BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
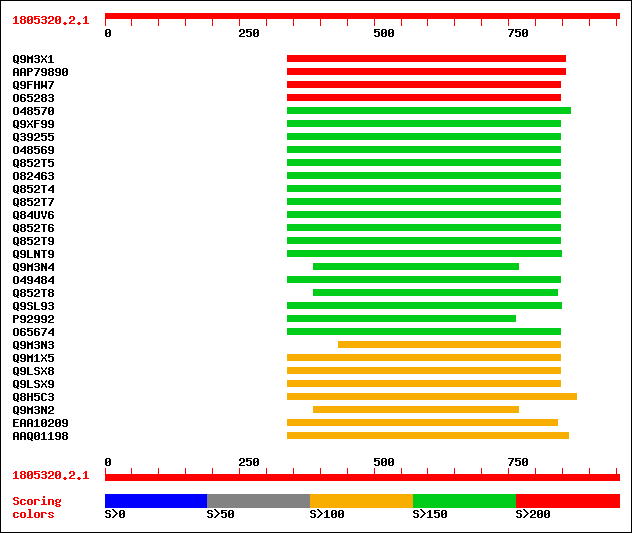
Query= 1805320.2.1
(955 letters)
Database: /db/trembl-ebi/tmp/swall
1,272,877 sequences; 406,886,216 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q9M3X1|Q9M3X1 Putative kinetochore protein. 222 5e-57
sptrnew|AAP79890|AAP79890 SKP1/ASK1-like protein. 219 4e-56
sptr|Q9FHW7|Q9FHW7 UIP2 (At5g42190). 205 9e-52
sptr|O65283|O65283 UIP2 (SKP1 homolog) (Fragment). 205 9e-52
sptr|O48570|O48570 Fimbriata-associated protein (Fragment). 194 1e-48
sptr|Q9XF99|Q9XF99 Skp1 (Fragment). 193 3e-48
sptr|Q39255|Q39255 SKP1P kinetochore protein homolog (T4O12.17) ... 193 3e-48
sptr|O48569|O48569 Fimbriata-associated protein (Fragment). 192 6e-48
sptr|Q852T5|Q852T5 Kinetochore protein. 191 1e-47
sptr|O82463|O82463 SKP1-like protein. 189 5e-47
sptr|Q852T4|Q852T4 Kinetochore protein. 189 5e-47
sptr|Q852T7|Q852T7 Kinetochore protein. 189 6e-47
sptr|Q84UV6|Q84UV6 SKP1. 188 8e-47
sptr|Q852T6|Q852T6 Kinetochore protein. 186 4e-46
sptr|Q852T9|Q852T9 Kinetochore protein. 183 4e-45
sptr|Q9LNT9|Q9LNT9 T20H2.8 protein (At1g20140/T20H2_8). 180 3e-44
sptr|Q9M3N4|Q9M3N4 Putative kinetochore protein (Fragment). 172 5e-42
sptr|O49484|O49484 Kinetochore (SKP1P) - like protein (SKP1P) (K... 171 1e-41
sptr|Q852T8|Q852T8 Kinetochore protein (Fragment). 171 2e-41
sptr|Q9SL93|Q9SL93 Putative kinetechore (Skp1p-like) protein (At... 170 3e-41
sptr|P92992|P92992 Homologue to SKP1. 169 7e-41
sptr|O65674|O65674 SKP1P - like protein (SKP1P-like protein). 167 2e-40
sptr|Q9M3N3|Q9M3N3 Putative kinetochore protein (Fragment). 149 7e-35
sptr|Q9M1X5|Q9M1X5 Skp1-like protein. 147 2e-34
sptr|Q9LSX8|Q9LSX8 Kinetechore (Skp1p-like) protein-like. 147 2e-34
sptr|Q9LSX9|Q9LSX9 Kinetechore (Skp1p-like) protein-like. 145 6e-34
sptr|Q8H5C3|Q8H5C3 Putative Skp1. 145 8e-34
sptr|Q9M3N2|Q9M3N2 Putative kinetochore protein (Fragment). 144 1e-33
sptrnew|EAA10209|EAA10209 AgCP15297 (Fragment). 144 2e-33
sptrnew|AAQ01198|AAQ01198 SKP1. 143 4e-33
>sptr|Q9M3X1|Q9M3X1 Putative kinetochore protein.
