BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
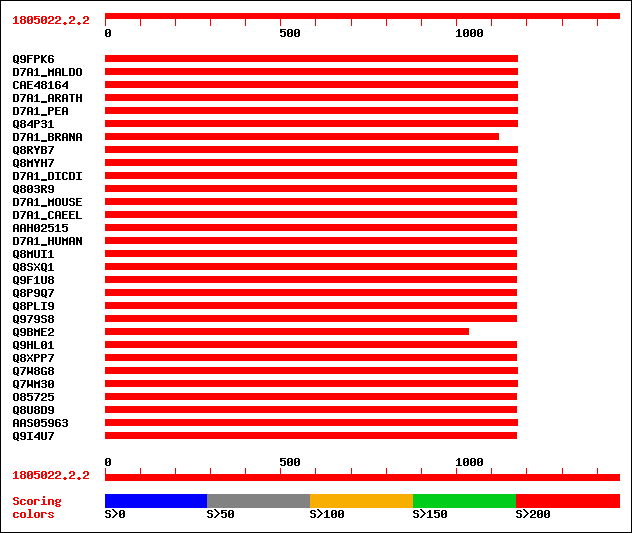
Query= 1805022.2.2
(1462 letters)
Database: /db/uniprot/tmp/swall
1,395,590 sequences; 442,889,342 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
sptr|Q9FPK6|Q9FPK6 Aldehyde dehydrogenase. 710 0.0
sw|Q9ZPB7|D7A1_MALDO Aldehyde dehydrogenase family 7 member A1 (... 639 0.0
sptrnew|CAE48164|CAE48164 Putative aldehyde dehydrogenase (EC 1.... 634 e-180
sw|Q9SYG7|D7A1_ARATH Aldehyde dehydrogenase family 7 member A1 (... 634 e-180
sw|P25795|D7A1_PEA Aldehyde dehydrogenase family 7 member A1 (EC... 610 e-173
sptr|Q84P31|Q84P31 Aldehyde dehydrogenase family 7 member A1 (EC... 604 e-172
sw|Q41247|D7A1_BRANA Aldehyde dehydrogenase family 7 member A1 (... 601 e-170
sptr|Q8RYB7|Q8RYB7 Aldehyde dehydrogenase ALDH2. 577 e-163
sptr|Q8MYH7|Q8MYH7 Similar to Arabidopsis thaliana (Mouse-ear cr... 475 e-133
sw|P83401|D7A1_DICDI Putative aldehyde dehydrogenase family 7 me... 475 e-133
sptr|Q803R9|Q803R9 Similar to aldehyde dehydrogenase 7 family, m... 473 e-132
sw|Q9DBF1|D7A1_MOUSE Aldehyde dehydrogenase family 7 member A1 (... 462 e-129
sw|P46562|D7A1_CAEEL Putative aldehyde dehydrogenase family 7 me... 453 e-126
sptrnew|AAH02515|AAH02515 Antiquitin (Fragment). 452 e-126
sw|P49419|D7A1_HUMAN Aldehyde dehydrogenase family 7 member A1 (... 452 e-126
sptr|Q8MUI1|Q8MUI1 Aldehyde dehydrogenase. 446 e-124
sptr|Q8SXQ1|Q8SXQ1 GH05218p (CG9629 protein). 404 e-111
sptr|Q9F1U8|Q9F1U8 Piperideine-6-carboxylate dehydrogenase. 390 e-107
sptr|Q8P9Q7|Q8P9Q7 Aldehyde dehydrogenase. 383 e-105
sptr|Q8PLI9|Q8PLI9 Aldehyde dehydrogenase. 380 e-104
sptr|Q979S8|Q979S8 Aldehyde dehydrogenase. 380 e-104
sptr|Q9BME2|Q9BME2 Turgor-like protein (Fragment). 377 e-103
sptr|Q9HL01|Q9HL01 Piperideine-6-carboxilic acid dehydrogenase r... 376 e-103
sptr|Q8XPP7|Q8XPP7 Putative transmembrane aldehyde dehydrogenase... 366 e-100
sptr|Q7W8G8|Q7W8G8 Probable aldehyde dehydrogenase. 362 6e-99
sptr|Q7WM30|Q7WM30 Probable aldehyde dehydrogenase. 362 6e-99
sptr|O85725|O85725 Semialdehyde dehydrogenase Pcd. 360 2e-98
sptr|Q8U8D9|Q8U8D9 Aldehyde dehydrogenase. 356 5e-97
sptrnew|AAS05963|AAS05963 AldB. 353 3e-96
sptr|Q9I4U7|Q9I4U7 Probable aldehyde dehydrogenase. 352 1e-95
>sptr|Q9FPK6|Q9FPK6 Aldehyde dehydrogenase.
Length = 509
Score = 710 bits (1832), Expect = 0.0
Identities = 348/391 (89%), Positives = 367/391 (93%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+EIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI
Sbjct: 119 QEIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 178
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
ALVCGNCVVWKGAPTTPLITIAMTKIVASVLE+NNLPG+IFT+FCGG +IGQAI+LD RI
Sbjct: 179 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLERNNLPGSIFTAFCGGADIGQAISLDTRI 238
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PLVSFTGST+ GLMVQQQV+ARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG
Sbjct: 239 PLVSFTGSTKVGLMVQQQVNARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 298
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTTCRRL+LHE+IY+TFLDQLVEVYKQVRIGDPLE GTLLGPLHTPAS++ FLKGIQT
Sbjct: 299 QRCTTCRRLLLHESIYRTFLDQLVEVYKQVRIGDPLENGTLLGPLHTPASRDAFLKGIQT 358
Query: 722 IKSQGGKILFGGSAIESEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEAIEIN 901
I+SQGGKIL+GGSAIESEGNFVQPTIVEI+PSAPVV+EELFGPVLYVMK Q LKEA+EIN
Sbjct: 359 IRSQGGKILYGGSAIESEGNFVQPTIVEISPSAPVVREELFGPVLYVMKVQNLKEAVEIN 418
Query: 902 NSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXXXX 1081
NSVPQGLSSSIFTKRPDIIFKW+GPHGSDCGIVNVNIPTNGAEI
Sbjct: 419 NSVPQGLSSSIFTKRPDIIFKWIGPHGSDCGIVNVNIPTNGAEIGGAFGGEKATGGGREA 478
Query: 1082 XSDSWKQYMRRATCTINYGSELPLAQGINFG 1174
SDSWKQYMRRATCTINYGSELPLAQGINFG
Sbjct: 479 GSDSWKQYMRRATCTINYGSELPLAQGINFG 509
>sw|Q9ZPB7|D7A1_MALDO Aldehyde dehydrogenase family 7 member A1 (EC
1.2.1.3) (Antiquitin 1) (Matured fruit 60 kDa protein)
(MF-60).
Length = 507
Score = 639 bits (1649), Expect = 0.0
Identities = 309/391 (79%), Positives = 344/391 (87%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+I MCD+AVGLSRQLNGSIIPSERP+HMM EVWNPLG+VGVITAFNFPCAVLGWNACI
Sbjct: 117 QEVIYMCDFAVGLSRQLNGSIIPSERPDHMMFEVWNPLGIVGVITAFNFPCAVLGWNACI 176
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
ALVCGNCVVWKGAPTTPL+TIA+TK++A VLEKNNLP AIFT+FCGG EIG+AIA D RI
Sbjct: 177 ALVCGNCVVWKGAPTTPLVTIAVTKLIAEVLEKNNLPAAIFTAFCGGAEIGEAIAKDTRI 236
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PLVSFTGS++ G VQQ V+ RFGKCLLELSGNNA+IVMDDAD+ LAVRS+ FAAVGTAG
Sbjct: 237 PLVSFTGSSKVGAKVQQIVTERFGKCLLELSGNNALIVMDDADVGLAVRSIFFAAVGTAG 296
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTTCRRL LHE+IYQ LD+LV +Y QV+IGDPLE+GTL+GP+HT AS+ENF KGI T
Sbjct: 297 QRCTTCRRLYLHESIYQNVLDKLVGLYNQVKIGDPLEEGTLVGPVHTKASRENFEKGIST 356
Query: 722 IKSQGGKILFGGSAIESEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEAIEIN 901
IKSQGGKIL GGS IES+GNFVQPTIVEI +A VVKEELFGPVLYVMKF+TL+EAI +N
Sbjct: 357 IKSQGGKILTGGSVIESDGNFVQPTIVEIASNASVVKEELFGPVLYVMKFKTLEEAIALN 416
Query: 902 NSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXXXX 1081
NSVPQGLSSSIFT +P+ IFKW+GPHGSDCGIVNVNIPTNGAEI
Sbjct: 417 NSVPQGLSSSIFTSKPNTIFKWIGPHGSDCGIVNVNIPTNGAEIGGAFGGEKATGGGREA 476
Query: 1082 XSDSWKQYMRRATCTINYGSELPLAQGINFG 1174
SDSWKQYMRR+TCTINYG+ELPLAQGINFG
Sbjct: 477 GSDSWKQYMRRSTCTINYGTELPLAQGINFG 507
>sptrnew|CAE48164|CAE48164 Putative aldehyde dehydrogenase (EC
1.2.1.3).
