BLASTX 2.2.6 [Apr-09-2003]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= 131537.2.8
(708 letters)
Database: /db/trembl-ebi/tmp/swall
1,288,562 sequences; 411,996,502 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
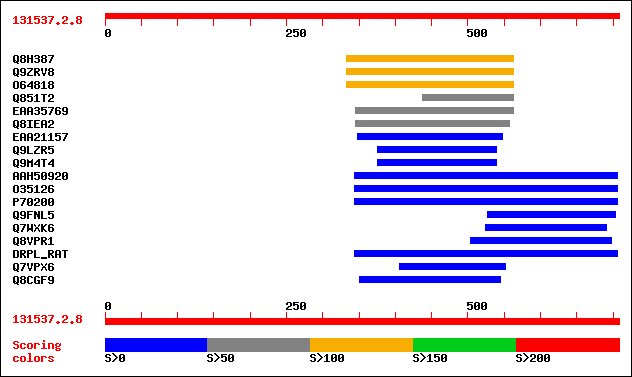
sptr|Q8H387|Q8H387 OJ1513_F02.31 protein. 147 2e-34
sptr|Q9ZRV8|Q9ZRV8 Hypothetical protein. 142 3e-33
sptr|O64818|O64818 At2g23090 protein (At2g23090/F21P24.15) (Hypo... 135 7e-31
sptr|Q851T2|Q851T2 Hypothetical protein. 74 2e-12
sptrnew|EAA35769|EAA35769 Predicted protein. 67 3e-10
sptr|Q8IEA2|Q8IEA2 Hypothetical protein, conserved. 63 3e-09
sptrnew|EAA21157|EAA21157 Hypothetical protein. 49 8e-05
sptr|Q9LZR5|Q9LZR5 Histone deacetylase-like protein. 34 2.1
sptr|Q9M4T4|Q9M4T4 Putative histone deacetylase HD2c (AT5g03740/... 34 2.1
sptrnew|AAH50920|AAH50920 Hypothetical protein. 33 3.6
sptr|O35126|O35126 DRPLA (Dentatorubral pallidoluysian atrophy). 33 3.6
sptr|P70200|P70200 DRPLA protein. 33 3.6
sptr|Q9FNL5|Q9FNL5 Similarity to guanylate binding protein. 33 4.7
sptr|Q7WXK6|Q7WXK6 Putative methyltransferase. 33 4.7
sptr|Q8VPR1|Q8VPR1 MC8. 32 6.1
sw|P54258|DRPL_RAT Atrophin-1 (Dentatorubral-pallidoluysian atro... 32 6.1
sptr|Q7VPX6|Q7VPX6 Hypothetical protein. 32 8.0
sptr|Q8CGF9|Q8CGF9 Similar to hypothetical protein MGC15548. 32 8.0
>sptr|Q8H387|Q8H387 OJ1513_F02.31 protein.
Length = 77
Score = 147 bits (370), Expect = 2e-34
Identities = 65/77 (84%), Positives = 73/77 (94%)
Frame = -2
Query: 563 MGGGNGQKSKMARERNLEKNKGAKGSQLETNKKAMSIQCKVCMQTFMCTTTEVKCREHAE 384
MGGGNGQKSKMARERN+EKNKGAKGSQLE NKKAM+IQCK+CMQTF+CTT+E KC+EHAE
Sbjct: 1 MGGGNGQKSKMARERNMEKNKGAKGSQLEANKKAMNIQCKICMQTFICTTSETKCKEHAE 60
Query: 383 AKHPKTDVYQCFPHLKK 333
AKHPK+D+ CFPHLKK
Sbjct: 61 AKHPKSDLTACFPHLKK 77
>sptr|Q9ZRV8|Q9ZRV8 Hypothetical protein.
Length = 78
Score = 142 bits (359), Expect = 3e-33
Identities = 68/78 (87%), Positives = 72/78 (92%), Gaps = 1/78 (1%)
Frame = -2
Query: 563 MGGGNGQKSKMARERNLEKNKGA-KGSQLETNKKAMSIQCKVCMQTFMCTTTEVKCREHA 387
MGGGNGQK+KMARERNLEK K A KGSQLETNKKAMSIQCKVCMQTF+CTT+EVKCREHA
Sbjct: 1 MGGGNGQKAKMARERNLEKQKNAGKGSQLETNKKAMSIQCKVCMQTFICTTSEVKCREHA 60
Query: 386 EAKHPKTDVYQCFPHLKK 333
EAKHPK+DV CFPHL K
Sbjct: 61 EAKHPKSDVLVCFPHLNK 78
>sptr|O64818|O64818 At2g23090 protein (At2g23090/F21P24.15)
(Hypothetical protein).