Length = 175
Score = 222 bits (566), Expect = 5e-57
Identities = 120/172 (69%), Positives = 121/172 (70%)
Frame = -2
Query: 855 GEKKMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCX 676
GEKKMITLKSSDG ESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYC
Sbjct: 8 GEKKMITLKSSDGEEFEVEEAVAMESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCN 67
Query: 675 XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYL 496
EDLKNWDA+FVKVDQATLFDLILAANYL
Sbjct: 68 KHVQAKPADAGASSDTASAAXGAPAAPA----EDLKNWDAEFVKVDQATLFDLILAANYL 123
Query: 495 NIKGLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
NIKGLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTP ENQWAFE
Sbjct: 124 NIKGLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPEEEEEIRRENQWAFE 175
>sptrnew|AAP79890|AAP79890 SKP1/ASK1-like protein.
Length = 175
Score = 219 bits (558), Expect = 4e-56
Identities = 118/172 (68%), Positives = 119/172 (69%)
Frame = -2
Query: 855 GEKKMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCX 676
GEKKMITLKSSDG ESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYC
Sbjct: 8 GEKKMITLKSSDGEEFEVEEAVAMESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCN 67
Query: 675 XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYL 496
EDLKNWDA+FVKVDQATLFDLILAANYL
Sbjct: 68 KHVQAKPADGAAAGAGAGASDAAPAAPA----EDLKNWDAEFVKVDQATLFDLILAANYL 123
Query: 495 NIKGLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
NIKGL DLTCQTVADMIKGKTPEEIRKTFNIKNDFTP ENQWAFE
Sbjct: 124 NIKGLPDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPEEEEEIRRENQWAFE 175
>sptr|Q9FHW7|Q9FHW7 UIP2 (At5g42190).
Length = 171
Score = 205 bits (521), Expect = 9e-52
Identities = 107/169 (63%), Positives = 112/169 (66%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
+ ITLKSSDG ESQTI+HMIEDDC DNGIPLPNV SKILSKVIEYC
Sbjct: 5 RKITLKSSDGENFEIDEAVALESQTIKHMIEDDCTDNGIPLPNVTSKILSKVIEYCKRHV 64
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
EDLK WD++F+KVDQ TLFDLILAANYLNIK
Sbjct: 65 EAAEKSETTADAAAATTTTTVASGSSD--EDLKTWDSEFIKVDQGTLFDLILAANYLNIK 122
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTP ENQWAFE
Sbjct: 123 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPEEEEEVRRENQWAFE 171
>sptr|O65283|O65283 UIP2 (SKP1 homolog) (Fragment).
Length = 172
Score = 205 bits (521), Expect = 9e-52
Identities = 107/169 (63%), Positives = 112/169 (66%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
+ ITLKSSDG ESQTI+HMIEDDC DNGIPLPNV SKILSKVIEYC
Sbjct: 6 RKITLKSSDGENFEIDEAVALESQTIKHMIEDDCTDNGIPLPNVTSKILSKVIEYCKRHV 65
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
EDLK WD++F+KVDQ TLFDLILAANYLNIK
Sbjct: 66 EAAEKSETTADAAAATTTTTVASGSSD--EDLKTWDSEFIKVDQGTLFDLILAANYLNIK 123
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTP ENQWAFE
Sbjct: 124 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPEEEEEVRRENQWAFE 172
>sptr|O48570|O48570 Fimbriata-associated protein (Fragment).
Length = 165
Score = 194 bits (494), Expect = 1e-48
Identities = 106/175 (60%), Positives = 113/175 (64%)
Frame = -2
Query: 864 AAEGEKKMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIE 685
+++ K ITL+SSDG SQTI+HMIEDDCADN IPLPNV KILSKVIE
Sbjct: 2 SSDNASKKITLRSSDGEVFEVDEAIALLSQTIKHMIEDDCADNVIPLPNVTGKILSKVIE 61
Query: 684 YCXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAA 505
YC +DLK +DADFVKVDQATLFDLILAA
Sbjct: 62 YCKRHVDADAAKSEEKVAAAAAGD-----------DDLKAFDADFVKVDQATLFDLILAA 110
Query: 504 NYLNIKGLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
NYLNIK LLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTP ENQWAFE
Sbjct: 111 NYLNIKTLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPEEEEEVRRENQWAFE 165
>sptr|Q9XF99|Q9XF99 Skp1 (Fragment).