Length = 508
Score = 634 bits (1635), Expect = e-180
Identities = 307/391 (78%), Positives = 342/391 (87%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+IDMCD+AVGLSRQLNGS+IPSERPNHMM+E+WNPLG+VGVITAFNFPCAVLGWNACI
Sbjct: 118 QEVIDMCDFAVGLSRQLNGSVIPSERPNHMMLEMWNPLGIVGVITAFNFPCAVLGWNACI 177
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
ALVCGNCVVWKGAPTTPLITIAMTK+VA VLEKNNLPGAIFT+ CGG EIG+AIA D RI
Sbjct: 178 ALVCGNCVVWKGAPTTPLITIAMTKLVAEVLEKNNLPGAIFTAMCGGAEIGEAIAKDTRI 237
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PLVSFTGS+R G MVQQ V+AR GK LLELSGNNAIIVMDDADIQLA RSVLFAAVGTAG
Sbjct: 238 PLVSFTGSSRVGSMVQQTVNARSGKTLLELSGNNAIIVMDDADIQLAARSVLFAAVGTAG 297
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTTCRRL+LHE++Y L+QL+ YKQV+IG+PLEKGTLLGPLHTP SK+NF KGI+
Sbjct: 298 QRCTTCRRLLLHESVYDKVLEQLLTSYKQVKIGNPLEKGTLLGPLHTPESKKNFEKGIEV 357
Query: 722 IKSQGGKILFGGSAIESEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEAIEIN 901
IKSQGGKIL GG A+E EGNFV+PTI+EI+ A VVKEELF PVLYV+KF++ EA+ IN
Sbjct: 358 IKSQGGKILTGGKAVEGEGNFVEPTIIEISADAAVVKEELFAPVLYVLKFKSFGEAVAIN 417
Query: 902 NSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXXXX 1081
NSVPQGLSSSIFT+ P+ IF+W+GP GSDCGIVNVNIPTNGAEI
Sbjct: 418 NSVPQGLSSSIFTRNPENIFRWIGPLGSDCGIVNVNIPTNGAEIGGAFGGEKATGGGREA 477
Query: 1082 XSDSWKQYMRRATCTINYGSELPLAQGINFG 1174
SDSWKQYMRR+TCTINYG+ELPLAQGINFG
Sbjct: 478 GSDSWKQYMRRSTCTINYGNELPLAQGINFG 508
>sw|Q9SYG7|D7A1_ARATH Aldehyde dehydrogenase family 7 member A1 (EC
1.2.1.3) (Antiquitin 1).
Length = 508
Score = 634 bits (1635), Expect = e-180
Identities = 307/391 (78%), Positives = 342/391 (87%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+IDMCD+AVGLSRQLNGS+IPSERPNHMM+E+WNPLG+VGVITAFNFPCAVLGWNACI
Sbjct: 118 QEVIDMCDFAVGLSRQLNGSVIPSERPNHMMLEMWNPLGIVGVITAFNFPCAVLGWNACI 177
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
ALVCGNCVVWKGAPTTPLITIAMTK+VA VLEKNNLPGAIFT+ CGG EIG+AIA D RI
Sbjct: 178 ALVCGNCVVWKGAPTTPLITIAMTKLVAEVLEKNNLPGAIFTAMCGGAEIGEAIAKDTRI 237
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PLVSFTGS+R G MVQQ V+AR GK LLELSGNNAIIVMDDADIQLA RSVLFAAVGTAG
Sbjct: 238 PLVSFTGSSRVGSMVQQTVNARSGKTLLELSGNNAIIVMDDADIQLAARSVLFAAVGTAG 297
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTTCRRL+LHE++Y L+QL+ YKQV+IG+PLEKGTLLGPLHTP SK+NF KGI+
Sbjct: 298 QRCTTCRRLLLHESVYDKVLEQLLTSYKQVKIGNPLEKGTLLGPLHTPESKKNFEKGIEV 357
Query: 722 IKSQGGKILFGGSAIESEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEAIEIN 901
IKSQGGKIL GG A+E EGNFV+PTI+EI+ A VVKEELF PVLYV+KF++ EA+ IN
Sbjct: 358 IKSQGGKILTGGKAVEGEGNFVEPTIIEISADAAVVKEELFAPVLYVLKFKSFGEAVAIN 417
Query: 902 NSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXXXX 1081
NSVPQGLSSSIFT+ P+ IF+W+GP GSDCGIVNVNIPTNGAEI
Sbjct: 418 NSVPQGLSSSIFTRNPENIFRWIGPLGSDCGIVNVNIPTNGAEIGGAFGGEKATGGGREA 477
Query: 1082 XSDSWKQYMRRATCTINYGSELPLAQGINFG 1174
SDSWKQYMRR+TCTINYG+ELPLAQGINFG
Sbjct: 478 GSDSWKQYMRRSTCTINYGNELPLAQGINFG 508
>sw|P25795|D7A1_PEA Aldehyde dehydrogenase family 7 member A1 (EC
1.2.1.3) (Turgor- responsive protein 26G) (Antiquitin 1).
Length = 507
Score = 610 bits (1573), Expect = e-173
Identities = 298/391 (76%), Positives = 332/391 (84%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+EIIDMCDY+VGLSRQLNGSIIPSERP HMM EVWNPLG+VGVITAFNFPCAVLGWNACI
Sbjct: 117 QEIIDMCDYSVGLSRQLNGSIIPSERPEHMMFEVWNPLGIVGVITAFNFPCAVLGWNACI 176
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
ALV GN VVWKGAPTTPLIT+A+TK++A V E+NNLPGAIFT+ CGG +IG AIA D RI
Sbjct: 177 ALVGGNTVVWKGAPTTPLITVAVTKLIAEVFERNNLPGAIFTALCGGADIGHAIAKDTRI 236
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PLVSFTGS++ G +VQQ VS RFGK LLELSGNNAIIVMDDADI LAVRS+ FAAVGTAG
Sbjct: 237 PLVSFTGSSKVGALVQQTVSQRFGKTLLELSGNNAIIVMDDADITLAVRSIFFAAVGTAG 296
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTTCRRL LHE++Y L+QL +YKQV+IG+PLE+GTL+GPLHT ++ ENF GI
Sbjct: 297 QRCTTCRRLYLHESVYANVLEQLTALYKQVKIGNPLEEGTLVGPLHTRSAVENFKNGISA 356
Query: 722 IKSQGGKILFGGSAIESEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEAIEIN 901
IKSQGGKI+ GGS +ESEGNFV PTIVEI+ A VVKEELF PVLYVMKF+ L+EAI +N
Sbjct: 357 IKSQGGKIVTGGSVLESEGNFVVPTIVEISADAAVVKEELFAPVLYVMKFKDLEEAIALN 416
Query: 902 NSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXXXX 1081
NSVPQGLSSSIFT++P IFKW+GP GSDCGIVNVNIPTNGAEI
Sbjct: 417 NSVPQGLSSSIFTQKPSTIFKWIGPSGSDCGIVNVNIPTNGAEIGGAFGGEKATGGGREA 476
Query: 1082 XSDSWKQYMRRATCTINYGSELPLAQGINFG 1174
SDSWKQYMRR+TCTINYGSELPLAQGINFG
Sbjct: 477 GSDSWKQYMRRSTCTINYGSELPLAQGINFG 507
>sptr|Q84P31|Q84P31 Aldehyde dehydrogenase family 7 member A1 (EC
1.2.1.3).
Length = 510
Score = 604 bits (1558), Expect = e-172
Identities = 305/394 (77%), Positives = 328/394 (83%), Gaps = 3/394 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+EIIDMC+Y VGLSRQLNGSIIPSERP+HMM EVWNPLG+VGVI+AFNFPCAVLGWNACI
Sbjct: 119 QEIIDMCNYGVGLSRQLNGSIIPSERPDHMMFEVWNPLGIVGVISAFNFPCAVLGWNACI 178
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
ALVCGNCVVWKGAPTTPLITIA+TK+VA VLE+N LPGAIFTSFCGG +IGQAIA D RI
Sbjct: 179 ALVCGNCVVWKGAPTTPLITIAVTKLVAEVLERNKLPGAIFTSFCGGADIGQAIAKDTRI 238
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAV-GTA 538
PLVSFTGS++ GLMVQQ V+ RFGKCLLELSGNNAIIVMDDAD V F G+
Sbjct: 239 PLVSFTGSSKVGLMVQQTVNERFGKCLLELSGNNAIIVMDDADSNWLY--VYFGCCCGST 296
Query: 539 GQRCTT--CRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKG 712
GQ T CRRL LHE+IY LDQLVEVYKQV+ G+PLEKGTL+GPLHT S ENF KG
Sbjct: 297 GQPVYTWPCRRLFLHESIYTDVLDQLVEVYKQVKTGNPLEKGTLVGPLHTRTSVENFQKG 356
Query: 713 IQTIKSQGGKILFGGSAIESEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEAI 892
I IKSQGGKIL GGS +ES GNFVQPTIVEI+P APVVKEELFGPVLYVMKFQTL EAI
Sbjct: 357 ISVIKSQGGKILTGGSVLESGGNFVQPTIVEISPDAPVVKEELFGPVLYVMKFQTLDEAI 416
Query: 893 EINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXX 1072
+NNSVPQGLSSSIFT+RP IFKW+GP GSDCGIVN NIPTNGAEI
Sbjct: 417 ALNNSVPQGLSSSIFTQRPGTIFKWIGPRGSDCGIVNANIPTNGAEIGGAFGGEKATGGG 476
Query: 1073 XXXXSDSWKQYMRRATCTINYGSELPLAQGINFG 1174
SDSWKQYMRR+TCTINYGSELPLAQGINFG
Sbjct: 477 REAGSDSWKQYMRRSTCTINYGSELPLAQGINFG 510
>sw|Q41247|D7A1_BRANA Aldehyde dehydrogenase family 7 member A1 (EC
1.2.1.3) (Antiquitin 1) (Brassica
turgor-responsive/drought-induced gene 26 protein)
(Btg-26).