Length = 78
Score = 135 bits (339), Expect = 7e-31
Identities = 65/78 (83%), Positives = 68/78 (87%), Gaps = 1/78 (1%)
Frame = -2
Query: 563 MGGGNGQKSKMARERNLEKNKGA-KGSQLETNKKAMSIQCKVCMQTFMCTTTEVKCREHA 387
MGGGN QKS MAR +NLEK K A KGSQLE NKKAMSIQCKVCMQTF+CTT+EVKCREHA
Sbjct: 1 MGGGNAQKSAMARAKNLEKAKAAGKGSQLEANKKAMSIQCKVCMQTFICTTSEVKCREHA 60
Query: 386 EAKHPKTDVYQCFPHLKK 333
EAKHPK DV CFPHLKK
Sbjct: 61 EAKHPKADVVACFPHLKK 78
>sptr|Q851T2|Q851T2 Hypothetical protein.
Length = 42
Score = 73.6 bits (179), Expect = 2e-12
Identities = 35/42 (83%), Positives = 38/42 (90%)
Frame = -2
Query: 563 MGGGNGQKSKMARERNLEKNKGAKGSQLETNKKAMSIQCKVC 438
MGGGNGQKS+MARERN+EK KGAKGSQLETNKKAM+IQ C
Sbjct: 1 MGGGNGQKSRMARERNMEKAKGAKGSQLETNKKAMNIQVGSC 42
>sptrnew|EAA35769|EAA35769 Predicted protein.
Length = 80
Score = 66.6 bits (161), Expect = 3e-10
Identities = 34/73 (46%), Positives = 43/73 (58%)
Frame = -2
Query: 563 MGGGNGQKSKMARERNLEKNKGAKGSQLETNKKAMSIQCKVCMQTFMCTTTEVKCREHAE 384
MGGGNG K+ RERN + SQL+TN AM+I C+ C TF+ T+ EHA+
Sbjct: 1 MGGGNGAKAAQKRERNAKNAAAGPKSQLKTNAAAMNIICQTCRATFLSTSRAKALDEHAQ 60
Query: 383 AKHPKTDVYQCFP 345
KH KT + CFP
Sbjct: 61 NKHNKT-LADCFP 72
>sptr|Q8IEA2|Q8IEA2 Hypothetical protein, conserved.
Length = 78
Score = 63.2 bits (152), Expect = 3e-09
Identities = 35/71 (49%), Positives = 44/71 (61%)
Frame = -2
Query: 557 GGNGQKSKMARERNLEKNKGAKGSQLETNKKAMSIQCKVCMQTFMCTTTEVKCREHAEAK 378
G N K ARER + KG K SQL+ NK+A+++ CKVC FM T + + EHAE K
Sbjct: 4 GSNACKRNQARERKNVEVKGGK-SQLKANKEALNVTCKVCYTVFMQTQSISQLAEHAENK 62
Query: 377 HPKTDVYQCFP 345
H K DV +CFP
Sbjct: 63 HNK-DVKECFP 72
>sptrnew|EAA21157|EAA21157 Hypothetical protein.
Length = 71
Score = 48.5 bits (114), Expect = 8e-05
Identities = 24/68 (35%), Positives = 38/68 (55%), Gaps = 1/68 (1%)
Frame = -2
Query: 548 GQKSKMARERNLEKNKGAK-GSQLETNKKAMSIQCKVCMQTFMCTTTEVKCREHAEAKHP 372
G + + R R+ ++N+ AK S ++ K +++ CK+C TFMCT + +EH E KH
Sbjct: 4 GNQRDVDRIRSSKRNEKAKPNSTVKDASKNLNVICKLCRHTFMCTVNQSILKEHHEKKHS 63
Query: 371 KTDVYQCF 348
K CF
Sbjct: 64 KHAYEDCF 71
>sptr|Q9LZR5|Q9LZR5 Histone deacetylase-like protein.
Length = 296
Score = 33.9 bits (76), Expect = 2.1
Identities = 19/55 (34%), Positives = 29/55 (52%)
Frame = -2
Query: 539 SKMARERNLEKNKGAKGSQLETNKKAMSIQCKVCMQTFMCTTTEVKCREHAEAKH 375
SK A + + + G Q +T K A + CK C +TF T+E+ + H +AKH
Sbjct: 241 SKQAGKNSGGGSTGETSKQQQTPKSAGAFGCKSCTRTF---TSEMGLQSHTKAKH 292
>sptr|Q9M4T4|Q9M4T4 Putative histone deacetylase HD2c
(AT5g03740/F17C15_160) (HDA11).