Length = 153
Score = 193 bits (491), Expect = 3e-48
Identities = 104/169 (61%), Positives = 114/169 (67%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
+ ITLKSSDG ESQTI+HMIEDDCAD+GIPLPNV SKIL+KVIEYC
Sbjct: 3 RKITLKSSDGETFEVDEAVALESQTIKHMIEDDCADSGIPLPNVTSKILAKVIEYCKKHV 62
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
++LK+WD++FVKVDQATLFDLILAANYLNIK
Sbjct: 63 DAAAAEDKPNE------------------DELKSWDSEFVKVDQATLFDLILAANYLNIK 104
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
LLDLTCQTVADMIKGKTPEEIRKTFNIKNDF+P ENQWAFE
Sbjct: 105 SLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFSPEEEEEVRRENQWAFE 153
>sptr|Q39255|Q39255 SKP1P kinetochore protein homolog (T4O12.17)
(Putative SKP1/ASK1 protein At1).
Length = 160
Score = 193 bits (490), Expect = 3e-48
Identities = 102/169 (60%), Positives = 109/169 (64%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
K I LKSSDG ESQTI HM+EDDC DNG+PLPNV SKIL+KVIEYC
Sbjct: 4 KKIVLKSSDGESFEVEEAVALESQTIAHMVEDDCVDNGVPLPNVTSKILAKVIEYCKRHV 63
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
+DLK WDADF+K+DQATLF+LILAANYLNIK
Sbjct: 64 EAAASKAEAVEGAATSD------------DDLKAWDADFMKIDQATLFELILAANYLNIK 111
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
LLDLTCQTVADMIKGKTPEEIR TFNIKNDFTP ENQWAFE
Sbjct: 112 NLLDLTCQTVADMIKGKTPEEIRTTFNIKNDFTPEEEEEVRRENQWAFE 160
>sptr|O48569|O48569 Fimbriata-associated protein (Fragment).
Length = 161
Score = 192 bits (488), Expect = 6e-48
Identities = 106/169 (62%), Positives = 111/169 (65%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
K ITL+SSDG ESQTI+HMIEDDCADN IPLPNV KILSKVIEYC
Sbjct: 4 KKITLRSSDGEVFEVEESLALESQTIKHMIEDDCADNVIPLPNVTGKILSKVIEYCKRHV 63
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
++LK +DADFVKVDQATLFDLILAANYLNIK
Sbjct: 64 DAAAAKADDKLASTGTSD-----------DELKAFDADFVKVDQATLFDLILAANYLNIK 112
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
LLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTP ENQWAFE
Sbjct: 113 TLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPEEEEEVRRENQWAFE 161
>sptr|Q852T5|Q852T5 Kinetochore protein.
Length = 160
Score = 191 bits (486), Expect = 1e-47
Identities = 102/169 (60%), Positives = 109/169 (64%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
K I LKSSDG ESQTI HM+EDDC DNGIPLPNV SKIL+KVIEYC
Sbjct: 4 KKIVLKSSDGESFEVDEAVALESQTIAHMVEDDCVDNGIPLPNVTSKILAKVIEYCKKHV 63
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
+DLK WDA+F+K+DQATLF+LILAANYLNIK
Sbjct: 64 DAAASKTEAVDGGASSD------------DDLKAWDAEFMKIDQATLFELILAANYLNIK 111
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
LLDLTCQTVADMIKGKTPEEIR TFNIKNDFTP ENQWAFE
Sbjct: 112 NLLDLTCQTVADMIKGKTPEEIRTTFNIKNDFTPEEEEEVRRENQWAFE 160
>sptr|O82463|O82463 SKP1-like protein.
Length = 153
Score = 189 bits (480), Expect = 5e-47
Identities = 104/169 (61%), Positives = 110/169 (65%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
KMI L+SSDG ESQTI+HMIEDDCAD IPLPNV SKIL+KVIEYC
Sbjct: 2 KMIVLRSSDGETFEVEESVALESQTIKHMIEDDCADTSIPLPNVTSKILAKVIEYCKRHV 61
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
+DLK +DADFVKVDQ+TLFDLILAANYLNIK
Sbjct: 62 DAASKTEDKAVE-----------------DDLKAFDADFVKVDQSTLFDLILAANYLNIK 104
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
LLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTP EN WAFE
Sbjct: 105 SLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPEEEEEVRRENAWAFE 153
>sptr|Q852T4|Q852T4 Kinetochore protein.
