGCRRFSASQAAP*SIRLISVPGTRVRWPRLRSRGRMQ 466
+T+R +R+ + +T + RR S S+ P + R + + R RSR R
Sbjct: 1930 VTRRRSRSRTPTTRRRSRSRTPPVTRRRSRSRTPPVTRRRSRSRTSPIT--RRRSRSRTS 1987
Query: 465 RIAS*E*RS*TSGRTPLASRSPRAGSGPASTR--WDQPPPSSRLATSWTRTPSVTRTRLR 292
+ RS TS P+ R R+ + P + R + PP+ R S +RTP + R R R
Sbjct: 1988 PVTRRRSRSRTS---PVTRRRSRSRTSPVTRRRSRSRTPPAIR-RRSRSRTPLLPRKRSR 2043
Query: 291 -------R*KAS*ATPRSRCRSRWLLTPSPPSIWR 208
R ++ TPR+ R + LT SPP+I R
Sbjct: 2044 SRSPLAIRRRSRSRTPRT-ARGKRSLTRSPPAIRR 2077
Score = 33.9 bits (76), Expect = 2.5
Identities = 41/114 (35%), Positives = 53/114 (46%)
Frame = -1
Query: 570 RRFSASQAAP*SIRLISVPGTRVRWPRLRSRGRMQRIAS*E*RS*TSGRTPLASRSPRAG 391
RR S S+ +P S R +R R RSR R ++ RS T T SRS R
Sbjct: 1885 RRRSRSRTSPVSRRRSR---SRTSVTRRRSRSRASPVSRRRSRSRTPPVTRRRSRS-RTP 1940
Query: 390 SGPASTRWDQPPPSSRLATSWTRTPSVTRTRLRR*KAS*ATPRSRCRSRWLLTP 229
+ +R PP + R + S RTP VTR R R S +P +R RSR +P
Sbjct: 1941 TTRRRSRSRTPPVTRRRSRS--RTPPVTRRRSR----SRTSPITRRRSRSRTSP 1988
Score = 33.1 bits (74), Expect = 4.2
Identities = 51/181 (28%), Positives = 67/181 (37%), Gaps = 30/181 (16%)
Frame = -1
Query: 681 RWQVTHHSS*TTMTKRGTRTLSSSTPGSTMELLATGCRRFSASQAAP*SIRLISVPGTRV 502
R Q + S T +RG S +P + RR S+ P S + +
Sbjct: 1800 RRQRSRSRSRVTRRRRGGSGYHSRSPARQESSRTSSRRRRGRSRTPPTSRKRSRSRTSPA 1859
Query: 501 RWPRLRSRG----------RMQRIAS*E*RS*TS--------GRTPLASRSPRAGSGPAS 376
W R RSR R I+ RS TS RT + R R+ + P S
Sbjct: 1860 PWKRSRSRASPATHRRSRSRTPLISRRRSRSRTSPVSRRRSRSRTSVTRRRSRSRASPVS 1919
Query: 375 TRWDQ---PPPSSRLATSWT---------RTPSVTRTRLRR*KAS*ATPRSRCRSRWLLT 232
R + PP + R + S T RTP VTR R R S P +R RSR +
Sbjct: 1920 RRRSRSRTPPVTRRRSRSRTPTTRRRSRSRTPPVTRRRSR----SRTPPVTRRRSRSRTS 1975
Query: 231 P 229
P
Sbjct: 1976 P 1976
>sptr|Q8NGC0|Q8NGC0 Seven transmembrane helix receptor.
Length = 311
Score = 35.8 bits (81), Expect = 0.65
Identities = 26/73 (35%), Positives = 37/73 (50%)
Frame = +3
Query: 195 LGSASARSKAVKE*GASVTCICSAALLSWLFTGATLFGLLKEFSSMTLPDVRTVAVDPSA 374
+ SA R KA + +T IC LF G TLF L+ SS +L RTVAV +
Sbjct: 228 MSSAQGRFKAFSTCASHLTAIC-------LFFGTTLFMYLRPRSSYSLTQDRTVAVIYTV 280
Query: 375 LTLAPSPLVETVR 413
+ +PL+ ++R
Sbjct: 281 VIPVLNPLMYSLR 293
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 587,644,043
Number of Sequences: 1272877
Number of extensions: 12502858
Number of successful extensions: 41991
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 39487
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 41844
length of database: 406,886,216
effective HSP length: 120
effective length of database: 254,140,976
effective search space used: 35325595664
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