Length = 493
Score = 601 bits (1549), Expect = e-170
Identities = 291/373 (78%), Positives = 324/373 (86%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+IDMCD+AVGLSRQLNGS+IPSERPNHMM+E+WNPLG+VGVITAFNFPCAVLGWNACI
Sbjct: 120 QEVIDMCDFAVGLSRQLNGSVIPSERPNHMMLEMWNPLGIVGVITAFNFPCAVLGWNACI 179
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
ALVCGNCVVWKGAPTTPLITIAMTK+VA VLEKN+LPGAIFT+ CGG EIG+AIA D RI
Sbjct: 180 ALVCGNCVVWKGAPTTPLITIAMTKLVAEVLEKNHLPGAIFTAMCGGAEIGEAIAKDTRI 239
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PLVSFTGS++ GL VQQ VSAR GK LLELSGNNAIIVMDDADIQLA RSVLFAAVGTAG
Sbjct: 240 PLVSFTGSSKVGLTVQQTVSARSGKTLLELSGNNAIIVMDDADIQLAARSVLFAAVGTAG 299
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTTCRRL+LHE++Y L+QL+ YKQV+IGDPLEKGTLLGPLHTP SK+NF KGI+
Sbjct: 300 QRCTTCRRLLLHESVYDKVLEQLLTSYKQVKIGDPLEKGTLLGPLHTPESKKNFEKGIEV 359
Query: 722 IKSQGGKILFGGSAIESEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEAIEIN 901
IKSQGGK+L GG A+E EGNFV+PTI+EI+ A VVKEELF PVLY +KF+T +EA+ IN
Sbjct: 360 IKSQGGKVLTGGKAVEGEGNFVEPTIIEISSDAAVVKEELFAPVLYALKFKTFEEAVAIN 419
Query: 902 NSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXXXX 1081
NSVPQGLSSSIFT+ PD IFKW+GP GSDCGIVNVNIPTNGAEI
Sbjct: 420 NSVPQGLSSSIFTRSPDNIFKWIGPMGSDCGIVNVNIPTNGAEIGGAFGGEKATGGGREA 479
Query: 1082 XSDSWKQYMRRAT 1120
SDSWKQYMRR+T
Sbjct: 480 GSDSWKQYMRRST 492
>sptr|Q8RYB7|Q8RYB7 Aldehyde dehydrogenase ALDH2.
Length = 516
Score = 577 bits (1488), Expect = e-163
Identities = 280/391 (71%), Positives = 319/391 (81%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E IDMCDYAVGLSRQL+GSIIPSERPNH MMEVWNPLG+VGVITAFNFPCAVLGWNACI
Sbjct: 125 QEFIDMCDYAVGLSRQLSGSIIPSERPNHAMMEVWNPLGIVGVITAFNFPCAVLGWNACI 184
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
ALVCGNCVVWKGAPTTPL+T+A T+I+A V E+N LPG IFT CGG EIG AIA D RI
Sbjct: 185 ALVCGNCVVWKGAPTTPLLTLATTRIIAEVFERNKLPGGIFTCACGGAEIGSAIAHDARI 244
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PLVSFTGST+ G++VQ V AR GK LLELSGNNAIIVMDDA + LA+R+VLFAAVGTAG
Sbjct: 245 PLVSFTGSTKVGMLVQSIVHARHGKVLLELSGNNAIIVMDDAVLPLAIRAVLFAAVGTAG 304
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTTCRRLI+HE +Y L+ L++ YKQV++G+ +E TL GPLH+ S F +GI+
Sbjct: 305 QRCTTCRRLIVHEKVYDDMLEGLLKAYKQVKVGNSVEGDTLCGPLHSKLSMSGFTEGIKK 364
Query: 722 IKSQGGKILFGGSAIESEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEAIEIN 901
IK+QGG IL GGS I GNFV+PT+VEI+ A VV+EELFGPVLYV K ++L+EAIE+N
Sbjct: 365 IKAQGGNILTGGSTINRAGNFVEPTLVEISHDAEVVREELFGPVLYVFKIKSLEEAIEMN 424
Query: 902 NSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXXXX 1081
NSVPQGLSSSIFT+ P+ IF W+GP GSDCGIVNVNIPTNGAEI
Sbjct: 425 NSVPQGLSSSIFTRNPETIFTWIGPTGSDCGIVNVNIPTNGAEIGGAFGGEKATGGGREA 484
Query: 1082 XSDSWKQYMRRATCTINYGSELPLAQGINFG 1174
SDSWKQYM RATCTINYG +LPLAQGINFG
Sbjct: 485 GSDSWKQYMHRATCTINYGKDLPLAQGINFG 515
>sptr|Q8MYH7|Q8MYH7 Similar to Arabidopsis thaliana (Mouse-ear cress).
aldehyde dehydrogenase family 7, member A1 (EC 1.2.1.3)
(Antiquitin 1).
Length = 509
Score = 475 bits (1222), Expect = e-133
Identities = 224/392 (57%), Positives = 292/392 (74%), Gaps = 2/392 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E ID+CDYA GLSR +NG ++PSERPNH++ME WNPLG+VG+ITAFNFPCAVLGWNA I
Sbjct: 117 QEFIDVCDYATGLSRSINGQVMPSERPNHILMETWNPLGLVGIITAFNFPCAVLGWNAAI 176
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCG-GTEIGQAIALDIR 358
+++CGN +WKGA TT LIT+A++KI+ VL +N++ A+ G G +G+ + D R
Sbjct: 177 SMICGNVQLWKGASTTSLITLAVSKIIEKVLVENDVDPAVCCVLIGPGRTVGEQMIQDKR 236
Query: 359 IPLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTA 538
L+SFTGST G + V FGK +LEL GNNAI+V +DADI+L +R+VLFA+VGT
Sbjct: 237 FGLISFTGSTEVGRRISSTVHGYFGKTILELGGNNAIVVAEDADIELVLRAVLFASVGTT 296
Query: 539 GQRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQ 718
GQRCTTCRRL +HE++Y T L++L + YK ++IG+PLE+G L+GPLHT ++ + F +G++
Sbjct: 297 GQRCTTCRRLFVHESLYDTILERLTKAYKTIKIGNPLEEGVLVGPLHTQSAVKEFTEGLE 356
Query: 719 TIKSQGGKILFGGSAIE-SEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEAIE 895
IK QGGK++ GG+ ++ S GNFV+PT+V I AP+VK ELF P+LY+MKF+ L +A
Sbjct: 357 EIKKQGGKVVIGGNKLDISGGNFVEPTVVAIEHDAPIVKTELFVPILYIMKFKNLDDAFA 416
Query: 896 INNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXX 1075
NN VPQGLSSS+FT IFKWLGP GSDCGIVNVN+ TNGAEI
Sbjct: 417 WNNEVPQGLSSSLFTNNQKNIFKWLGPTGSDCGIVNVNVATNGAEIGGAFGGEKETGGGR 476
Query: 1076 XXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SDSWKQY RR+T TINYG+ +PL+QGINF
Sbjct: 477 ESGSDSWKQYCRRSTNTINYGNTMPLSQGINF 508
>sw|P83401|D7A1_DICDI Putative aldehyde dehydrogenase family 7 member
A1 homolog (EC 1.2.1.3) (Antiquitin 1).
Length = 509
Score = 475 bits (1222), Expect = e-133
Identities = 224/392 (57%), Positives = 292/392 (74%), Gaps = 2/392 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E ID+CDYA GLSR +NG ++PSERPNH++ME WNPLG+VG+ITAFNFPCAVLGWNA I
Sbjct: 117 QEFIDVCDYATGLSRSINGQVMPSERPNHILMETWNPLGLVGIITAFNFPCAVLGWNAAI 176
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCG-GTEIGQAIALDIR 358
+++CGN +WKGA TT LIT+A++KI+ VL +N++ A+ G G +G+ + D R
Sbjct: 177 SMICGNVQLWKGASTTSLITLAVSKIIEKVLVENDVDPAVCCVLIGPGRTVGEQMIQDKR 236
Query: 359 IPLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTA 538
L+SFTGST G + V FGK +LEL GNNAI+V +DADI+L +R+VLFA+VGT
Sbjct: 237 FGLISFTGSTEVGRRISSTVHGYFGKTILELGGNNAIVVAEDADIELVLRAVLFASVGTT 296
Query: 539 GQRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQ 718
GQRCTTCRRL +HE++Y T L++L + YK ++IG+PLE+G L+GPLHT ++ + F +G++
Sbjct: 297 GQRCTTCRRLFVHESLYDTILERLTKAYKTIKIGNPLEEGVLVGPLHTQSAVKEFTEGLE 356
Query: 719 TIKSQGGKILFGGSAIE-SEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEAIE 895
IK QGGK++ GG+ ++ S GNFV+PT+V I AP+VK ELF P+LY+MKF+ L +A
Sbjct: 357 EIKKQGGKVVIGGNKLDISGGNFVEPTVVAIEHDAPIVKTELFVPILYIMKFKNLDDAFA 416
Query: 896 INNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXX 1075
NN VPQGLSSS+FT IFKWLGP GSDCGIVNVN+ TNGAEI
Sbjct: 417 WNNEVPQGLSSSLFTNNQKNIFKWLGPTGSDCGIVNVNVATNGAEIGGAFGGEKETGGGR 476
Query: 1076 XXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SDSWKQY RR+T TINYG+ +PL+QGINF
Sbjct: 477 ESGSDSWKQYCRRSTNTINYGNTMPLSQGINF 508
>sptr|Q803R9|Q803R9 Similar to aldehyde dehydrogenase 7 family, member
A1.