Length = 160
Score = 189 bits (480), Expect = 5e-47
Identities = 100/169 (59%), Positives = 109/169 (64%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
K I LKSSDG ESQTI HM+EDDC DNG+PLPNV SKIL+KVIEYC
Sbjct: 4 KKIVLKSSDGESFEVDEAVALESQTIAHMVEDDCVDNGVPLPNVTSKILAKVIEYCKKHV 63
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
+DLK WDA+F+K+DQATLF+LILAANYLNIK
Sbjct: 64 DAAASKSEAVDGGGSSD------------DDLKAWDAEFMKIDQATLFELILAANYLNIK 111
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
LLDLTCQTVADMIKGKTPEEIR TFNIKNDF+P ENQWAFE
Sbjct: 112 NLLDLTCQTVADMIKGKTPEEIRTTFNIKNDFSPEEEEEVRRENQWAFE 160
>sptr|Q852T7|Q852T7 Kinetochore protein.
Length = 160
Score = 189 bits (479), Expect = 6e-47
Identities = 100/169 (59%), Positives = 109/169 (64%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
K I LKSSDG ESQTI HM+EDDC DNG+PLPNV SKIL+KVIEYC
Sbjct: 4 KKIVLKSSDGESFEVDEAVALESQTIAHMVEDDCVDNGVPLPNVTSKILAKVIEYCKKHV 63
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
+DLK WDA+F+K+DQATLF+LILAANYLNIK
Sbjct: 64 DAVASKSEAVDGGGSSD------------DDLKAWDAEFMKIDQATLFELILAANYLNIK 111
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
LLDLTCQTVADMIKGKTPEEIR TFNIKNDF+P ENQWAFE
Sbjct: 112 NLLDLTCQTVADMIKGKTPEEIRTTFNIKNDFSPEEEEEVRRENQWAFE 160
>sptr|Q84UV6|Q84UV6 SKP1.
Length = 153
Score = 188 bits (478), Expect = 8e-47
Identities = 104/169 (61%), Positives = 110/169 (65%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
KMI L+SSDG ESQTI+HMIEDDCAD IPLPNV SKIL+KVIEYC
Sbjct: 2 KMIVLRSSDGETFEVEESVALESQTIKHMIEDDCADTSIPLPNVTSKILAKVIEYCKRHV 61
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
+DLK +DADFVKVDQ+TLFDLILAANYLNIK
Sbjct: 62 DAASKTEDKAVE-----------------DDLKAFDADFVKVDQSTLFDLILAANYLNIK 104
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
LLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTP EN WAFE
Sbjct: 105 RLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPEEEEEVRRENAWAFE 153
>sptr|Q852T6|Q852T6 Kinetochore protein.
Length = 161
Score = 186 bits (472), Expect = 4e-46
Identities = 98/169 (57%), Positives = 108/169 (63%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
K I LKSSDG ESQT+ HM+EDDC +NGIPLPNV SKIL+KVIEYC
Sbjct: 4 KKIVLKSSDGESFEVDEAVARESQTLAHMVEDDCIENGIPLPNVTSKILAKVIEYCKKHV 63
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
+DLK WD +F+K+DQATLF+LILAANYLNIK
Sbjct: 64 DAAAAKTEGAVDGAASSD-----------DDLKAWDTEFMKIDQATLFELILAANYLNIK 112
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
LLDLTCQTVADMIKGKTPEEIR TFNIKNDF+P ENQWAFE
Sbjct: 113 NLLDLTCQTVADMIKGKTPEEIRTTFNIKNDFSPEEEEEVRRENQWAFE 161
>sptr|Q852T9|Q852T9 Kinetochore protein.
Length = 161
Score = 183 bits (464), Expect = 4e-45
Identities = 97/169 (57%), Positives = 106/169 (62%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
K I LKSSD ESQT+ HM+EDDC DNGIPLPNV KIL+KVIEYC
Sbjct: 4 KKIVLKSSDDESFEVDEAVARESQTLAHMVEDDCTDNGIPLPNVTGKILAKVIEYCKKHV 63
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
EDLK WDA+F+ +DQATLF+LILAANYLNIK
Sbjct: 64 DAAAAKTEATADGGAPSD-----------EDLKAWDAEFMNIDQATLFELILAANYLNIK 112
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
LLDLTCQTVADMIKGKTP+EIR TFNIKNDF+P ENQWAFE
Sbjct: 113 NLLDLTCQTVADMIKGKTPDEIRTTFNIKNDFSPEEEEEVRRENQWAFE 161
>sptr|Q9LNT9|Q9LNT9 T20H2.8 protein (At1g20140/T20H2_8).