Length = 294
Score = 33.9 bits (76), Expect = 2.1
Identities = 19/55 (34%), Positives = 29/55 (52%)
Frame = -2
Query: 539 SKMARERNLEKNKGAKGSQLETNKKAMSIQCKVCMQTFMCTTTEVKCREHAEAKH 375
SK A + + + G Q +T K A + CK C +TF T+E+ + H +AKH
Sbjct: 239 SKQAGKNSGGGSTGETSKQQQTPKSAGAFGCKSCTRTF---TSEMGLQSHTKAKH 290
>sptrnew|AAH50920|AAH50920 Hypothetical protein.
Length = 1175
Score = 33.1 bits (74), Expect = 3.6
Identities = 30/127 (23%), Positives = 44/127 (34%), Gaps = 6/127 (4%)
Frame = -3
Query: 706 AGTDPPERSGLPFPRSFP*TTNLTRSGAGRPPGEVRVPAVDRRRIHPPWAVATARSP--R 533
A PP P P P + + +G PPG + P + P A ++A P R
Sbjct: 315 ASASPPGMGAQPIPGHLPSPHAMGQGMSGLPPGPEKGPTLAPSPHPLPPASSSAPGPPMR 374
Query: 532 WPASATWRRTXXXXXXXXXXXXXXXXXXAKCACKHSCVPRLK*SAGSTP----RPSIPRQ 365
+P S++ A + HS P S + P +PS+P Q
Sbjct: 375 YPYSSSSSSAAASSSSSSSSASQYPASQALPSYPHSFPPPTSMSVSNQPPKYTQPSLPSQ 434
Query: 364 TCTSASP 344
S P
Sbjct: 435 AVWSQGP 441
>sptr|O35126|O35126 DRPLA (Dentatorubral pallidoluysian atrophy).
Length = 1175
Score = 33.1 bits (74), Expect = 3.6
Identities = 30/127 (23%), Positives = 44/127 (34%), Gaps = 6/127 (4%)
Frame = -3
Query: 706 AGTDPPERSGLPFPRSFP*TTNLTRSGAGRPPGEVRVPAVDRRRIHPPWAVATARSP--R 533
A PP P P P + + +G PPG + P + P A ++A P R
Sbjct: 315 ASASPPGMGAQPIPGHLPSPHAMGQGMSGLPPGPEKGPTLAPSPHPLPPASSSAPGPPMR 374
Query: 532 WPASATWRRTXXXXXXXXXXXXXXXXXXAKCACKHSCVPRLK*SAGSTP----RPSIPRQ 365
+P S++ A + HS P S + P +PS+P Q
Sbjct: 375 YPYSSSSSSAAASSSSSSSSASQYPASQALPSYPHSFPPPTSMSVSNQPPKYTQPSLPSQ 434
Query: 364 TCTSASP 344
S P
Sbjct: 435 AVWSQGP 441
>sptr|P70200|P70200 DRPLA protein.
Length = 1175
Score = 33.1 bits (74), Expect = 3.6
Identities = 30/127 (23%), Positives = 44/127 (34%), Gaps = 6/127 (4%)
Frame = -3
Query: 706 AGTDPPERSGLPFPRSFP*TTNLTRSGAGRPPGEVRVPAVDRRRIHPPWAVATARSP--R 533
A PP P P P + + +G PPG + P + P A ++A P R
Sbjct: 315 ASASPPGMGAQPIPGHLPSPHAMGQGMSGLPPGPEKGPTLAPSPHPLPPASSSAPGPPMR 374
Query: 532 WPASATWRRTXXXXXXXXXXXXXXXXXXAKCACKHSCVPRLK*SAGSTP----RPSIPRQ 365
+P S++ A + HS P S + P +PS+P Q
Sbjct: 375 YPYSSSSSSAAASSSSSSSSASQYPASQALPSYPHSFPPPTSMSVSNQPPKYTQPSLPSQ 434
Query: 364 TCTSASP 344
S P
Sbjct: 435 AVWSQGP 441
>sptr|Q9FNL5|Q9FNL5 Similarity to guanylate binding protein.
Length = 1060
Score = 32.7 bits (73), Expect = 4.7
Identities = 22/62 (35%), Positives = 31/62 (50%), Gaps = 3/62 (4%)
Frame = -3
Query: 703 GTDPPERSGLPFPRSFP*TTNLTRSGAGRPPGEVRVPAVDRR---RIHPPWAVATARSPR 533
G D P S P PRS+P T+ + S PP +R+ D + R+ P AVAT + +
Sbjct: 9 GKDSPADSASPSPRSYPSTSPASSSAVTGPPRPIRLVYCDEKGKFRMDPE-AVATLQLVK 67
Query: 532 WP 527
P
Sbjct: 68 EP 69
>sptr|Q7WXK6|Q7WXK6 Putative methyltransferase.