Length = 511
Score = 473 bits (1218), Expect = e-132
Identities = 228/391 (58%), Positives = 288/391 (73%), Gaps = 1/391 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E +D+CDYAVGLSR + G I+PSERP H+++E WNP+G+VG+ITAFNFP AV GWN I
Sbjct: 120 QEYVDVCDYAVGLSRMIGGPILPSERPGHVLIEQWNPVGLVGIITAFNFPVAVYGWNNAI 179
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
AL+CGN +WKGAPTTPL ++A+TKIVA VLE+N+LPGAI + CGG +IG A+A D R+
Sbjct: 180 ALICGNACLWKGAPTTPLTSVAVTKIVAEVLEQNHLPGAICSMTCGGADIGMAMAKDERV 239
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
L+SFTGST G V V RFG+ LLEL GNNAIIV +DAD+ L V S +FA+VGTAG
Sbjct: 240 GLLSFTGSTHVGKQVAMMVQERFGRQLLELGGNNAIIVFEDADLSLVVPSAVFASVGTAG 299
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTT RRL+LHE+I+ ++++ + YKQ+RIGDP + TL GPLHT + + +L I+
Sbjct: 300 QRCTTTRRLMLHESIHDEVVERIAKAYKQIRIGDPWDPNTLYGPLHTKQAVQQYLAAIEQ 359
Query: 722 IKSQGGKILFGGSAIESEGNFVQPTIVEITP-SAPVVKEELFGPVLYVMKFQTLKEAIEI 898
K QGG ++ GG ++ GN+V+PTI+ P +A +V E F P+LYV+KF+T +EA
Sbjct: 360 AKQQGGTLVCGGKIMDRPGNYVEPTIITGLPHNASIVHTETFVPILYVLKFKTEEEAFSW 419
Query: 899 NNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXXX 1078
NN V QGLSSSIFTK +F+WLGP GSDCGIVNVNIPT+GAEI
Sbjct: 420 NNEVKQGLSSSIFTKDMGRVFRWLGPKGSDCGIVNVNIPTSGAEIGGAFGGEKHTGGGRE 479
Query: 1079 XXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SDSWKQYMRR+TCTINY +LPLAQGI F
Sbjct: 480 SGSDSWKQYMRRSTCTINYSKDLPLAQGIKF 510
>sw|Q9DBF1|D7A1_MOUSE Aldehyde dehydrogenase family 7 member A1 (EC
1.2.1.3) (Antiquitin 1).
Length = 510
Score = 462 bits (1189), Expect = e-129
Identities = 224/391 (57%), Positives = 279/391 (71%), Gaps = 1/391 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E +D+CDYA GLSR + G +PSERP H ++E+WNPLG+VG+ITAFNFP AV GWN I
Sbjct: 119 QEYVDVCDYAAGLSRMIGGPTLPSERPGHALIEMWNPLGLVGIITAFNFPVAVFGWNNAI 178
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
AL+ GN +WKGAPTT L+++A+TKI+A VLE N LPGAI + CGG +IG +A D R+
Sbjct: 179 ALITGNVCLWKGAPTTSLVSVAVTKIIAQVLEDNLLPGAICSLVCGGADIGTTMARDERV 238
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
L+SFTGST+ G V V RFGK LLEL GNNAII +DAD+ L V SVLFAAVGTAG
Sbjct: 239 NLLSFTGSTQVGKEVALMVQERFGKSLLELGGNNAIIAFEDADLSLVVPSVLFAAVGTAG 298
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTT RRL LHE+I+ +D+L Y Q+R+G+P + L GPLHT + F++ ++
Sbjct: 299 QRCTTVRRLFLHESIHNEVVDRLRSAYSQIRVGNPWDPNILYGPLHTKQAVSMFVRAVEE 358
Query: 722 IKSQGGKILFGGSAIESEGNFVQPTIVE-ITPSAPVVKEELFGPVLYVMKFQTLKEAIEI 898
K QGG +++GG ++ GN+V+PTIV + AP+V +E F P+LYV KFQ +E E
Sbjct: 359 AKKQGGTVVYGGKVMDHPGNYVEPTIVTGLAHDAPIVHQETFAPILYVFKFQDEEEVFEW 418
Query: 899 NNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXXX 1078
NN V QGLSSSIFTK IF+WLGP GSDCGIVNVNIPT+GAEI
Sbjct: 419 NNEVKQGLSSSIFTKDLGRIFRWLGPKGSDCGIVNVNIPTSGAEIGGAFGGEKHTGGGRE 478
Query: 1079 XXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+WKQYMRR+TCTINY + LPLAQGI F
Sbjct: 479 SGSDAWKQYMRRSTCTINYSTSLPLAQGIKF 509
>sw|P46562|D7A1_CAEEL Putative aldehyde dehydrogenase family 7 member
A1 homolog (EC 1.2.1.3).
Length = 514
Score = 453 bits (1165), Expect = e-126
Identities = 226/393 (57%), Positives = 277/393 (70%), Gaps = 3/393 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E +D+CDYA GLSR L G I PSERP H ++E WNPLGVVGVI+AFNFPCAV GWN +
Sbjct: 121 QEYVDICDYATGLSRSLEGKIFPSERPGHALLEQWNPLGVVGVISAFNFPCAVYGWNNAL 180
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
ALV GN VVWK AP+TPL IA+TK+V VL NN+ A+ + CG ++GQA+ D R+
Sbjct: 181 ALVTGNSVVWKPAPSTPLTAIAVTKLVEEVLVANNVNPALCSLVCGEGDVGQALVKDKRV 240
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
LVSFTGS+ G +V QQV ARFGK LLEL GNNAIIV +DAD+ + V + +FAAVGTAG
Sbjct: 241 NLVSFTGSSEIGKIVGQQVQARFGKLLLELGGNNAIIVNEDADLNMVVPATVFAAVGTAG 300
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQV--RIGDPLEKGTLLGPLHTPASKENFLKGI 715
QRCTT RRLI+H+ +Y L++L + Y Q RIG PL+ T++GPLH + + +
Sbjct: 301 QRCTTTRRLIVHDKVYDQVLERLKKAYAQFESRIGCPLDSNTIIGPLHNQQAVGKYKASV 360
Query: 716 QTIKSQGGKILFGGSAIESEGNFVQPTIVE-ITPSAPVVKEELFGPVLYVMKFQTLKEAI 892
+ GGKI +GG +E +GNFV PTIV + +PVV E F P+LYV+KF TL+EAI
Sbjct: 361 AEAVASGGKIEYGGKVLERDGNFVLPTIVTGLKHDSPVVLRETFAPILYVLKFSTLEEAI 420
Query: 893 EINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXX 1072
INN V QGLSSS+FT +FKW+GP GSDCGIVNVNIPT+GAEI
Sbjct: 421 AINNEVDQGLSSSLFTTNIQNVFKWMGPKGSDCGIVNVNIPTSGAEIGGAFGGEKETGGG 480
Query: 1073 XXXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SDSW+QYMRR+TCTINY ELPLAQGI F
Sbjct: 481 RESGSDSWRQYMRRSTCTINYSKELPLAQGIKF 513
>sptrnew|AAH02515|AAH02515 Antiquitin (Fragment).
Length = 542
Score = 452 bits (1163), Expect = e-126
Identities = 219/391 (56%), Positives = 276/391 (70%), Gaps = 1/391 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E +D+CDYAVGLSR + G I+PSER H ++E WNP+G+VG+ITAFNFP AV GWN I
Sbjct: 151 QEYVDICDYAVGLSRMIGGPILPSERSGHALIEQWNPVGLVGIITAFNFPVAVYGWNNAI 210
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
A++CGN +WKGAPTT LI++A+TKI+A VLE N LPGAI + CGG +IG A+A D R+
Sbjct: 211 AMICGNVCLWKGAPTTSLISVAVTKIIAKVLEDNKLPGAICSLTCGGADIGTAMAKDERV 270
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
L+SFTGST+ G V V RFG+ LLEL GNNAII +DAD+ L V S LFAAVGTAG
Sbjct: 271 NLLSFTGSTQVGKQVGLMVQERFGRSLLELGGNNAIIAFEDADLSLVVPSALFAAVGTAG 330
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTT RRL +HE+I+ +++L + Y Q+R+G+P + L GPLHT + FL ++
Sbjct: 331 QRCTTARRLFIHESIHDEVVNRLKKAYAQIRVGNPWDPNVLYGPLHTKQAVSMFLGAVEE 390
Query: 722 IKSQGGKILFGGSAIESEGNFVQPTIVE-ITPSAPVVKEELFGPVLYVMKFQTLKEAIEI 898
K +GG +++GG ++ GN+V+PTIV + A + E F P+LYV KFQ +E
Sbjct: 391 AKKEGGTVVYGGKVMDRPGNYVEPTIVTGLGHDASIAHTETFAPILYVFKFQNEEEVFAW 450
Query: 899 NNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXXX 1078
NN V QGLSSSIFTK IF+WLGP GSDCGIVNVNIPT+GAEI
Sbjct: 451 NNEVKQGLSSSIFTKDLGRIFRWLGPKGSDCGIVNVNIPTSGAEIGGAFGGEKHTGGGRE 510
Query: 1079 XXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+WKQYMRR+TCTINY +LPLAQGI F
Sbjct: 511 SGSDAWKQYMRRSTCTINYSKDLPLAQGIKF 541
>sw|P49419|D7A1_HUMAN Aldehyde dehydrogenase family 7 member A1 (EC
1.2.1.3) (Antiquitin 1).