Length = 163
Score = 180 bits (456), Expect = 3e-44
Identities = 94/170 (55%), Positives = 108/170 (63%)
Frame = -2
Query: 849 KKMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXX 670
KKMI LKSSDG +SQTI+HMIEDDCADNGIPLPNV IL+KVIEYC
Sbjct: 5 KKMIILKSSDGESFEIEEAVAVKSQTIKHMIEDDCADNGIPLPNVTGAILAKVIEYCKKH 64
Query: 669 XXXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNI 490
++LKNWD++FVKVDQ TLFDLILAANYLNI
Sbjct: 65 VEAAAEAGGDKDFYGSAE-----------NDELKNWDSEFVKVDQPTLFDLILAANYLNI 113
Query: 489 KGLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
GLLDLTC+ VAD ++GKTPE++R FNIKND+TP EN+WAFE
Sbjct: 114 GGLLDLTCKAVADQMRGKTPEQMRAHFNIKNDYTPEEEAEVRNENKWAFE 163
>sptr|Q9M3N4|Q9M3N4 Putative kinetochore protein (Fragment).
Length = 117
Score = 172 bits (437), Expect = 5e-42
Identities = 89/127 (70%), Positives = 94/127 (74%)
Frame = -2
Query: 768 RHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX 589
+HMIEDDCADNGIPLPNVNSKILSKVIEYC
Sbjct: 1 KHMIEDDCADNGIPLPNVNSKILSKVIEYCNKHVQAKPADAAGPGASDALPPA------- 53
Query: 588 XSGEDLKNWDADFVKVDQATLFDLILAANYLNIKGLLDLTCQTVADMIKGKTPEEIRKTF 409
+DLKNWDA+FVKVDQATLFDLILAAN+LNIKGLLDLTCQTVADMIKGKTPEEIRKTF
Sbjct: 54 ---DDLKNWDAEFVKVDQATLFDLILAANFLNIKGLLDLTCQTVADMIKGKTPEEIRKTF 110
Query: 408 NIKNDFT 388
NIKNDF+
Sbjct: 111 NIKNDFS 117
>sptr|O49484|O49484 Kinetochore (SKP1P) - like protein (SKP1P)
(Kinetochore (SKP1P)-like protein).
Length = 152
Score = 171 bits (433), Expect = 1e-41
Identities = 93/169 (55%), Positives = 103/169 (60%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
KMI L SSDG +SQTI HM+EDDC +GIPL NV SKIL KVIEYC
Sbjct: 4 KMIVLMSSDGQSFEVEEAVAIQSQTIAHMVEDDCVADGIPLANVESKILVKVIEYCKKHH 63
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
EDL NWD F+ ++Q+T+F+LILAANYLNIK
Sbjct: 64 VDEANPISE--------------------EDLNNWDEKFMDLEQSTIFELILAANYLNIK 103
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
LLDLTCQTVADMIKGKTPEEIR TFNI+NDFTP ENQWAFE
Sbjct: 104 SLLDLTCQTVADMIKGKTPEEIRSTFNIENDFTPEEEEAVRKENQWAFE 152
>sptr|Q852T8|Q852T8 Kinetochore protein (Fragment).
Length = 145
Score = 171 bits (432), Expect = 2e-41
Identities = 91/151 (60%), Positives = 99/151 (65%)
Frame = -2
Query: 840 ITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXXXX 661
+ LKSSDG ESQTI HM+EDD DNGIPLPNV SKIL+KVIEYC
Sbjct: 1 MVLKSSDGESFEVDEAVALESQTIAHMVEDDGVDNGIPLPNVTSKILAKVIEYCKKHVDA 60
Query: 660 XXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIKGL 481
+DLK WDA+F+K+DQATLF+LILAANYLNIK L
Sbjct: 61 AASKTEAVDGGASSD------------DDLKAWDAEFMKIDQATLFELILAANYLNIKNL 108
Query: 480 LDLTCQTVADMIKGKTPEEIRKTFNIKNDFT 388
LDLTCQTVADMIKGKTPEEIR TFNIKNDFT
Sbjct: 109 LDLTCQTVADMIKGKTPEEIRTTFNIKNDFT 139
>sptr|Q9SL93|Q9SL93 Putative kinetechore (Skp1p-like) protein
(At2g25700/F3N11.15).