Length = 64
Score = 32.7 bits (73), Expect = 4.7
Identities = 22/60 (36%), Positives = 26/60 (43%), Gaps = 4/60 (6%)
Frame = -3
Query: 691 PERSGLPFP-RSFP*TTNLTRSGAGRPPGEVRVPAVD---RRRIHPPWAVATARSPRWPA 524
PE GL R P + + AG P R+ AVD R PW V R P+WPA
Sbjct: 2 PEEQGLGHHHRLEPYSIAIALLDAGVPTARFRIDAVDVSARALARAPWRVRRQRLPQWPA 61
>sptr|Q8VPR1|Q8VPR1 MC8.
Length = 787
Score = 32.3 bits (72), Expect = 6.1
Identities = 23/65 (35%), Positives = 27/65 (41%)
Frame = -3
Query: 697 DPPERSGLPFPRSFP*TTNLTRSGAGRPPGEVRVPAVDRRRIHPPWAVATARSPRWPASA 518
DP R+ PR P +R AGRP VR P R+ PP + T R P
Sbjct: 77 DPRGRTAHARPRPRP----ASREEAGRPSPRVRRPPSVETRMPPPPSTKTGRGRGRPGGG 132
Query: 517 TWRRT 503
RRT
Sbjct: 133 MGRRT 137
>sw|P54258|DRPL_RAT Atrophin-1 (Dentatorubral-pallidoluysian atrophy
protein).
Length = 1183
Score = 32.3 bits (72), Expect = 6.1
Identities = 29/127 (22%), Positives = 43/127 (33%), Gaps = 6/127 (4%)
Frame = -3
Query: 706 AGTDPPERSGLPFPRSFP*TTNLTRSGAGRPPGEVRVPAVDRRRIHPPWAVATARSP--R 533
A PP P P P + + +G PPG + P + P A ++A P R
Sbjct: 314 ASASPPGMGAQPIPGHLPSPHAMGQGMSGLPPGPEKGPTLAPSPHPLPPASSSAPGPPMR 373
Query: 532 WPASATWRRTXXXXXXXXXXXXXXXXXXAKCACKHSCVPRLK*SAGSTP----RPSIPRQ 365
+P S+ + + HS P S + P +PS+P Q
Sbjct: 374 YPYSSCSSSSVAASSSSSAATSQYPASQTLPSYPHSFPPPTSMSVSNQPPKYTQPSLPSQ 433
Query: 364 TCTSASP 344
S P
Sbjct: 434 AVWSQGP 440
>sptr|Q7VPX6|Q7VPX6 Hypothetical protein.
Length = 148
Score = 32.0 bits (71), Expect = 8.0
Identities = 16/50 (32%), Positives = 28/50 (56%), Gaps = 1/50 (2%)
Frame = +1
Query: 406 TSVVVHMNVCMHTLHWMLMAFLLVSSWLPL-APLFFSKLRSRAILDFWPL 552
T+ V+ +++C +T + ++ L S+W PL PLF +L L WP+
Sbjct: 55 TTRVITLSICFYTFYLQIIFLFLYSAWKPLRQPLFCHRL-----LIIWPI 99
>sptr|Q8CGF9|Q8CGF9 Similar to hypothetical protein MGC15548.
Length = 585
Score = 32.0 bits (71), Expect = 8.0
Identities = 20/65 (30%), Positives = 28/65 (43%)
Frame = -2
Query: 545 QKSKMARERNLEKNKGAKGSQLETNKKAMSIQCKVCMQTFMCTTTEVKCREHAEAKHPKT 366
Q K + E LE LE +K +S QC C ++F V+ R H K +T
Sbjct: 490 QPMKKSEEETLETEGTGASDLLEKSKAKLSFQCGDCEKSFQRHDHLVRHRRHCHLK-DET 548
Query: 365 DVYQC 351
+QC
Sbjct: 549 RPFQC 553
Database: /db/trembl-ebi/tmp/swall
Posted date: Oct 24, 2003 8:22 PM
Number of letters in database: 411,996,502
Number of sequences in database: 1,288,562
Lambda K H
0.318 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 588,806,902
Number of Sequences: 1288562
Number of extensions: 12614479
Number of successful extensions: 43830
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 42152
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 43810
length of database: 411,996,502
effective HSP length: 119
effective length of database: 258,657,624
effective search space used: 30004284384
frameshift window, decay const: 50, 0.1
T: 12
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)