Length = 510
Score = 452 bits (1163), Expect = e-126
Identities = 219/391 (56%), Positives = 276/391 (70%), Gaps = 1/391 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E +D+CDYAVGLSR + G I+PSER H ++E WNP+G+VG+ITAFNFP AV GWN I
Sbjct: 119 QEYVDICDYAVGLSRMIGGPILPSERSGHALIEQWNPVGLVGIITAFNFPVAVYGWNNAI 178
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
A++CGN +WKGAPTT LI++A+TKI+A VLE N LPGAI + CGG +IG A+A D R+
Sbjct: 179 AMICGNVCLWKGAPTTSLISVAVTKIIAKVLEDNKLPGAICSLTCGGADIGTAMAKDERV 238
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
L+SFTGST+ G V V RFG+ LLEL GNNAII +DAD+ L V S LFAAVGTAG
Sbjct: 239 NLLSFTGSTQVGKQVGLMVQERFGRSLLELGGNNAIIAFEDADLSLVVPSALFAAVGTAG 298
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTT RRL +HE+I+ +++L + Y Q+R+G+P + L GPLHT + FL ++
Sbjct: 299 QRCTTARRLFIHESIHDEVVNRLKKAYAQIRVGNPWDPNVLYGPLHTKQAVSMFLGAVEE 358
Query: 722 IKSQGGKILFGGSAIESEGNFVQPTIVE-ITPSAPVVKEELFGPVLYVMKFQTLKEAIEI 898
K +GG +++GG ++ GN+V+PTIV + A + E F P+LYV KFQ +E
Sbjct: 359 AKKEGGTVVYGGKVMDRPGNYVEPTIVTGLGHDASIAHTETFAPILYVFKFQNEEEVFAW 418
Query: 899 NNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXXXX 1078
NN V QGLSSSIFTK IF+WLGP GSDCGIVNVNIPT+GAEI
Sbjct: 419 NNEVKQGLSSSIFTKDLGRIFRWLGPKGSDCGIVNVNIPTSGAEIGGAFGGEKHTGGGRE 478
Query: 1079 XXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+WKQYMRR+TCTINY +LPLAQGI F
Sbjct: 479 SGSDAWKQYMRRSTCTINYSKDLPLAQGIKF 509
>sptr|Q8MUI1|Q8MUI1 Aldehyde dehydrogenase.
Length = 514
Score = 446 bits (1146), Expect = e-124
Identities = 217/393 (55%), Positives = 278/393 (70%), Gaps = 3/393 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E +D+CDYAVGLSR +G +IPSERP H+++E WNPLGVVGVI+AFNFPCAV GWN+ +
Sbjct: 121 QEYVDICDYAVGLSRMFSGKVIPSERPGHVLLEQWNPLGVVGVISAFNFPCAVYGWNSAL 180
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
A+V GN ++WK AP+TPL IA+TK++ VL +N LPG I + G ++GQA+ D R+
Sbjct: 181 AMVTGNSLIWKPAPSTPLTAIAITKLIEEVLSENKLPGGICSLVTGEADVGQALTKDERV 240
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
LVSFTGST G +V QQV RFGK LLEL GNNA++V DDAD+ + V + +FAAVGTAG
Sbjct: 241 NLVSFTGSTEVGRIVGQQVQGRFGKVLLELGGNNALLVNDDADLDMVVPATVFAAVGTAG 300
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQV--RIGDPLEKGTLLGPLHTPASKENFLKGI 715
QRCT+ RRLI+HE+I+ ++++ + Y Q+ RIGDPLE T++GPLH + + +
Sbjct: 301 QRCTSTRRLIVHESIHDQVVERMKKAYAQLESRIGDPLESNTIIGPLHNQQAVLKYKATV 360
Query: 716 QTIKSQGGKILFGGSAIESEGNFVQPTIVE-ITPSAPVVKEELFGPVLYVMKFQTLKEAI 892
QGGKI GG +E +GNF PTIV + A VV E F P+LYV+K + L E I
Sbjct: 361 AEAIVQGGKIECGGKIVEGDGNFALPTIVTGLKHDAEVVLRETFAPILYVIKMKNLDEGI 420
Query: 893 EINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXX 1072
+ NN V QGLSSS+FT+ + IFKW+GP GSDCGIVN+NIPT+GAEI
Sbjct: 421 KFNNEVKQGLSSSLFTQNLENIFKWMGPKGSDCGIVNINIPTSGAEIGGAFGGEKETGGG 480
Query: 1073 XXXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+WKQYMRR+TCTINY LPLAQGI F
Sbjct: 481 RESGSDAWKQYMRRSTCTINYSKNLPLAQGIKF 513
>sptr|Q8SXQ1|Q8SXQ1 GH05218p (CG9629 protein).
Length = 540
Score = 404 bits (1039), Expect = e-111
Identities = 201/393 (51%), Positives = 271/393 (68%), Gaps = 3/393 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E ID+CDYAVGLSR +G +I SER +H ++E W PLGVVGVI+A+NFP AV GWNA I
Sbjct: 146 QEFIDICDYAVGLSRIYSGQLINSERADHSILEAWRPLGVVGVISAYNFPNAVFGWNAAI 205
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
AL GN V+WKGAP+TPL+++A TKIVA VL +NNLP + T GGT++GQ + D R+
Sbjct: 206 ALTTGNSVLWKGAPSTPLVSVATTKIVAEVLRRNNLP-PVVTLCQGGTDVGQTLVADKRV 264
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
LVSFTGS + G V +V RFGK +LEL GNNA+I+ + A++++A+ + LF +GT+G
Sbjct: 265 NLVSFTGSCQTGRDVGVEVQRRFGKVILELGGNNALIIDESANVKMALDAALFGCIGTSG 324
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQV--RIGDPLEKGTLLGPLHTPASKENFLKGI 715
QRCTT RR+I+HE ++ F+ +LV YKQ+ +IG LE TL+GP+HT + EN+ I
Sbjct: 325 QRCTTTRRIIVHEKLHDQFVKELVGKYKQLISKIGHQLEAQTLVGPVHTQQNVENYKAAI 384
Query: 716 QTIKSQGGKILFGGSAIESEGNFVQPTIVEITP-SAPVVKEELFGPVLYVMKFQTLKEAI 892
KS GG + FGG+ I+ +G +V+PT++ P A VV E F P++Y++K + + +AI
Sbjct: 385 AEAKSLGGTVAFGGNVIQRDGFYVEPTVITGLPHDASVVHRETFAPIVYILKAKNVDQAI 444
Query: 893 EINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXX 1072
E NN V QGLSS+IFT+ FKW+G GSDCGIVN+N TNGAEI
Sbjct: 445 EWNNEVEQGLSSAIFTENIGQAFKWIGAKGSDCGIVNINTTTNGAEIGGAFGGEKATGGG 504
Query: 1073 XXXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+WKQY +RAT T+N+ EL AQG+ F
Sbjct: 505 RESGSDAWKQYCKRATITVNHSGELACAQGVVF 537
>sptr|Q9F1U8|Q9F1U8 Piperideine-6-carboxylate dehydrogenase.
Length = 510
Score = 390 bits (1002), Expect = e-107
Identities = 202/393 (51%), Positives = 259/393 (65%), Gaps = 3/393 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+ID+ D+AVG SR L G + SERP H M E + PLG+VG+I+AFNFP AV WN+ +
Sbjct: 116 QEMIDIADFAVGQSRMLYGYTMHSERPGHRMYEQYQPLGIVGIISAFNFPVAVWAWNSFL 175
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
A +CG+ +WK + TPL IA +I L + P F GT + + + D R+
Sbjct: 176 AAICGDVCIWKPSNKTPLTAIASMRICNEALREGGFPDIFFLINDAGTALSEKLVEDKRV 235
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PL+SFTGST+ G +V Q+V+AR G+CLLEL GNNAII+ + AD++LAV ++F AVGTAG
Sbjct: 236 PLISFTGSTQVGRIVNQKVAARLGRCLLELGGNNAIILDETADLKLAVPGIVFGAVGTAG 295
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQV--RIGDPLEKGTLLGPLHTPASKENFLKGI 715
QRCTT RRLI+HE+IY L L++ YKQV +IGDPL+ L+GPL++P + + FL I
Sbjct: 296 QRCTTTRRLIVHESIYDNVLATLIKAYKQVEGKIGDPLDAANLMGPLNSPEAVQQFLASI 355
Query: 716 QTIKSQGGKILFGGSAIESEGNFVQPTIVE-ITPSAPVVKEELFGPVLYVMKFQTLKEAI 892
+ K+ GG + GG+AI+ GNFV P IV + S VV+ E F P+LYVMK+ TL EAI
Sbjct: 356 EKAKAAGGTVQTGGTAIDRPGNFVLPAIVTGLKNSDEVVQHETFAPILYVMKYSTLDEAI 415
Query: 893 EINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXX 1072
E+ N VPQGLSSSIFT K+L GSDCGI NVNI T+GAEI
Sbjct: 416 EMQNGVPQGLSSSIFTTNLKAAEKFLSAAGSDCGIANVNIGTSGAEIGGAFGGEKETGGG 475
Query: 1073 XXXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+WK YMRR T TINY LPLAQGI F
Sbjct: 476 RESGSDAWKVYMRRQTNTINYSDSLPLAQGIKF 508
>sptr|Q8P9Q7|Q8P9Q7 Aldehyde dehydrogenase.