Length = 163
Score = 170 bits (430), Expect = 3e-41
Identities = 90/170 (52%), Positives = 102/170 (60%)
Frame = -2
Query: 849 KKMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXX 670
KKMI LKSSDG ESQTI+HMIEDDC DNGIPLPNV IL+KVIEYC
Sbjct: 5 KKMIILKSSDGESFEVEEAVAVESQTIKHMIEDDCVDNGIPLPNVTGAILAKVIEYCKKH 64
Query: 669 XXXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNI 490
+LK WD DFVKVD TLFDL+ AANYLNI
Sbjct: 65 VEAAAEAGGDKDFYGSTE-----------NHELKTWDNDFVKVDHPTLFDLLRAANYLNI 113
Query: 489 KGLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
GLLDLTC+ VAD ++GKTP ++R+ FNIKND+TP EN+WAFE
Sbjct: 114 SGLLDLTCKAVADQMRGKTPAQMREHFNIKNDYTPEEEAEVRNENRWAFE 163
>sptr|P92992|P92992 Homologue to SKP1.
Length = 129
Score = 169 bits (427), Expect = 7e-41
Identities = 87/141 (61%), Positives = 94/141 (66%)
Frame = -2
Query: 762 MIEDDCADNGIPLPNVNSKILSKVIEYCXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXS 583
M+EDDC DNG+ LPNV SKIL+KVIEYC
Sbjct: 1 MVEDDCVDNGVLLPNVTSKILAKVIEYCKRHVEAAASKAEAVEGAATSD----------- 49
Query: 582 GEDLKNWDADFVKVDQATLFDLILAANYLNIKGLLDLTCQTVADMIKGKTPEEIRKTFNI 403
+DLK WDADF+K+DQATLF+LILAANYLNIK LLDLTCQTVADMIKGKTPEEIR TFNI
Sbjct: 50 -DDLKAWDADFMKIDQATLFELILAANYLNIKNLLDLTCQTVADMIKGKTPEEIRTTFNI 108
Query: 402 KNDFTPXXXXXXXXENQWAFE 340
KNDFTP ENQWAFE
Sbjct: 109 KNDFTPEEEEEVRRENQWAFE 129
>sptr|O65674|O65674 SKP1P - like protein (SKP1P-like protein).
Length = 152
Score = 167 bits (423), Expect = 2e-40
Identities = 91/169 (53%), Positives = 101/169 (59%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
KMI L SSDG +SQTI HM+EDDC +GIPL NV SKIL KVIEYC
Sbjct: 4 KMIVLMSSDGQSFEVEEAVAIQSQTIAHMVEDDCVADGIPLANVESKILVKVIEYCKKYH 63
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
EDL WD F+ ++Q+T+F+LILAANYLNIK
Sbjct: 64 VDEANPISE--------------------EDLNKWDEKFMDLEQSTIFELILAANYLNIK 103
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
L DLTCQTVADMIKGKTPEEIR TFNI+NDFTP ENQWAFE
Sbjct: 104 SLFDLTCQTVADMIKGKTPEEIRSTFNIENDFTPEEEEAVRKENQWAFE 152
>sptr|Q9M3N3|Q9M3N3 Putative kinetochore protein (Fragment).
Length = 124
Score = 149 bits (375), Expect = 7e-35
Identities = 83/138 (60%), Positives = 89/138 (64%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
K ITLKSSDG ESQ I+HMIEDDCAD+GIPLPNV SKIL+KVIE+C
Sbjct: 5 KKITLKSSDGEAFEVDEAVALESQAIKHMIEDDCADSGIPLPNVTSKILAKVIEFCKKHV 64
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
++LK WDADFVKVDQ TLFDLILAANYLNIK
Sbjct: 65 XAAASDDKPTE------------------DELKAWDADFVKVDQVTLFDLILAANYLNIK 106
Query: 486 GLLDLTCQTVADMIKGKT 433
LLDLTCQTVADMIKGKT
Sbjct: 107 NLLDLTCQTVADMIKGKT 124
>sptr|Q9M1X5|Q9M1X5 Skp1-like protein.
Length = 154
Score = 147 bits (372), Expect = 2e-34
Identities = 83/171 (48%), Positives = 96/171 (56%), Gaps = 2/171 (1%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
KM+ L SSDG +SQTI HMIEDDC NG+P+ NV ILSKVIEYC
Sbjct: 3 KMVMLLSSDGESFQVEEAVAVQSQTIAHMIEDDCVANGVPIANVTGVILSKVIEYCKKHV 62
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKV--DQATLFDLILAANYLN 493
++LK WDA+F+K +TLFD++LAANYLN
Sbjct: 63 VSDSPTEESK-------------------DELKKWDAEFMKALEQSSTLFDVMLAANYLN 103
Query: 492 IKGLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
IK LLDL CQTVADMI GK P+EIR I+NDFTP ENQWAFE
Sbjct: 104 IKDLLDLGCQTVADMITGKKPDEIRALLGIENDFTPEEEEEIRKENQWAFE 154
>sptr|Q9LSX8|Q9LSX8 Kinetechore (Skp1p-like) protein-like.
Length = 152
Score = 147 bits (372), Expect = 2e-34
Identities = 83/169 (49%), Positives = 92/169 (54%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEYCXXXX 667
K I LKSSDG + QTI HM EDDC DNGIPLP V KIL VIEYC
Sbjct: 4 KKIILKSSDGHSFEVEEEAACQCQTIAHMSEDDCTDNGIPLPEVTGKILEMVIEYCNKHH 63
Query: 666 XXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNIK 487
EDLK WD +F++ Q+T+FDLI+AANYLNIK
Sbjct: 64 VDAANPCSD--------------------EDLKKWDKEFMEKYQSTIFDLIMAANYLNIK 103
Query: 486 GLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
LLDL CQTVADMIK T E RK FNI+ND+T ENQW FE
Sbjct: 104 SLLDLACQTVADMIKDNTVEHTRKFFNIENDYTHEEEEAVRRENQWGFE 152
>sptr|Q9LSX9|Q9LSX9 Kinetechore (Skp1p-like) protein-like.
Length = 153
Score = 145 bits (367), Expect = 6e-34
Identities = 81/170 (47%), Positives = 96/170 (56%), Gaps = 1/170 (0%)
Frame = -2
Query: 846 KMITLKSSDGXXXXXXXXXXXESQTI-RHMIEDDCADNGIPLPNVNSKILSKVIEYCXXX 670
K I LKSSDG + Q I HM E+DC DNGIPLPNV KIL+ VIEYC
Sbjct: 4 KKIILKSSDGHSFEVEEEAARQCQIIIAHMSENDCTDNGIPLPNVTGKILAMVIEYCNKH 63
Query: 669 XXXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLNI 490
+DLK WD +F++ D +T+FDLI AANYLNI
Sbjct: 64 HVDAANPCSD--------------------DDLKKWDKEFMEKDTSTIFDLIKAANYLNI 103
Query: 489 KGLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
K L DL CQTVA++IKG TPE+IR+ FNI+ND TP EN+WAFE
Sbjct: 104 KSLFDLACQTVAEIIKGNTPEQIREFFNIENDLTPEEEAAIRRENKWAFE 153
>sptr|Q8H5C3|Q8H5C3 Putative Skp1.
Length = 164
Score = 145 bits (366), Expect = 8e-34
Identities = 79/179 (44%), Positives = 100/179 (55%)
Frame = -2
Query: 876 IESMAAEGEKKMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILS 697
+ AA I L SSDG SQ + +MIEDDC NG+PLPNV SK+L+
Sbjct: 8 VADAAAGSGSDTILLISSDGEHFNVPSAAASLSQLVSNMIEDDCTTNGVPLPNVASKVLA 67
Query: 696 KVIEYCXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDL 517
KVIEYC +DLK++DA+F+ VD+ L+DL
Sbjct: 68 KVIEYCIKHAAAGEEEE----------------------KDLKSFDAEFIDVDKNMLYDL 105
Query: 516 ILAANYLNIKGLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
+LA+N++NIK LLDL CQ A++IKGK+PE+IRK F IKNDFTP EN WAFE
Sbjct: 106 LLASNFMNIKSLLDLCCQHTANLIKGKSPEQIRKEFGIKNDFTPEEEEEIRKENTWAFE 164
>sptr|Q9M3N2|Q9M3N2 Putative kinetochore protein (Fragment).