Length = 510
Score = 383 bits (983), Expect = e-105
Identities = 200/393 (50%), Positives = 259/393 (65%), Gaps = 3/393 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+ID+ D+AVG SR L G + SERP H M E + PLG+VG+I+AFNFP AV WNA +
Sbjct: 116 QEMIDIADFAVGQSRMLYGYTMHSERPGHRMYEQYQPLGLVGIISAFNFPVAVWAWNAFL 175
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
A +CG+ +WK + TPL IA KI + L++ P F GT + + + D R+
Sbjct: 176 AAICGDICIWKPSNKTPLTAIATMKICNAALKEAGFPDIFFLINDAGTSLSEKLVEDGRV 235
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PL+SFTGST+ G +V ++V+ R G+CLLEL GNNAII+ + AD++LAV ++F AVGTAG
Sbjct: 236 PLISFTGSTQIGRVVAEKVARRLGRCLLELGGNNAIILDETADLKLAVPGIVFGAVGTAG 295
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQV--RIGDPLEKGTLLGPLHTPASKENFLKGI 715
QRCTT RRLI+HE+IY L LV+ YKQ+ +IGDP + L+GPL++ + E FL+ I
Sbjct: 296 QRCTTTRRLIVHESIYDNVLATLVKAYKQLDSKIGDPTDPANLMGPLNSRGAVEQFLESI 355
Query: 716 QTIKSQGGKILFGGSAIESEGNFVQPTIVE-ITPSAPVVKEELFGPVLYVMKFQTLKEAI 892
K+ GG + GG+AI+ GNFV P IV + S VV+ E F P+LYVMK+ TL EAI
Sbjct: 356 AKAKAAGGTVEVGGTAIDRPGNFVLPAIVTGLKNSDAVVQHETFAPILYVMKYSTLDEAI 415
Query: 893 EINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXX 1072
++ N VPQGLSSSIFT+ K+L GSDCGI NVNI T+GAEI
Sbjct: 416 DLQNGVPQGLSSSIFTQNLKAAEKFLSAAGSDCGIANVNIGTSGAEIGGAFGGEKETGGG 475
Query: 1073 XXXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+WK YMRR T TINY LPLAQGI F
Sbjct: 476 RESGSDAWKVYMRRQTNTINYSDSLPLAQGIKF 508
>sptr|Q8PLI9|Q8PLI9 Aldehyde dehydrogenase.
Length = 510
Score = 380 bits (977), Expect = e-104
Identities = 199/393 (50%), Positives = 256/393 (65%), Gaps = 3/393 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+ID+ D+AVG SR L G + SERP H M E + PLG+VG+I+AFNFP AV WNA +
Sbjct: 116 QEMIDIADFAVGQSRMLYGYTMHSERPGHRMYEQYQPLGLVGIISAFNFPVAVWAWNAFL 175
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
A +CG+ +WK + TPL IA KI + L+ P F GT + + + D R+
Sbjct: 176 AAICGDICIWKPSNKTPLTAIATMKICNAALKDAGFPDIFFLINDAGTALSEKLVEDTRV 235
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PL+SFTGST+ G +V ++V+ R G+CLLEL GNNAII+ + AD++LA+ ++F AVGTAG
Sbjct: 236 PLISFTGSTQIGRVVAEKVARRLGRCLLELGGNNAIILDETADLKLAIPGIVFGAVGTAG 295
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQV--RIGDPLEKGTLLGPLHTPASKENFLKGI 715
QRCTT RRLI+H +IY T L LV+ YKQ+ +IGDP + L+GPL++ + FL I
Sbjct: 296 QRCTTTRRLIVHASIYDTVLATLVKAYKQLDSKIGDPTDPANLMGPLNSQGAVAQFLDSI 355
Query: 716 QTIKSQGGKILFGGSAIESEGNFVQPTIVE-ITPSAPVVKEELFGPVLYVMKFQTLKEAI 892
K+ GG + GG+AI+ GNFV P IV + S VV+ E F P+LYVMK+ TL EAI
Sbjct: 356 AKAKAAGGTVEVGGTAIDRPGNFVLPAIVTGLQNSDAVVQHETFAPILYVMKYSTLDEAI 415
Query: 893 EINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXXX 1072
E+ N VPQGLSSSIFT+ K+L GSDCGI NVNI T+GAEI
Sbjct: 416 ELQNGVPQGLSSSIFTQNLKAAEKFLSAAGSDCGIANVNIGTSGAEIGGAFGGEKETGGG 475
Query: 1073 XXXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+WK YMRR T TINY LPLAQGI F
Sbjct: 476 RESGSDAWKVYMRRQTNTINYSDSLPLAQGIKF 508
>sptr|Q979S8|Q979S8 Aldehyde dehydrogenase.
Length = 514
Score = 380 bits (975), Expect = e-104
Identities = 195/394 (49%), Positives = 255/394 (64%), Gaps = 4/394 (1%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+ID+ D A+GLSRQL G I SERP H M E W PLG + VIT+FNFP +V WN+ I
Sbjct: 119 QEMIDISDLALGLSRQLYGLTIASERPYHRMYEQWVPLGPIAVITSFNFPASVWSWNSFI 178
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
A V G+ V+WK + L +A+ K++ VL++NN+P + G+E G I D RI
Sbjct: 179 AAVTGDVVIWKPSSKAALTAVAVMKVIERVLKRNNVPNIFYLMVSSGSEGGDWITNDPRI 238
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PLVSFTGS G + + V+ R GK +LEL GNN IV D ADI LA+R V+F A+ TAG
Sbjct: 239 PLVSFTGSVPVGRKISENVAKRLGKSILELGGNNGAIVSDKADIDLALRGVVFGALATAG 298
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTT RR+I+ E IY F+ +LV Y +V++GDP EKG L+GPL + ++ I
Sbjct: 299 QRCTTTRRVIVSEKIYDEFVKKLVNAYSKVKVGDPREKGVLVGPLIDEDAVRDYEGAISE 358
Query: 722 IKSQGGKILFGGSAIE----SEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEA 889
QGGK+L+GG I +G++V+PTI+E P+ P+VKEE F P+LYVMK++T++EA
Sbjct: 359 AVKQGGKVLYGGKKISLKGYEKGHYVEPTIIEANPNMPIVKEETFAPILYVMKYKTIEEA 418
Query: 890 IEINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXX 1069
+EI+NSVPQGLSSSIFT +L P+GSDCG+ NVN T GAEI
Sbjct: 419 LEIHNSVPQGLSSSIFTTDLREEEAFLSPYGSDCGLANVNTSTAGAEIGGAFGGEKDTGG 478
Query: 1070 XXXXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+WK YMRR T T N+G LPLAQ + F
Sbjct: 479 GRESGSDAWKFYMRRQTVTKNWGQTLPLAQDVVF 512
>sptr|Q9BME2|Q9BME2 Turgor-like protein (Fragment).
Length = 473
Score = 377 bits (967), Expect = e-103
Identities = 193/345 (55%), Positives = 243/345 (70%), Gaps = 1/345 (0%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E ID+CD A GLSR + G ++PSERP H MME WNPLG+VG ITAFNFP AV GWNA I
Sbjct: 128 QEYIDICDMATGLSRTIEGKVLPSERPGHFMMENWNPLGLVGCITAFNFPVAVSGWNATI 187
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCG-GTEIGQAIALDIR 358
A +CG+ ++WK A TT L IA IV VL+K +I T G G +G + D R
Sbjct: 188 AFICGDLMIWKPALTTCLCAIATQIIVNEVLQKFGF-NSIQTLCAGDGPNVGALLVKDPR 246
Query: 359 IPLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTA 538
+ L+SFTGST G V Q+V+ RFG+ +LEL GNNA ++M DAD++LA+++ +FAAVGTA
Sbjct: 247 LQLISFTGSTAVGRQVSQEVAKRFGRTILELGGNNAAVIMPDADLELALKACVFAAVGTA 306
Query: 539 GQRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQ 718
GQRCTT RR+I+HE+I+ D+ VE Y TLLGPLHT S+ FL+GI+
Sbjct: 307 GQRCTTLRRIIIHESIH----DKFVEAYD-----------TLLGPLHTKHSQNQFLEGIE 351
Query: 719 TIKSQGGKILFGGSAIESEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEAIEI 898
IKSQGGK+L GG+ + EGN+V+PTI+ I A VK ELF PV+YVMKF TL+EAI+
Sbjct: 352 AIKSQGGKVLVGGNIVPGEGNYVEPTIISINHDAECVKHELFAPVVYVMKFNTLEEAIQY 411
Query: 899 NNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEI 1033
NN VPQGLSS++FT+ F W+GP GSDCGIVN NI T+GAEI
Sbjct: 412 NNEVPQGLSSTLFTRDIQNYFNWVGPAGSDCGIVNCNIGTSGAEI 456
>sptr|Q9HL01|Q9HL01 Piperideine-6-carboxilic acid dehydrogenase
related protein.