Length = 113
Score = 144 bits (364), Expect = 1e-33
Identities = 76/128 (59%), Positives = 89/128 (69%), Gaps = 1/128 (0%)
Frame = -2
Query: 768 RHMIEDDCADN-GIPLPNVNSKILSKVIEYCXXXXXXXXXXXXXXXXXXXXXXXXXXXXX 592
+HMIEDDCAD GIPLPNV S+IL+KVIEYC
Sbjct: 1 KHMIEDDCADETGIPLPNVTSRILAKVIEYCKKHVEAPKIDEYGMPVD------------ 48
Query: 591 XXSGEDLKNWDADFVKVDQATLFDLILAANYLNIKGLLDLTCQTVADMIKGKTPEEIRKT 412
G+D+K WDA+FVKVDQ TLFDLILAANYL+IK LLDLTC+TVA+M+ G+TP+EIR+T
Sbjct: 49 ---GKDMKKWDAEFVKVDQDTLFDLILAANYLDIKSLLDLTCKTVANMMDGRTPDEIRRT 105
Query: 411 FNIKNDFT 388
FNIKNDFT
Sbjct: 106 FNIKNDFT 113
>sptrnew|EAA10209|EAA10209 AgCP15297 (Fragment).
Length = 177
Score = 144 bits (363), Expect = 2e-33
Identities = 83/171 (48%), Positives = 100/171 (58%), Gaps = 4/171 (2%)
Frame = -2
Query: 840 ITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNG----IPLPNVNSKILSKVIEYCXX 673
I L+SSDG S TI+ M+ED D G +PLPNVNS IL KV+++
Sbjct: 19 IKLQSSDGEIFDTDVQIAKCSGTIKTMLEDLGMDEGDDEVVPLPNVNSAILRKVLQWATF 78
Query: 672 XXXXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAANYLN 493
+D+ +WDADF+KVDQ TLF+LILAANYL+
Sbjct: 79 HKDDPIPVEDDDSKEKRT-------------DDISSWDADFLKVDQGTLFELILAANYLD 125
Query: 492 IKGLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
IKGLLD+TC+TVA+MIKGKTPEEIRKTFNIKNDFTP EN+W E
Sbjct: 126 IKGLLDVTCKTVANMIKGKTPEEIRKTFNIKNDFTPAEEEQVRKENEWCEE 176
>sptrnew|AAQ01198|AAQ01198 SKP1.
Length = 169
Score = 143 bits (360), Expect = 4e-33
Identities = 76/174 (43%), Positives = 100/174 (57%)
Frame = -2
Query: 861 AEGEKKMITLKSSDGXXXXXXXXXXXESQTIRHMIEDDCADNGIPLPNVNSKILSKVIEY 682
A+ +KMI L SSDG +S+T+ HMIEDDC DNG+PLPNV + +L+KV+EY
Sbjct: 5 ADNGEKMILLISSDGERFELSEAAASQSKTLSHMIEDDCTDNGVPLPNVTAVVLAKVVEY 64
Query: 681 CXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXSGEDLKNWDADFVKVDQATLFDLILAAN 502
E+LK++DA+FV VD+ +F+LILAAN
Sbjct: 65 FKKHAAVTPKPATEAVAADKAKRE----------EELKSFDAEFVDVDRTMVFELILAAN 114
Query: 501 YLNIKGLLDLTCQTVADMIKGKTPEEIRKTFNIKNDFTPXXXXXXXXENQWAFE 340
+LN + LLDLTCQ AD+IK + EE+R+ FNI NDFTP EN WAF+
Sbjct: 115 FLNAQDLLDLTCQHAADLIKDMSVEEVREVFNITNDFTPEEEAEVRKENAWAFD 168
Database: /db/trembl-ebi/tmp/swall
Posted date: Sep 26, 2003 8:29 PM
Number of letters in database: 406,886,216
Number of sequences in database: 1,272,877
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 574,519,441
Number of Sequences: 1272877
Number of extensions: 10270845
Number of successful extensions: 28396
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 27222
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 28341
length of database: 406,886,216
effective HSP length: 122
effective length of database: 251,595,222
effective search space used: 49061068290
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