Length = 512
Score = 376 bits (966), Expect = e-103
Identities = 197/394 (50%), Positives = 251/394 (63%), Gaps = 4/394 (1%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+ID+ D A+GLSRQL G I SERP H M E W PLG V VIT+FNFP +V WNA I
Sbjct: 117 QEMIDISDLALGLSRQLYGLTIASERPYHRMYEQWVPLGPVAVITSFNFPASVWSWNAFI 176
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
A V G+ V+WK + L +A+ K++ VL++NN P F G+E G I D RI
Sbjct: 177 AAVDGDVVIWKPSSKAALTAVAVMKVIERVLKRNNAPNIFFLMVSSGSEGGDWITNDPRI 236
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PLVSFTGS G + + V+ R GK +LEL GNN IV D ADI LA+R V+F A+ TAG
Sbjct: 237 PLVSFTGSVPVGKKISENVAKRLGKSILELGGNNGAIVSDKADIDLALRGVVFGALATAG 296
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTT RR+I+ E IY FL++L Y +++GDP EKG L+GPL + +++ I
Sbjct: 297 QRCTTTRRVIVQEKIYDQFLEKLKNAYSTIKVGDPREKGVLVGPLIDEDAVKDYEHAIDL 356
Query: 722 IKSQGGKILFGGSAIESE----GNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEA 889
QGGK++FGG I + G++V PTI+E P P+VKEE F +LYVMK++T++EA
Sbjct: 357 AIKQGGKLIFGGKRITIKGLEGGHYVMPTIIEAKPDMPIVKEETFASILYVMKYRTIEEA 416
Query: 890 IEINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXX 1069
IEI+NSVPQGLSSSIFT +L P+GSDCG+ NVN T GAEI
Sbjct: 417 IEIHNSVPQGLSSSIFTTDLREEEAFLSPYGSDCGLANVNTSTAGAEIGGAFGGEKDTGG 476
Query: 1070 XXXXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+WK YMRR T T N+G LPLAQ + F
Sbjct: 477 GRESGSDAWKYYMRRQTVTKNWGQTLPLAQDVVF 510
>sptr|Q8XPP7|Q8XPP7 Putative transmembrane aldehyde dehydrogenase
oxidoreductase protein (EC 1.2.1.-).
Length = 504
Score = 366 bits (939), Expect = e-100
Identities = 190/400 (47%), Positives = 257/400 (64%), Gaps = 10/400 (2%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+ID+CD+AVGLSRQL G I SERP H MME W+P+GVVGVI+AFNFP AV WN+ +
Sbjct: 104 QEMIDICDFAVGLSRQLYGLTIASERPGHRMMETWHPVGVVGVISAFNFPAAVWAWNSAL 163
Query: 182 ALVCGNCVVWKGAPTTPLITIA----MTKIVASV--LEKNNLPGAIFTSFCGGTEIGQAI 343
A VCG+ VVWK + TPL +A K VA+ + P + GG E G+A+
Sbjct: 164 AFVCGDSVVWKPSEKTPLTALACDALFRKAVAAFGKAHQGTAPDGLHALIVGGREAGEAL 223
Query: 344 ALDIRIPLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFA 523
R+P+VS TGSTR G +V +V R G+ +LEL GNNA+IV AD++LA R++ FA
Sbjct: 224 VQSKRVPVVSATGSTRMGRLVSARVGERLGRAILELGGNNAMIVAPSADLELATRAITFA 283
Query: 524 AVGTAGQRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENF 703
AVGTAGQRCTT RRLI+HE++ +++L +Y V +G+PL++GTL+GPL +
Sbjct: 284 AVGTAGQRCTTLRRLIVHESVAANLVERLKRIYGSVTVGNPLQEGTLVGPLIDAGAYAAM 343
Query: 704 LKGIQTIKSQGGKILFGGSAIESEGN----FVQPTIVEITPSAPVVKEELFGPVLYVMKF 871
+ ++ +QGG++ GG E +V+P +VE+ V+++E F P+LYV+ +
Sbjct: 344 ARALERADAQGGRV-HGGERTHLEAGPEAYYVKPALVEMPAQTEVMQQETFAPILYVLTY 402
Query: 872 QTLKEAIEINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXX 1051
+TL EAI + N VPQGLSS+IFT+ + +L GSDCGI NVNI T+GAEI
Sbjct: 403 RTLDEAIALQNGVPQGLSSAIFTRDLNEAEWFLSAAGSDCGIANVNIGTSGAEIGGAFGG 462
Query: 1052 XXXXXXXXXXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+WK YMRRAT TINY + LPLAQG+ F
Sbjct: 463 EKETGGGRESGSDAWKAYMRRATNTINYSNRLPLAQGVRF 502
>sptr|Q7W8G8|Q7W8G8 Probable aldehyde dehydrogenase.
Length = 500
Score = 362 bits (930), Expect = 6e-99
Identities = 193/396 (48%), Positives = 249/396 (62%), Gaps = 5/396 (1%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+ID+CD+AVGLSRQL G I SERP H MME W+PLGVVGVI+AFNFP AV WNA +
Sbjct: 105 QEMIDICDFAVGLSRQLYGLTIASERPGHRMMETWHPLGVVGVISAFNFPVAVWSWNAAL 164
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEK-NNLPGAIFTSFCGGTEIGQAIALDIR 358
ALVCGN VVWK + TPL +A ++A L+ P + GG ++G+A+
Sbjct: 165 ALVCGNAVVWKPSEKTPLTALACQALLARALQDYGEAPAGLAEVLIGGRQLGEALVDSHD 224
Query: 359 IPLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTA 538
+ LVS TGSTR G V +V+ RFG+ LLEL GNNAIIV AD+++A R ++F A+GTA
Sbjct: 225 VALVSATGSTRMGRQVGPRVAQRFGRVLLELGGNNAIIVAPSADLEMATRGIVFGAIGTA 284
Query: 539 GQRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQ 718
GQRCTT RRLI H +I T + +L + Y +GDPL+ GTL+GPL A+ + +
Sbjct: 285 GQRCTTTRRLIAHTDIADTLVARLKQAYASATVGDPLQPGTLVGPLIDRAAYDAMQAALT 344
Query: 719 TIKSQGGKILFGGSAIESE----GNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKE 886
+ QGG++ GG + +E +V+P IVE+ VV E F P+LYVM++
Sbjct: 345 AAREQGGQV-SGGERLLAERYPDAWYVRPAIVEMPGQTDVVCHETFAPILYVMRYGDFPA 403
Query: 887 AIEINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXX 1066
A+ ++N VPQGLSS+IFT +L GSDCGI NVNI T+GAEI
Sbjct: 404 ALAMHNGVPQGLSSAIFTNDLREAEAFLSSAGSDCGIANVNIGTSGAEIGGAFGGEKETG 463
Query: 1067 XXXXXXSDSWKQYMRRATCTINYGSELPLAQGINFG 1174
SD+WK YMRRAT T+NY LPLAQGI FG
Sbjct: 464 GGRESGSDAWKHYMRRATNTVNYSRALPLAQGIRFG 499
>sptr|Q7WM30|Q7WM30 Probable aldehyde dehydrogenase.
Length = 500
Score = 362 bits (930), Expect = 6e-99
Identities = 193/396 (48%), Positives = 249/396 (62%), Gaps = 5/396 (1%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+ID+CD+AVGLSRQL G I SERP H MME W+PLGVVGVI+AFNFP AV WNA +
Sbjct: 105 QEMIDICDFAVGLSRQLYGLTIASERPGHRMMETWHPLGVVGVISAFNFPVAVWSWNAAL 164
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEK-NNLPGAIFTSFCGGTEIGQAIALDIR 358
ALVCGN VVWK + TPL +A ++A L+ P + GG ++G+A+
Sbjct: 165 ALVCGNAVVWKPSEKTPLTALACQALLARALQDYGEAPAGLAEVLIGGRQLGEALVDSHD 224
Query: 359 IPLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTA 538
+ LVS TGSTR G V +V+ RFG+ LLEL GNNAIIV AD+++A R ++F A+GTA
Sbjct: 225 VALVSATGSTRMGRQVGPRVAQRFGRVLLELGGNNAIIVAPSADLEMATRGIVFGAIGTA 284
Query: 539 GQRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQ 718
GQRCTT RRLI H +I T + +L + Y +GDPL+ GTL+GPL A+ + +
Sbjct: 285 GQRCTTTRRLIAHTDIADTLVARLKQAYASATVGDPLQPGTLVGPLIDRAAYDAMQAALT 344
Query: 719 TIKSQGGKILFGGSAIESE----GNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKE 886
+ QGG++ GG + +E +V+P IVE+ VV E F P+LYVM++
Sbjct: 345 AAREQGGQV-SGGERLLAERYPDAWYVRPAIVEMPGQTDVVCHETFAPILYVMRYGDFPA 403
Query: 887 AIEINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXX 1066
A+ ++N VPQGLSS+IFT +L GSDCGI NVNI T+GAEI
Sbjct: 404 ALAMHNGVPQGLSSAIFTNDLREAEAFLSSAGSDCGIANVNIGTSGAEIGGAFGGEKETG 463
Query: 1067 XXXXXXSDSWKQYMRRATCTINYGSELPLAQGINFG 1174
SD+WK YMRRAT T+NY LPLAQGI FG
Sbjct: 464 GGRESGSDAWKHYMRRATNTVNYSRALPLAQGIRFG 499
>sptr|O85725|O85725 Semialdehyde dehydrogenase Pcd.
Length = 512
Score = 360 bits (925), Expect = 2e-98
Identities = 181/394 (45%), Positives = 249/394 (63%), Gaps = 4/394 (1%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+ID+CD+AVGLSRQL G +PSERP H +ME W+PLGVVGVI+AFNFP AV WNA +
Sbjct: 117 QEMIDICDFAVGLSRQLYGRTMPSERPGHRLMETWHPLGVVGVISAFNFPVAVWAWNAAV 176
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
ALVCG+ VVWK + TPL +A ++ + P + G ++G+ + R+
Sbjct: 177 ALVCGDTVVWKPSELTPLTALACAALLDLAIADAGAPKGLNQVVVGAADVGERLVDSPRV 236
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
PLVS TGSTR G V +V+ARFG+ +LEL GNNA +V AD+ L V + +FAA GTAG
Sbjct: 237 PLVSATGSTRMGRAVGPRVAARFGRTILELGGNNAAVVTPSADLDLTVNAAVFAAAGTAG 296
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTT RRLI+HE+I T +++L ++++ IGDP + TL+GPL A+ + ++
Sbjct: 297 QRCTTLRRLIVHEDIADTVVERLTAAFERLPIGDPFQDTTLVGPLVNEAAFGRMREAVER 356
Query: 722 IKSQGGKILFGGSA----IESEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEA 889
++GG + GG +V+P +V + VV+EE F P+LYV+ ++ L EA
Sbjct: 357 ATAEGGTLCAGGERQFPDAAPGAYYVRPALVRMPAQTAVVREETFAPILYVLTYRDLDEA 416
Query: 890 IEINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXX 1069
I +NN VPQGLS+ IFT ++L P G+DCGI NVNI T+GAEI
Sbjct: 417 IRLNNEVPQGLSAGIFTADQSEAERFLAPDGADCGIANVNIGTSGAEIGGAFGGEKETGG 476
Query: 1070 XXXXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+W+ YMRRAT T+NY + LAQG++F
Sbjct: 477 GRESGSDAWRAYMRRATNTVNYSGRVALAQGVDF 510
>sptr|Q8U8D9|Q8U8D9 Aldehyde dehydrogenase.
Length = 511
Score = 356 bits (914), Expect = 5e-97
Identities = 186/394 (47%), Positives = 253/394 (64%), Gaps = 4/394 (1%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+ID+CD+AVGLSRQL G I +ERP H MME W+PLGVVGVI+AFNFP AV WNA +
Sbjct: 116 QEMIDICDFAVGLSRQLYGLTIATERPGHRMMETWHPLGVVGVISAFNFPVAVWSWNAAL 175
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEK-NNLPGAIFTSFCGGTEIGQAIALDIR 358
ALVCGN VVWK + TPL +A+ I + + P + G +G+A+ +
Sbjct: 176 ALVCGNSVVWKPSEKTPLTALAVQGIFERAAARFGDAPEGLSQLLIGDRAVGEAMVDYPK 235
Query: 359 IPLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTA 538
+PLVS TGSTR G V +++ RF + +LEL GNNA IV AD+ +A+R++ F A+GTA
Sbjct: 236 VPLVSATGSTRMGRDVGPRLAKRFARAILELGGNNAGIVCPSADLDMALRAIAFGAMGTA 295
Query: 539 GQRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQ 718
GQRCTT RRL +H+++Y+ + +L + Y V +G+PLE L+GPL A+ + K I+
Sbjct: 296 GQRCTTLRRLFVHDSVYEELVPRLKKAYASVSVGNPLESSALVGPLVDKAAFDGMQKAIE 355
Query: 719 TIKSQGGKILFGGSAIE---SEGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKEA 889
K+ GG ++ GG ++ ++ +V+P +VE+ V EE F P+LYVMK+ L +A
Sbjct: 356 AAKAAGG-VVHGGDRVDTGAADAYYVKPALVEMPKQVGPVLEETFAPILYVMKYSDLDQA 414
Query: 890 IEINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXXX 1069
I+ +N+V GLSSSIFT+ ++L GSDCGI NVNI T+GAEI
Sbjct: 415 IDAHNAVAAGLSSSIFTRDIQESERFLSSEGSDCGIANVNIGTSGAEIGGAFGGEKETGG 474
Query: 1070 XXXXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SDSWK YMRRAT TINY LPLAQG++F
Sbjct: 475 GRESGSDSWKAYMRRATNTINYSKALPLAQGVSF 508
>sptrnew|AAS05963|AAS05963 AldB.
Length = 509
Score = 353 bits (907), Expect = 3e-96
Identities = 185/396 (46%), Positives = 250/396 (63%), Gaps = 5/396 (1%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+ID+C +AVGLSRQL G I SERP H + E W+PLGVVGVITAFNFP AV WNA +
Sbjct: 116 QEMIDICQFAVGLSRQLYGRTIASERPGHRLSETWHPLGVVGVITAFNFPVAVWAWNAAL 175
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEKNNLPGAIFTSFCGGTEIGQAIALDIRI 361
ALVCG+ VVWK + TPL +A ++A P + G E+G+ + D R+
Sbjct: 176 ALVCGDTVVWKPSELTPLTALATQALIARAAADVGAPPGVSAVVLGDRELGELLVDDERV 235
Query: 362 PLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGTAG 541
LVS TGS R G V +V+ARFG+ LLEL GNNA +V AD+ LAVR+V+FAA GTAG
Sbjct: 236 ALVSATGSVRMGRAVGPRVAARFGRVLLELGGNNAAVVTPSADLDLAVRAVVFAAAGTAG 295
Query: 542 QRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGIQT 721
QRCTT RRLI+H ++ + ++V +++ IGDP E GTL+GPL + + + +++
Sbjct: 296 QRCTTLRRLIVHRSVADEVVARVVSALRRLPIGDPFEPGTLVGPLIHETAYRDMVGALES 355
Query: 722 IKSQGGKILFGGS----AIESEGN-FVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKE 886
++ GG+++ G A+E + +V P +V + P+V E F P+LYV+ + L +
Sbjct: 356 ARADGGEVIDGDRRHPFAVEHPSSYYVAPAVVRMPAQTPIVATETFAPILYVLTYDRLDD 415
Query: 887 AIEINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXX 1066
AI +NN+VPQGLSS+IFT ++L SDCGI NVNI T+GAEI
Sbjct: 416 AIALNNAVPQGLSSAIFTTDLREAERFLDE--SDCGIANVNIGTSGAEIGGAFGGEKQTG 473
Query: 1067 XXXXXXSDSWKQYMRRATCTINYGSELPLAQGINFG 1174
SDSWK YMR++T T+NY LPLAQG+ FG
Sbjct: 474 GGRESGSDSWKAYMRQSTNTVNYSGALPLAQGVEFG 509
>sptr|Q9I4U7|Q9I4U7 Probable aldehyde dehydrogenase.
Length = 529
Score = 352 bits (902), Expect = 1e-95
Identities = 191/395 (48%), Positives = 241/395 (61%), Gaps = 5/395 (1%)
Frame = +2
Query: 2 REIIDMCDYAVGLSRQLNGSIIPSERPNHMMMEVWNPLGVVGVITAFNFPCAVLGWNACI 181
+E+ID+CD+AVGLSRQL G I SERP H M E W+PLGVVGVI+AFNFP AV WN +
Sbjct: 135 QEMIDICDFAVGLSRQLYGLTIASERPGHHMRETWHPLGVVGVISAFNFPVAVWSWNTAL 194
Query: 182 ALVCGNCVVWKGAPTTPLITIAMTKIVASVLEK--NNLPGAIFTSFCGGTEIGQAIALDI 355
ALVCGN VVWK + TPL +A + + P + G E G+A+
Sbjct: 195 ALVCGNSVVWKPSEKTPLTALACQALFERAARAFGGDAPAHLSQLLIGDREAGEALVDAR 254
Query: 356 RIPLVSFTGSTRAGLMVQQQVSARFGKCLLELSGNNAIIVMDDADIQLAVRSVLFAAVGT 535
R+ LVS TGSTR G V +V+ARF +C+LEL GNNA+I+ AD+ LAVR +LF AVGT
Sbjct: 255 RVALVSATGSTRMGREVAPRVAARFARCILELGGNNAMILAPSADLDLAVRGILFGAVGT 314
Query: 536 AGQRCTTCRRLILHENIYQTFLDQLVEVYKQVRIGDPLEKGTLLGPLHTPASKENFLKGI 715
AGQRCTT RRLI HE++ +++L Y +VRIG PLE G L+GPL S +
Sbjct: 315 AGQRCTTLRRLIAHESVKDEIVERLKAAYSRVRIGHPLE-GNLVGPLIDERSYLAMQDAL 373
Query: 716 QTIKSQGGKILFGGSAIES---EGNFVQPTIVEITPSAPVVKEELFGPVLYVMKFQTLKE 886
+ QGG++ G ++ + +V P IVE+ VV+ E F P+LYV+ ++ E
Sbjct: 374 ARAREQGGRVFGGERQLQERYPDAYYVSPAIVEMPGQTEVVRTETFAPILYVVGYRDFDE 433
Query: 887 AIEINNSVPQGLSSSIFTKRPDIIFKWLGPHGSDCGIVNVNIPTNGAEIXXXXXXXXXXX 1066
A+ +NN VPQGLSS IFT + G GSDCGI NVNI T+GAEI
Sbjct: 434 ALRLNNEVPQGLSSCIFTTDLREAELFQGAAGSDCGIANVNIGTSGAEIGGAFGGEKETG 493
Query: 1067 XXXXXXSDSWKQYMRRATCTINYGSELPLAQGINF 1171
SD+WK YMRR T T+NY ELPLAQGI F
Sbjct: 494 GGRESGSDAWKAYMRRQTNTVNYSRELPLAQGITF 528
Database: /db/uniprot/tmp/swall
Posted date: Mar 5, 2004 7:45 PM
Number of letters in database: 442,889,342
Number of sequences in database: 1,395,590
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,226,724,253
Number of Sequences: 1395590
Number of extensions: 26910823
Number of successful extensions: 67487
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 60373
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 64410
length of database: 442,889,342
effective HSP length: 127
effective length of database: 265,649,412
effective search space used: 95368138908
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

